Team:Imperial College/Growth Assays
From 2013.igem.org
| Line 56: | Line 56: | ||
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
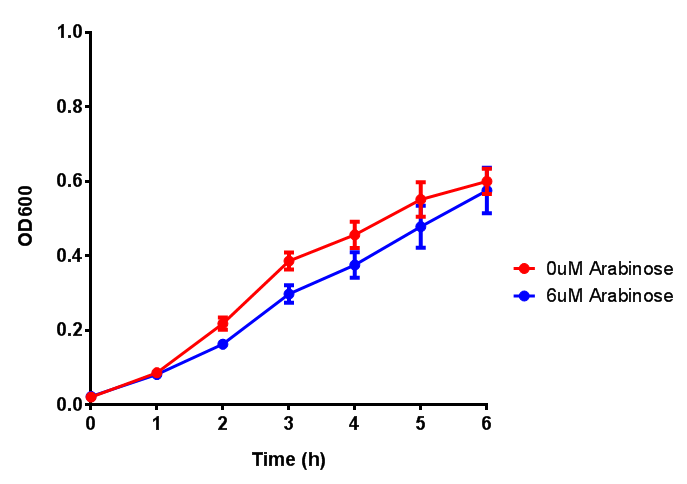
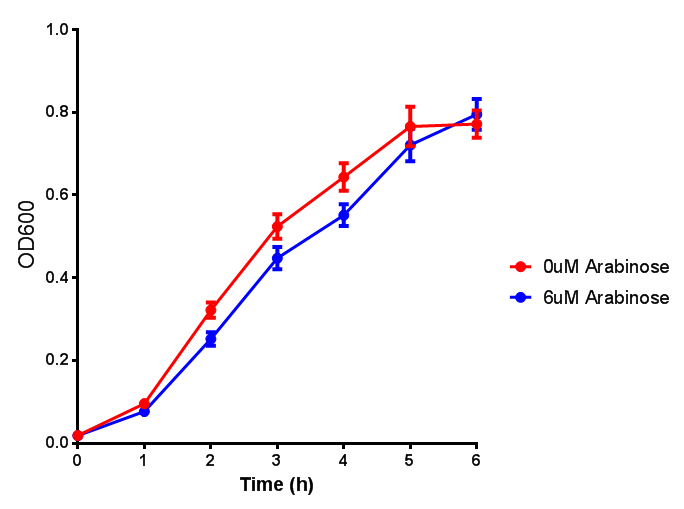
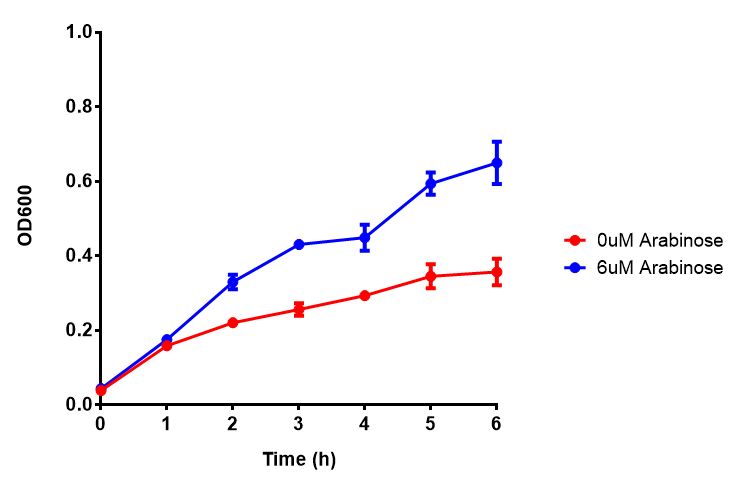
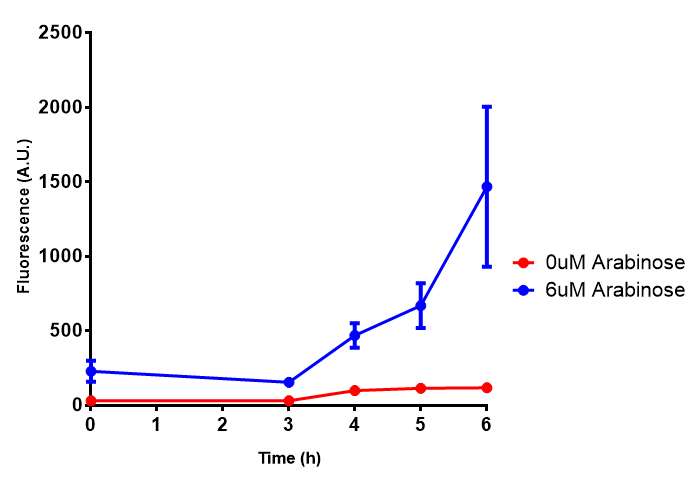
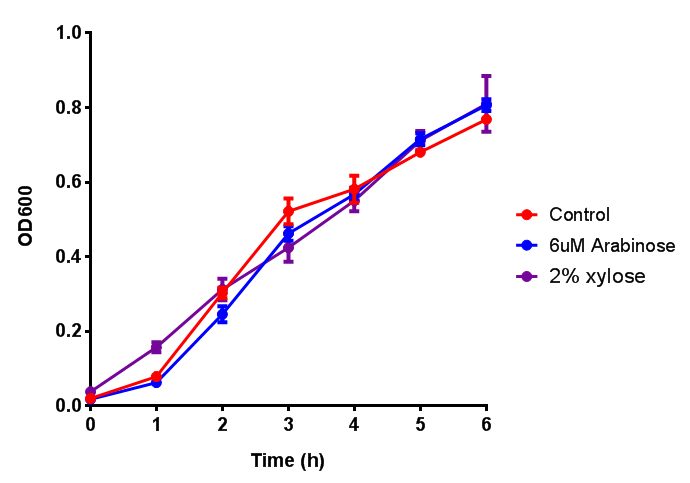
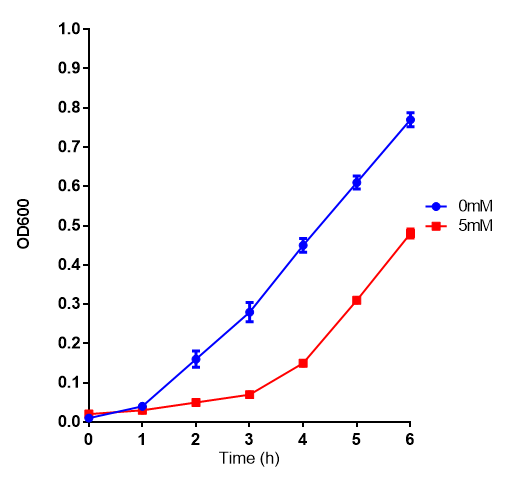
| - | | [[File:Bdh2.png|thumbnail|right|450px|<b>Bdh2 with pelB secretion tag growth assay.</b> E. coli (MG1655) transformed with pelB-bdh2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149013 BBa_K1149013] were grown with either 0 μM or 6 μM Arabinose to induce bdh2 and sfGFP expression. Bdh2 induction shows reduced growth of MG1655 but after 6h they reach the same OD. Growth was at 37°C with shaking. Error bars are SEM, n=4. ]] | + | | [[File:Bdh2.png|thumbnail|right|450px|<b>Bdh2 with pelB secretion tag growth assay.</b> E. coli (MG1655) transformed with pelB-bdh2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149013 BBa_K1149013] were grown with either 0 μM or 6 μM Arabinose to induce bdh2 and sfGFP expression. Bdh2 induction shows reduced growth of MG1655 but after 6h they reach the same OD, the decrease in growth is confirmed as not significant by a two-tailed t-test where p = 0.6964 > 0.05 (critical value). Growth was at 37°C with shaking. Error bars are SEM, n=4. ]] |
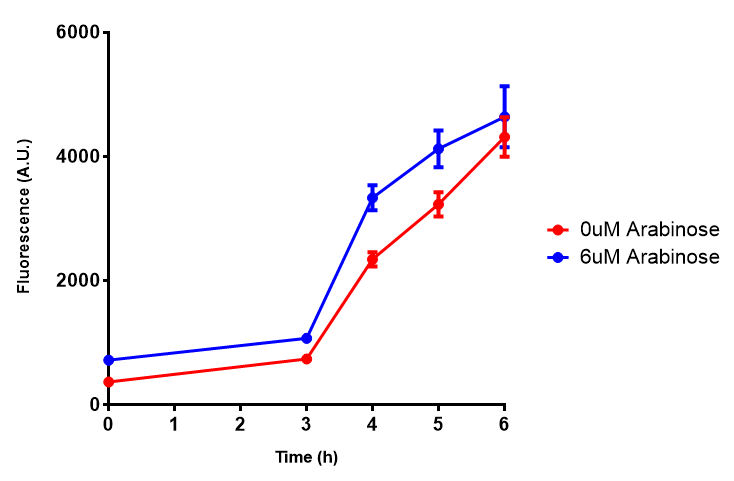
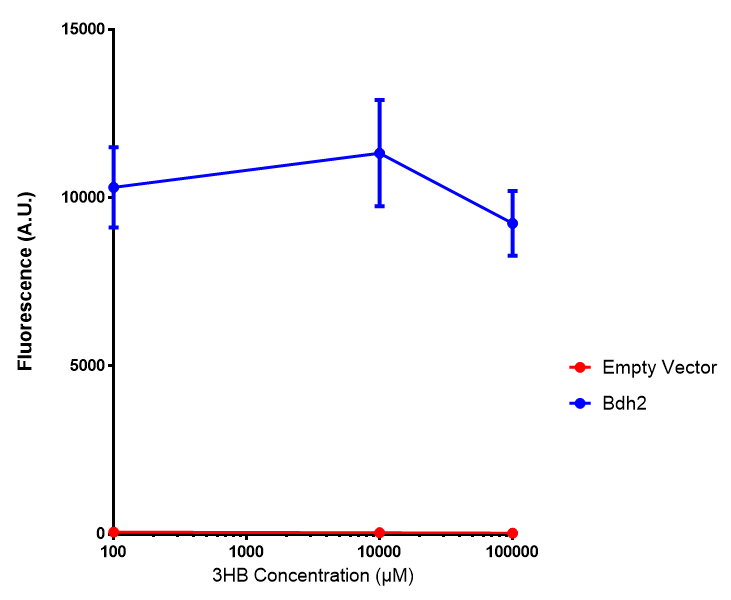
| [[File:Bdh2_fluor.png|thumbnail|right|450px|<b>Bdh2 with pelB secretion tag induction assay.</b> E. coli (MG1655) transformed with pelB-bdh2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149013 BBa_K1149013] were grown with either 0 μM or 6 μM Arabinose to induce bdh2 and sfGFP expression. We see that induction leads to stronger expression of bdh2 than without, although the promoter is leaky as the curve follows a similar pathway. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] | | [[File:Bdh2_fluor.png|thumbnail|right|450px|<b>Bdh2 with pelB secretion tag induction assay.</b> E. coli (MG1655) transformed with pelB-bdh2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149013 BBa_K1149013] were grown with either 0 μM or 6 μM Arabinose to induce bdh2 and sfGFP expression. We see that induction leads to stronger expression of bdh2 than without, although the promoter is leaky as the curve follows a similar pathway. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] | ||
|- | |- | ||
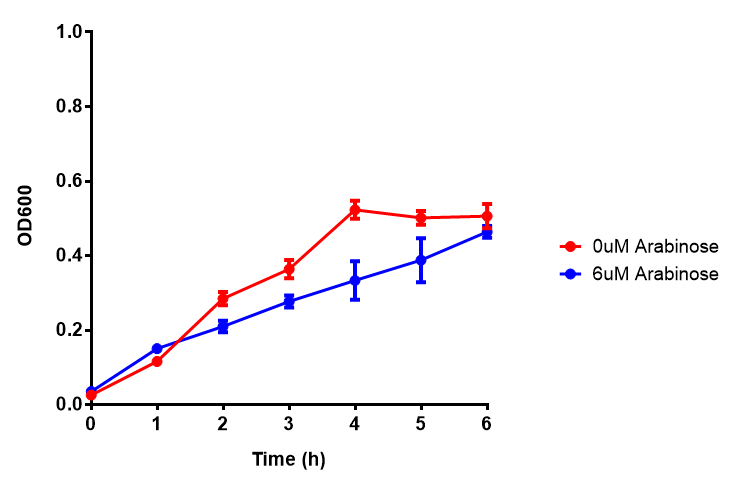
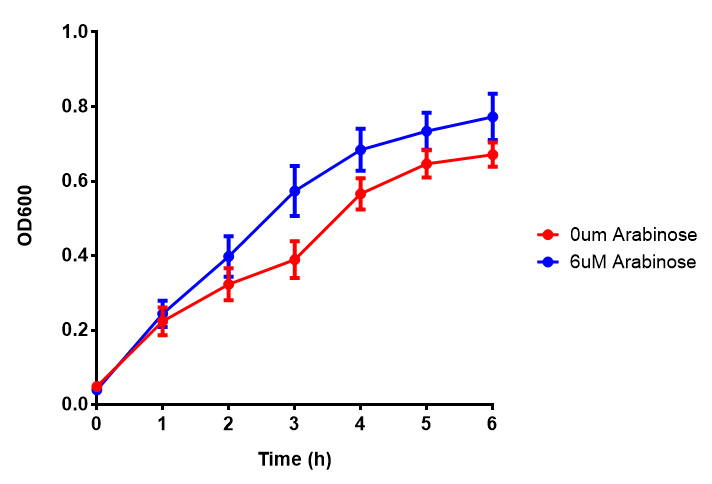
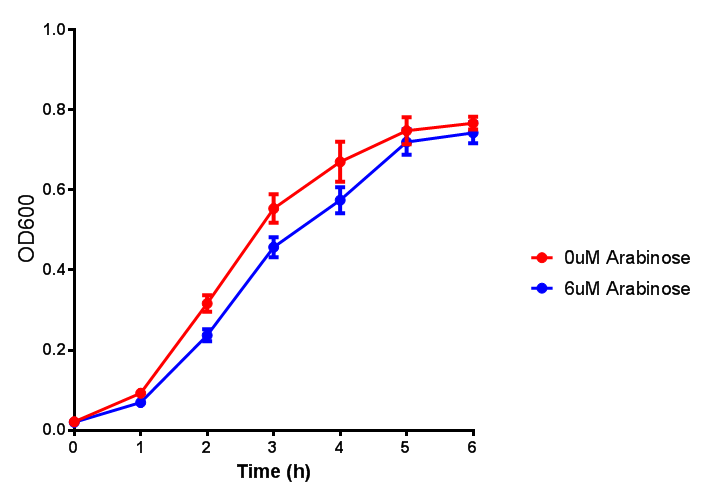
| - | |[[File:Bdh2-2.png|thumbnail|left|450px|<b>Bdh2 with no pelB secretion tag growth assay</b> MG1655 with intracellular bdh2 were grown over 6h to gauge the effect on growth when induced. Graph shows that while they reach the same end point, growth is initially faster without induction for [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149050 BBa_K1149050]. Error bars are SEM, n=4.]] | + | |[[File:Bdh2-2.png|thumbnail|left|450px|<b>Bdh2 with no pelB secretion tag growth assay</b> MG1655 with intracellular bdh2 were grown over 6h to gauge the effect on growth when induced. Graph shows that while they reach the same end point, growth is initially faster without induction for [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149050 BBa_K1149050], though this was not significant as a two-tailed t-test gave p = 0.4930 > 0.05, allowing us to accept the null hypothesis. Error bars are SEM, n=4.]] |
|[[File:Bdh2-2_fluor.png|thumbnail|left|450px|<b>Bdh2 with no pelB secretion tag induction assay</b> In addition to testing [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149050 BBa_K1149050] growth, we looked at induction whereby we see that more sfGFP is produced when induced. Although the promoter is leaky as the curve of intracellular bdh2, as seen with bdh2-pelB is followed tightly by the non-induced cells. Error bars are SEM, n=4.]] | |[[File:Bdh2-2_fluor.png|thumbnail|left|450px|<b>Bdh2 with no pelB secretion tag induction assay</b> In addition to testing [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149050 BBa_K1149050] growth, we looked at induction whereby we see that more sfGFP is produced when induced. Although the promoter is leaky as the curve of intracellular bdh2, as seen with bdh2-pelB is followed tightly by the non-induced cells. Error bars are SEM, n=4.]] | ||
|- | |- | ||
| Line 67: | Line 67: | ||
|} | |} | ||
| - | Ideally you would normalise the fluorescence with OD600 to give fluorescence/cell in order to get the actual difference in induction expression. | + | Ideally you would normalise the fluorescence with OD600 to give fluorescence/cell in order to get the actual difference in induction expression. A two-tailed t-test shows that p = 0.4308 > 0.05, therefore there is no statistical difference between induced and uninduced. |
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| - | | [[File:ESTCS2.png|thumbnail|right|450px|<b>ETSCS2 growth assay.</b> E. coli (MG1655) transformed with ESTCS2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149002 BBa_K1149002] were grown with either | + | |[[File:ESTCS2.png|thumbnail|right|450px|<b>ETSCS2 growth assay.</b> <i>E. coli</i> (MG1655) transformed with ESTCS2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149002 BBa_K1149002] were grown with either 0 μM or 6 μM Arabinose to induce ESTC2 and sfGFP expression. The growth curves appear very similar, which is confirmed by a two-tailed t-test p-value = 0.8118, thus no reason to say that there is any growth inhibition. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] |
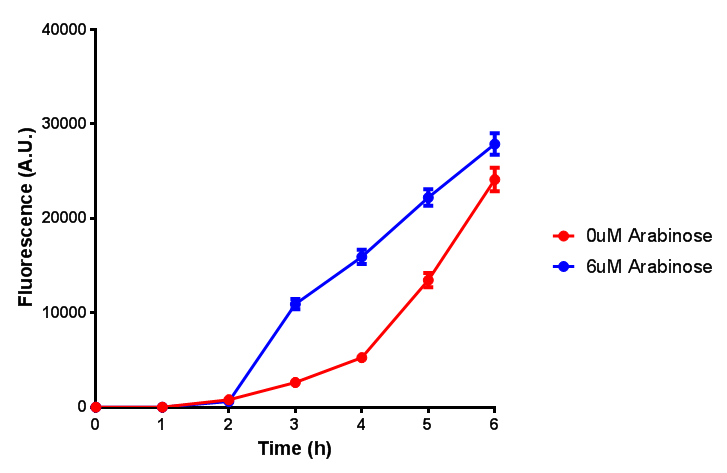
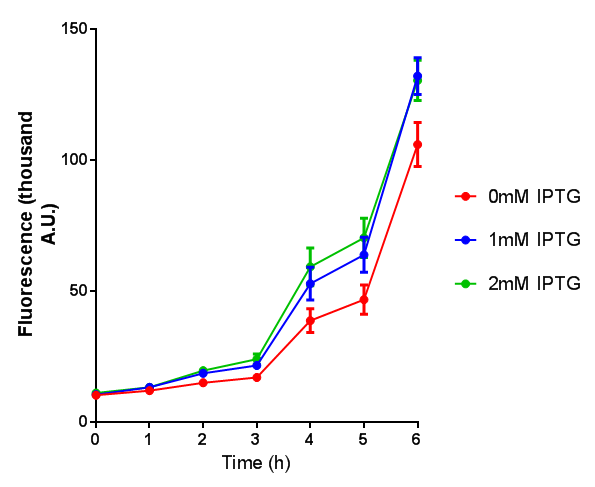
| - | | [[File:ESTCS2_fluor.png|thumbnail|right|450px|<b>ETSCS2 induction assay.</b> E. coli (MG1655) transformed with ESTCS2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149002 BBa_K1149002] were grown with either 0uM or 6uM Arabinose to induce ESTC2 and sfGFP expression. Induction by Arabinose produces a strong response, with induced transformants fluorescing more. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] | + | |[[File:ESTCS2_fluor.png|thumbnail|right|450px|<b>ETSCS2 induction assay.</b> <i>E. coli</i> (MG1655) transformed with ESTCS2 [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1149002 BBa_K1149002] were grown with either 0uM or 6uM Arabinose to induce ESTC2 and sfGFP expression. Induction by Arabinose produces a strong response, with induced transformants fluorescing more. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] |
|- | |- | ||
| - | |[[File:PueA.png|thumbnail|right|450px|<b>PueA growth assay</b> E.]] | + | |[[File:PueA.png|thumbnail|right|450px|<b>PueA growth assay.</b> MG1655 <i>E. coli</i> were grown for 6h with 0 μM or 6 μM Arabinose. There was no difference between no induction and induction as a t-test gave p = 0.5559 > 0.05. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] |
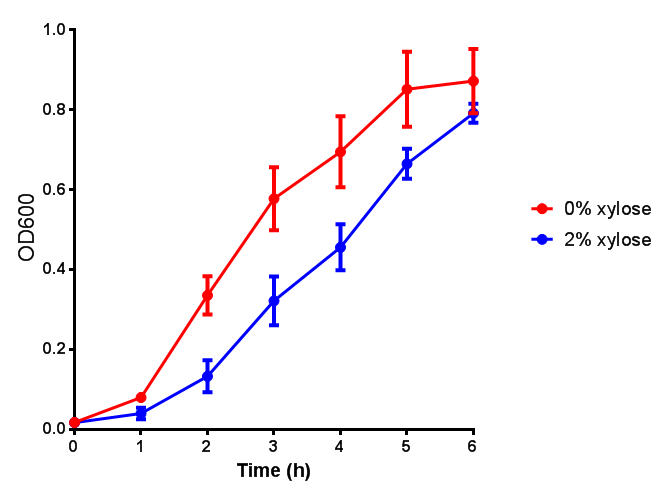
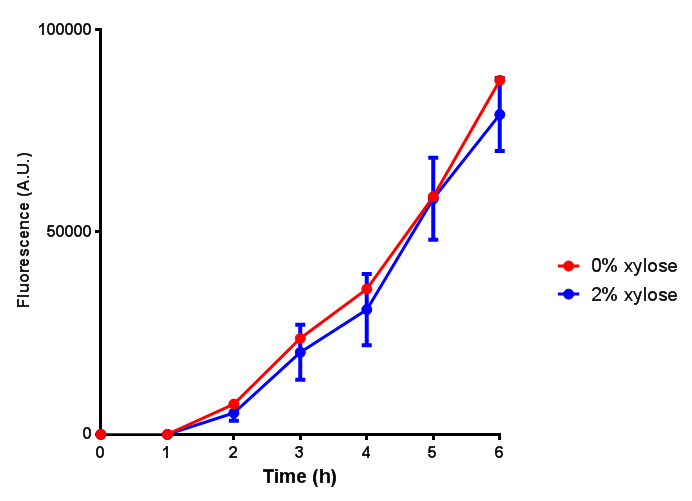
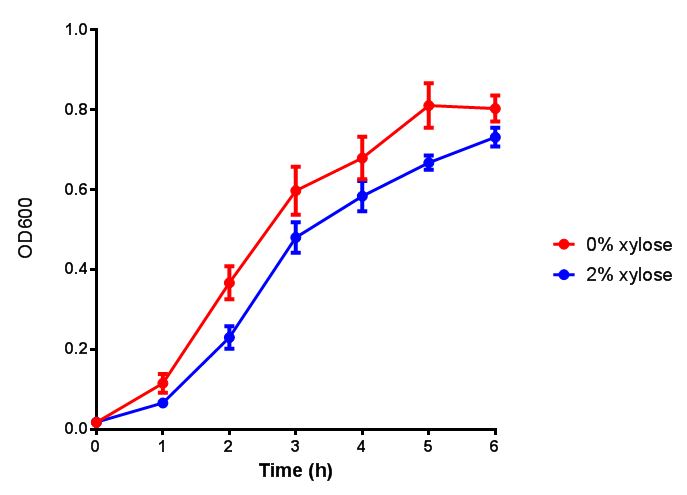
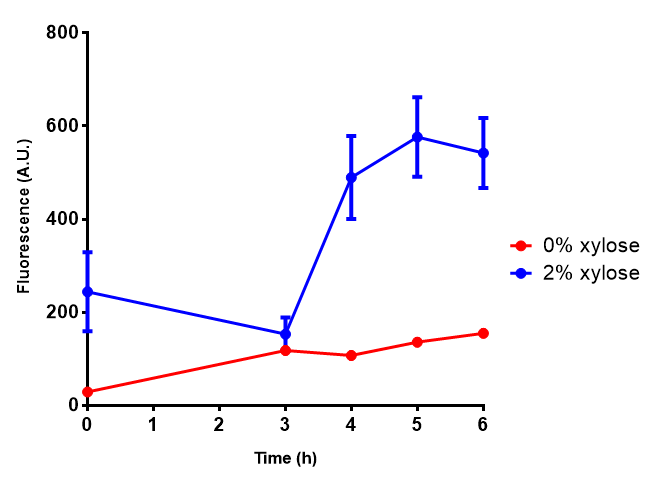
| - | |[[File:PueB.png|thumbnail|right|450px|<b>PueB growth assay</b> E. coli (MG1655) transformed with PueB (BBa_K1149004) were grown with either 0% or 2% Xylose to induce PueB expression. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] | + | |[[File:PueB.png|thumbnail|right|450px|<b>PueB growth assay.</b> <i>E. coli</i> (MG1655) transformed with PueB (BBa_K1149004) were grown with either 0% or 2% Xylose to induce PueB expression. There was no difference between no induction and induction as a t-test gave p = 0.6040 > 0.05. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] |
|- | |- | ||
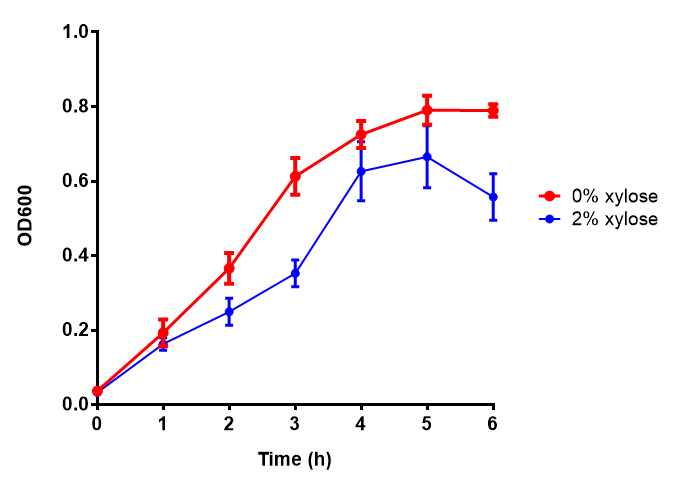
| - | |[[File:phaz1.png|thumbnail|left|450px|<b>Phaz1 growth assay</b> E. | + | |[[File:phaz1.png|thumbnail|left|450px|<b>Phaz1 growth assay.</b> MG1655 <i>E. coli</i> were grown for 6h with 0% and 2% Xylose to induce Phaz1. A two-tailed t-test, with p = 0.4181 shows that there is no significant difference between induction and no induction with regards to growth inhibition, as the value exceeds the critical value of 0.05. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] |
| - | |[[File:PulA.png|thumbnail|left|450px|<b>PulA growth assay</b> E. coli (MG1655) transformed with PulA (BBa_K1149006) were grown with either | + | |[[File:PulA.png|thumbnail|left|450px|<b>PulA growth assay.</b> <i>E. coli</i> (MG1655) transformed with PulA (BBa_K1149006) were grown with either 0 μM or 6 μM Arabinose to induce PulA expression. There was no growth inhibition as p = 0.7648 > 0.05, therefore induction did not cause significant growth decrease. Growth was at 37°C with shaking. Error bars are SEM, n=4.]] |
|- | |- | ||
|[[File:PK.png|thumbnail|left|450px|<b>Proteinase K growth assay</b> E. c]] | |[[File:PK.png|thumbnail|left|450px|<b>Proteinase K growth assay</b> E. c]] | ||
Revision as of 17:40, 1 October 2013
Contents |
Growth and Toxicity Assays
This page includes all of our experimental growth, toxicity and sole carbon source assay data.
Growth assays with different experimental media
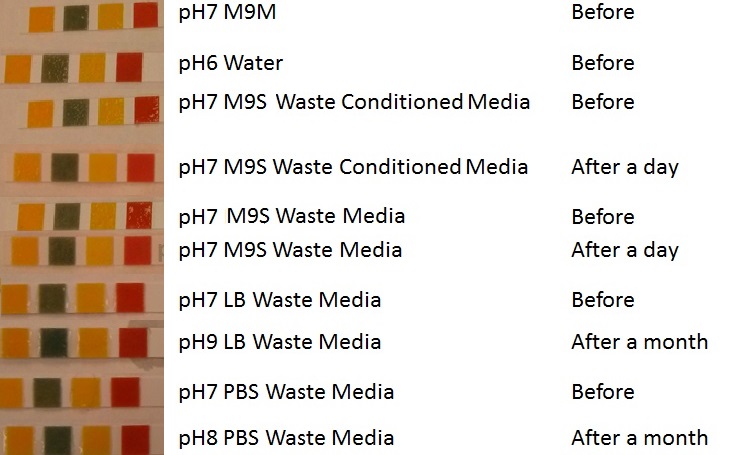
In additional to standard LB and minimal media, several novel experimental media were developed in order to characterise Biobricks within a mixed waste/landfill setting. These media were characterised through an examination of pH and through an array of growth assays with the project chassis, E.coli (MG1655).
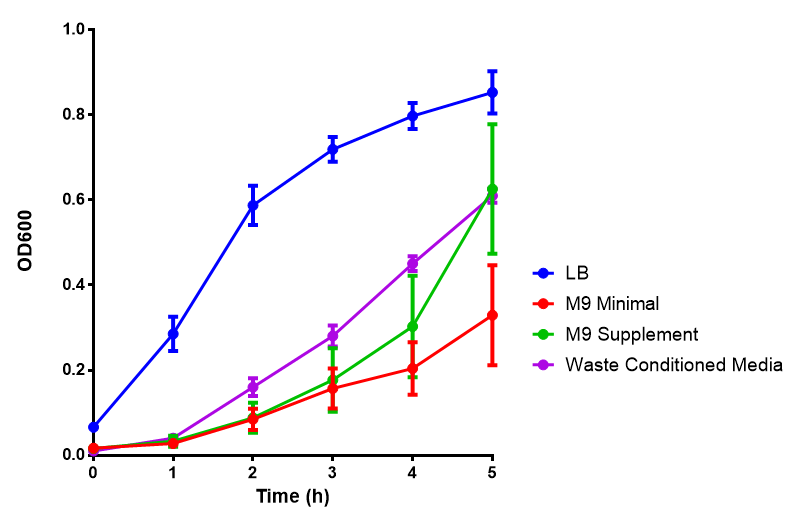 Media characterisation. E. coli strain MG1655 were transformed with a control plasmid and grown in different experimental media over a period of 5 hours. LB media, minimal media (M9M), supplemented minimal media (M9S), as described here or waste conditioned media (WCM), which is made from sterile filtrated mixed waste, see here. OD600 measured, error bars are S.E.M., n=4. |
Conclusion: MG1655 E. coli are viable and grow in all of our experimental medias. We have established a novel media that is optimised for characterisation of biobricks within a mixed waste/landfill context.
Long term waste growth assays
These assays were designed to test whether our chassis, E. coli (MG1655) could grow directly with waste over a long period of time.
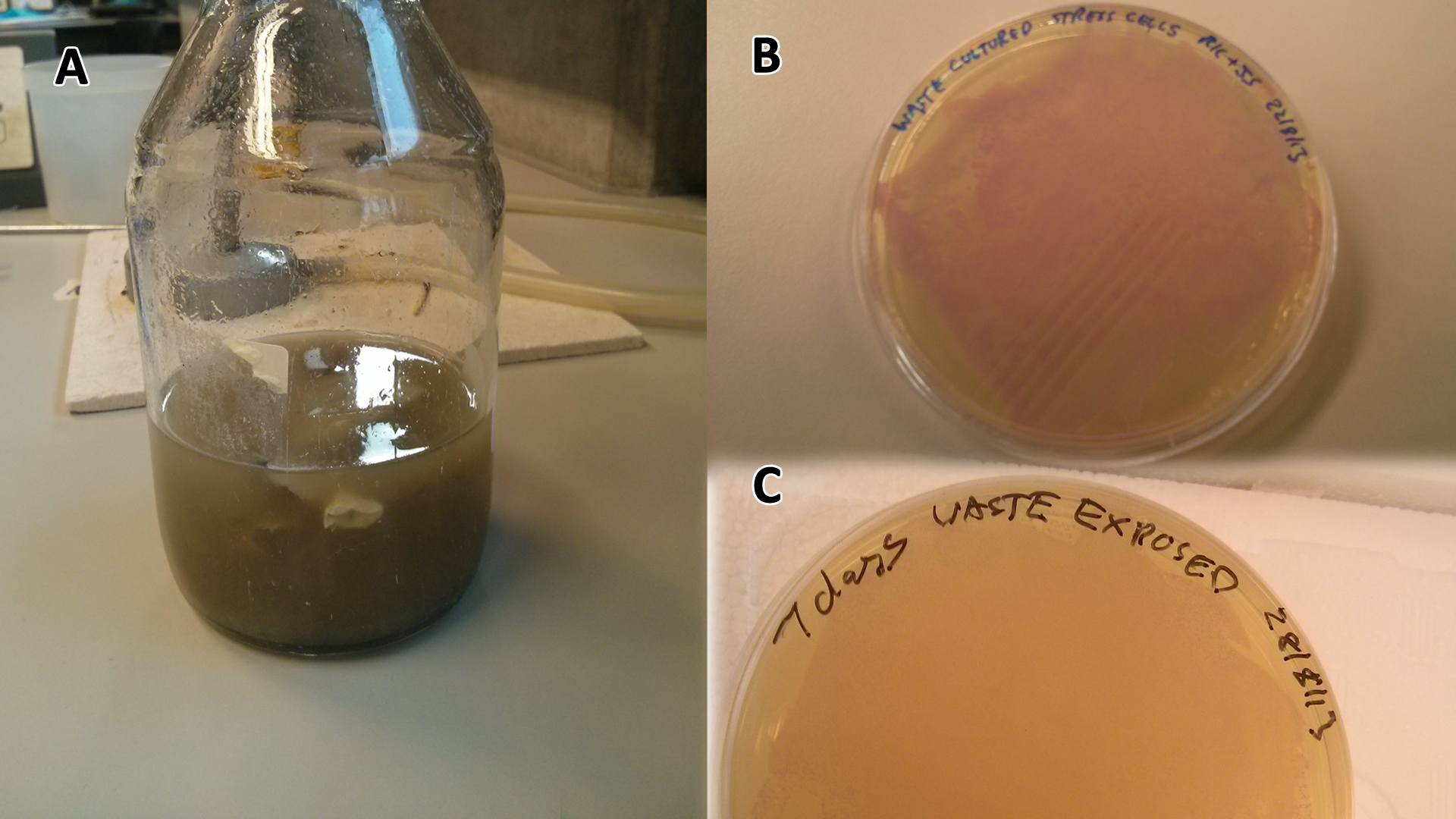
Waste media
Conclusion: MG1655 E. coli are viable and grow on mixed waste alone. Therefore we have established that our chassis could survive in a mixed waste bio-reactor context, which is validation of our concept to industrially implement our system.
Waste conditioned media
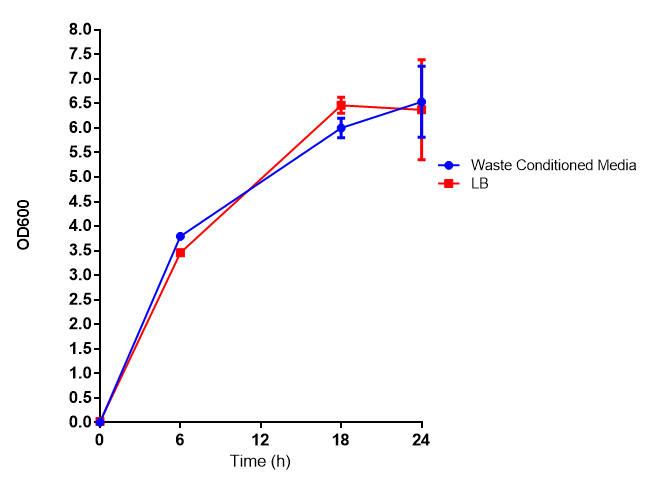
These assays were designed to test whether our chassis, E. coli (MG1655) could grow with waste conditioned media (WCM) over a period of 24-48 hours. Waste conditioned media is a filter sterilised version of the waste media and was designed for several reasons; Firstly we were unsure whether mixed waste would be toxic to Ecoli and hence a less concentrated version may be more suitable and secondly large chunks of waste would prevent accurate OD600 measurements and therefore we decided to filter out the largest chunks.
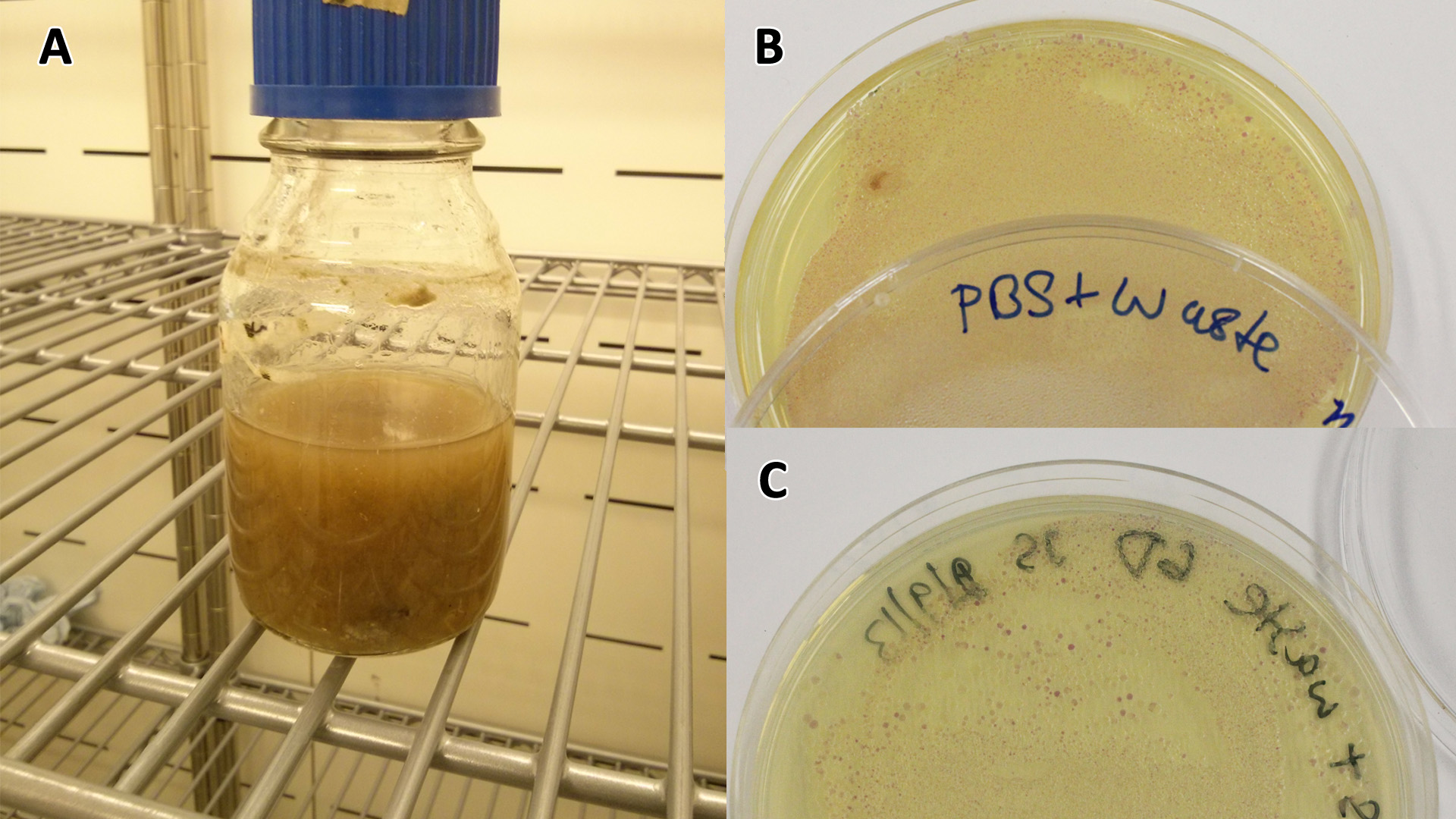
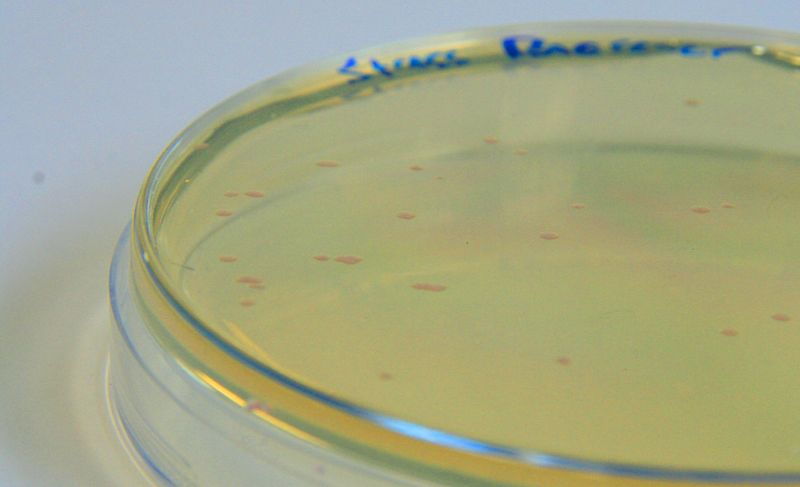
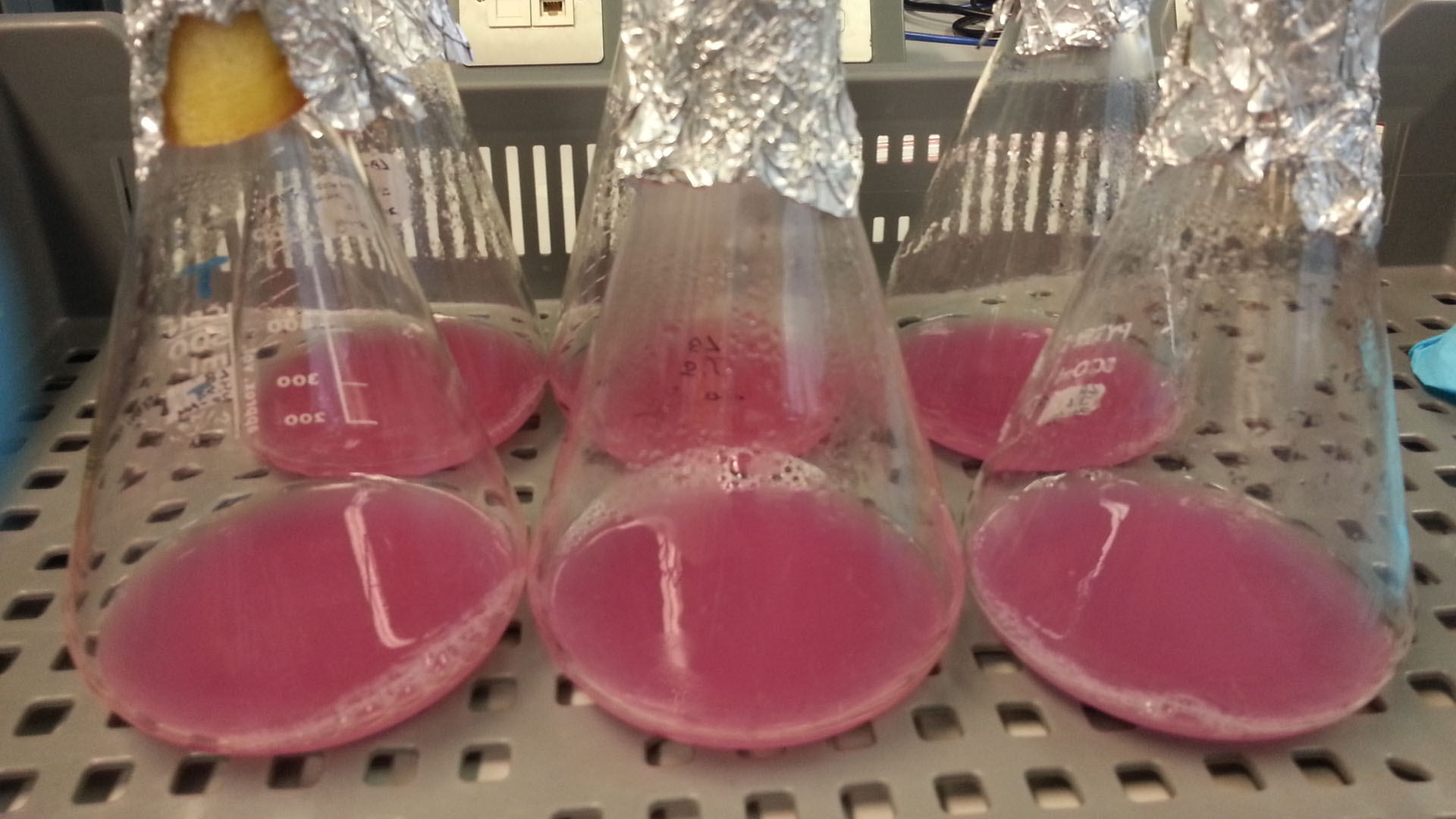 Photograph of waste conditioned media cultures mCherry stress biosensor (BBa_K639003) transformed MG1655 were grown with LB-WCM at 37ºC, with shaking for 48 hours. | |
Conclusion: MG1655 transformed with either empty vector (EV) control or mCherry stress biosensor (BBa_K639003) vector are viable and can grow in waste conditioned media. Therefore waste conditioned media is an appropriate and novel experimental media with which to characterise biobricks within a mixed waste/landfill context. These data are also characterisation of an existing biobrick (BBa_K639003)
Growth and induction assays of our Biobricks
Growth and induction assays of our project biobricks. Several of our constructs contain sfGFP within an operon and therefore fluorescence can be utilised to determine if expression is being induced by either addition of Arabinose or Xylose as appropriate to the construct.
Ideally you would normalise the fluorescence with OD600 to give fluorescence/cell in order to get the actual difference in induction expression. A two-tailed t-test shows that p = 0.4308 > 0.05, therefore there is no statistical difference between induced and uninduced.
Empty Vector Control
Growth assay characterisation of existing biobricks
Stress biosensor characterisation (BBa_K639003)
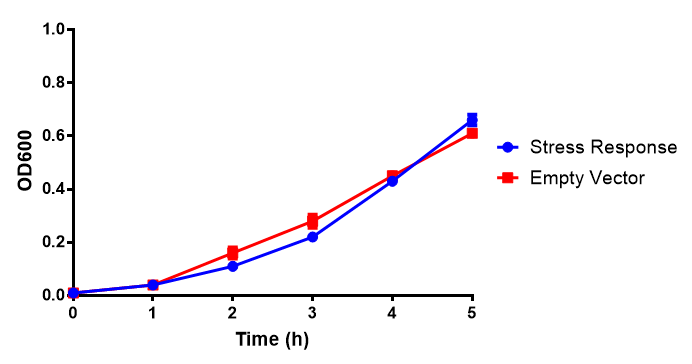
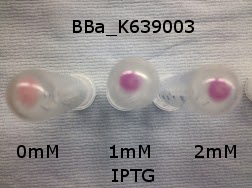
Originally we intended on using [http://parts.igem.org/Part:BBa_K639003 BBa_K639003] to detect whether our cells were stressed when grown with an array of potentially toxic plastics and degradation products. However, as the data below shows, the promoter is leaky and expresses mCherry in a non stressed state. As an alternative we utilised the stress sensor as a marker for our chassis E. coli (MG1655) for an array of qualitative and quantitative waste growth and toxicity assays.
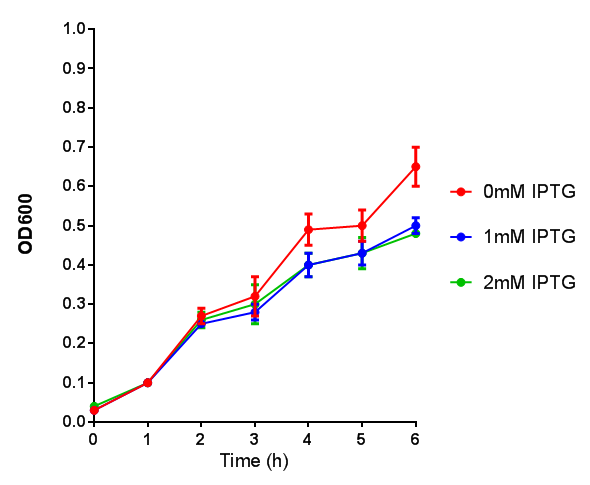
Note: The stress sensor induces mCherry production through a mechanism involving the ppGpp stress response. Induction with IPTG bypassess this mechanism through an inhibition of LacI, resulting in mCherry expression.
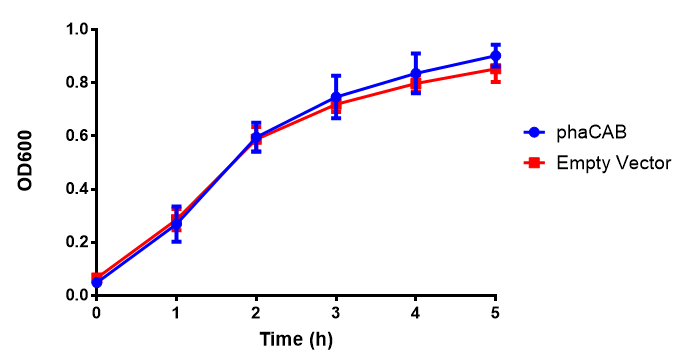
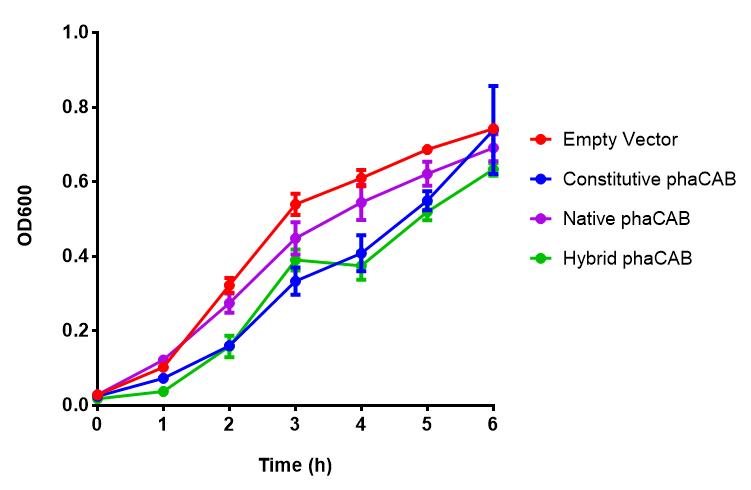
phaCAB biobrick characterisation
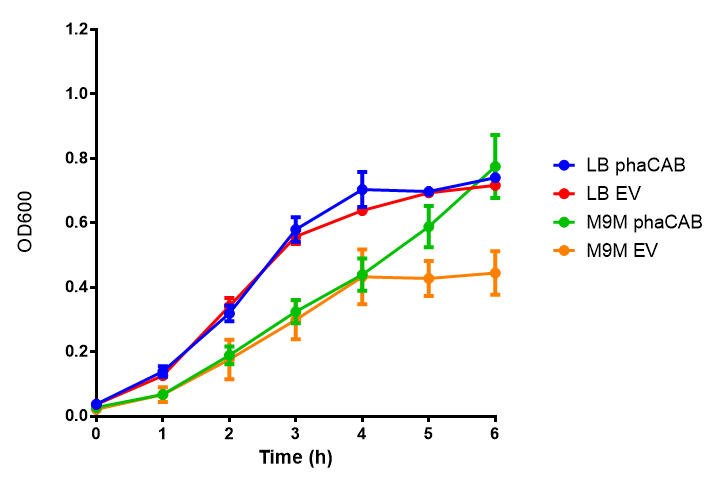
LB
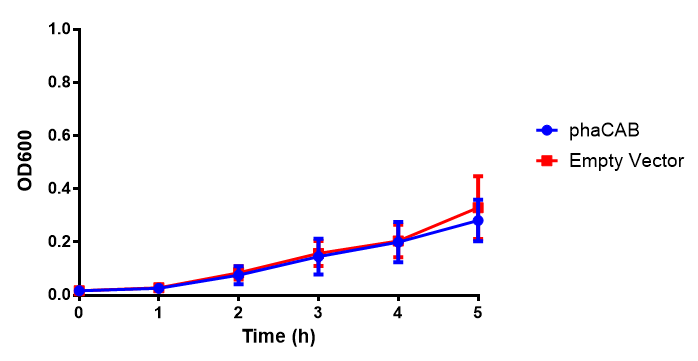
M9 Minimal
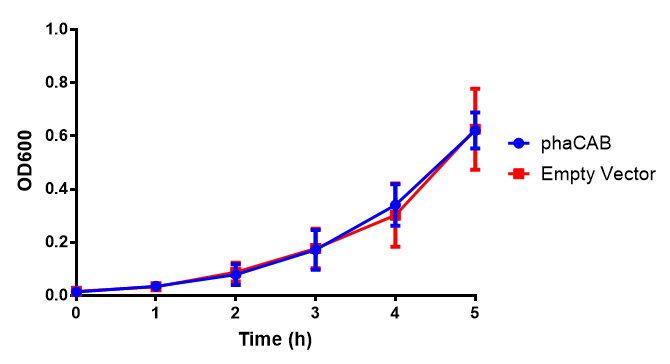
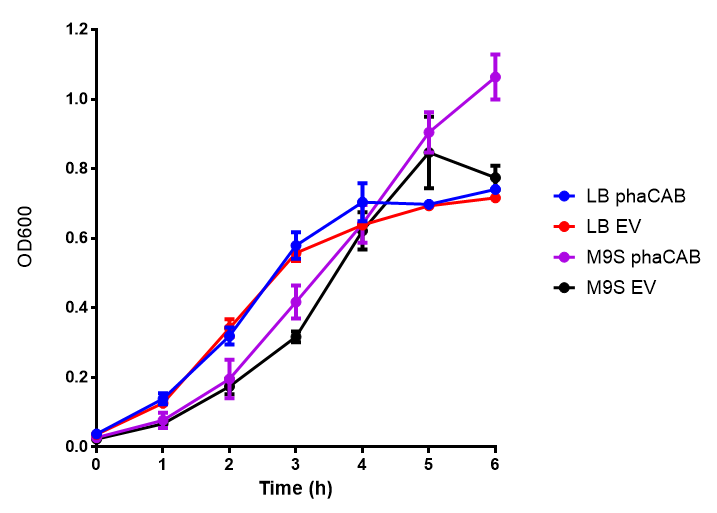
M9 Supplemented
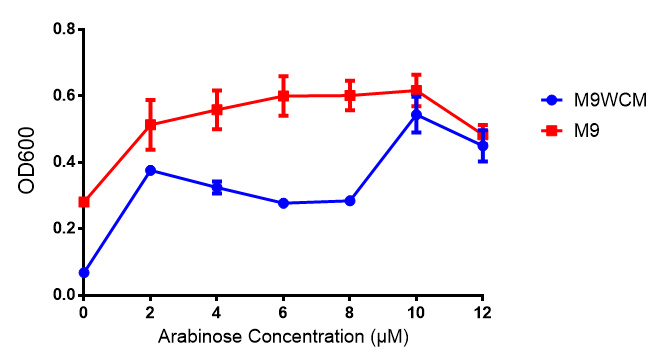
pBAD characterisation
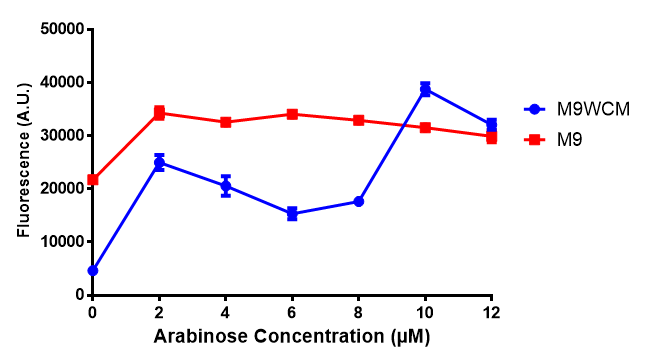
The null hypothesis, arabinose concentration does not influence growth of MG1655 in M9 minimal (M9M) media and M9 minimal waste conditioned media (M9MWCM) was tested using a two-tailed t-test. This must be rejected because p < 0.0001, thus arabinose concentration has an influence on growth in M9M and M9MWCM.
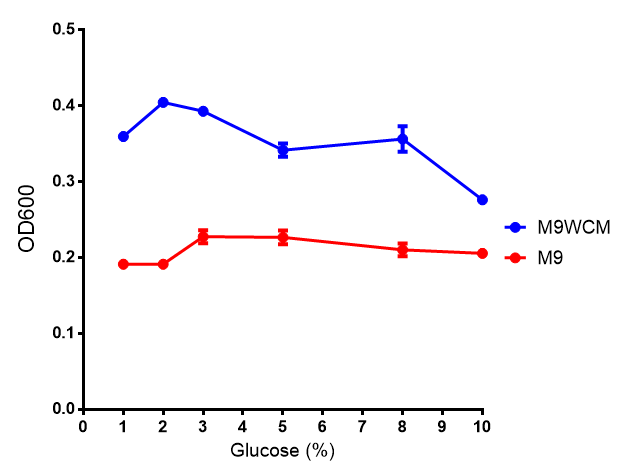
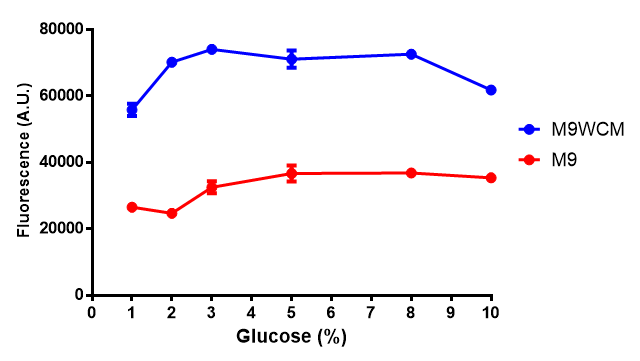
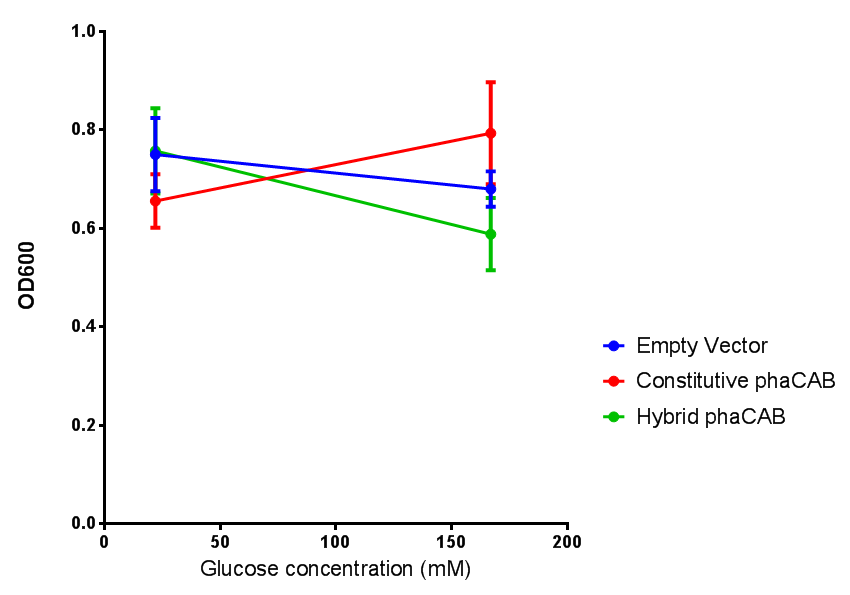
The null hypothesis, glucose concentration does not influence growth of MG1655 in M9 minimal (M9M) media and M9 minimal waste conditioned media (M9MWCM) was tested using a two-tailed t-test. This must be rejected because p < 0.0217, thus glucose concentration has an influence on growth in M9M and M9MWCM.
Glucose
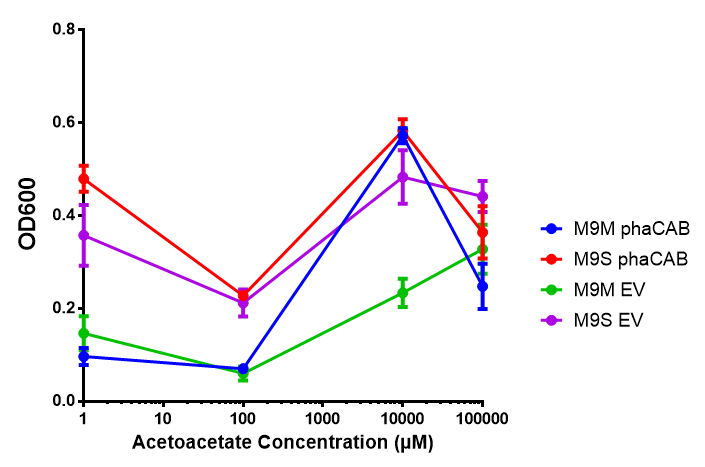
ANOVA analysis shows that the null hypothesis that there is no significant difference between M9M and LB in empty vector and phaCAB is true (F 3,24 = 0.8451, p < 0.4827). In addition to this the null hypothesis that there is no significant difference between M9S and LB similarly must be rejected as p > 0.05 (F 3,24 = 0.9802, p < 0.06009)
Plastic Toxicity Assays
L-lactic Acid
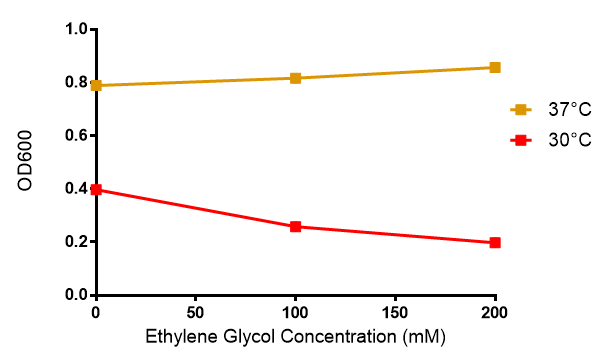
Ethylene glycol
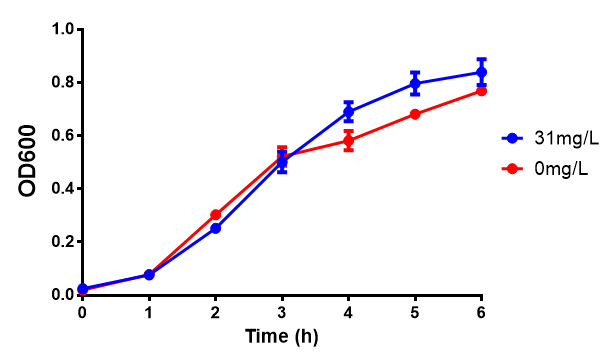
Reduced growth at 30oC likely due to decreased efficiency of MG1655 ethylene glycol break down enzymes. These enzymes (see UC Davis 2012) are endogenously expressed and detoxify Ethylene Glycol.
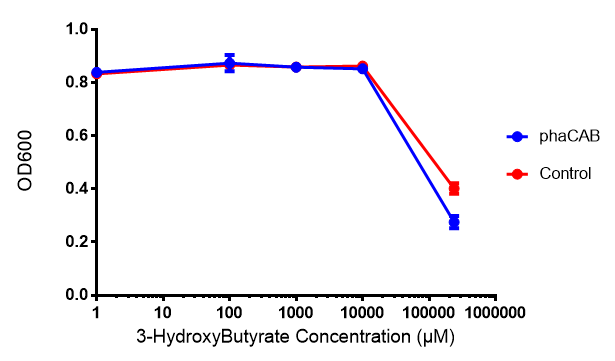
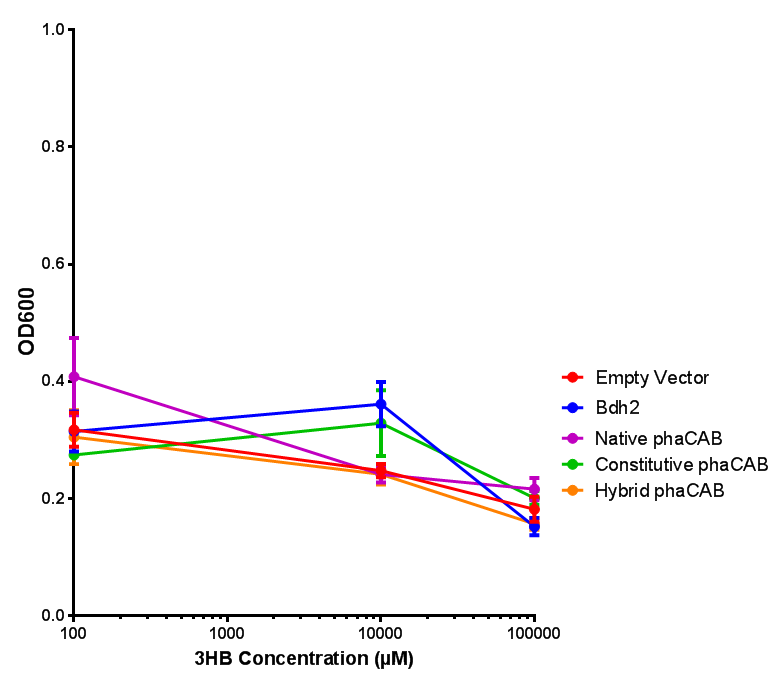
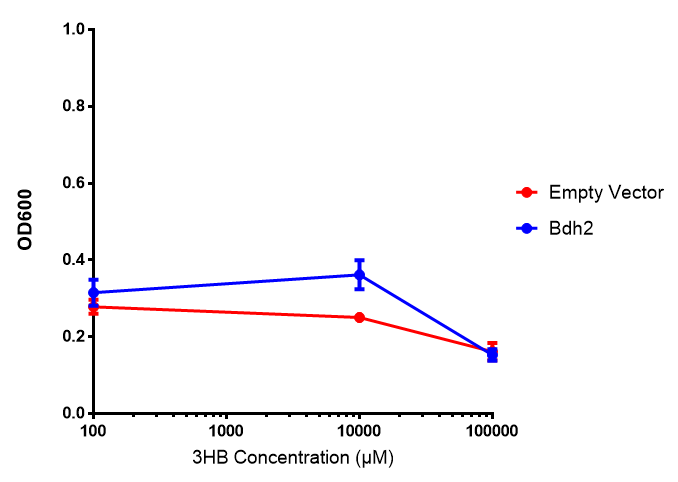
3-hydroxybutyrate (3HB)
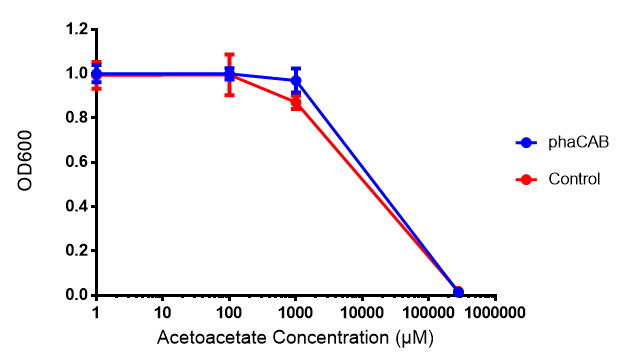
Acetoacetate
Poly(3-hydroxybutyrate) P(3HB)
Poly(lactic acid) (PLA)
Sole carbon source
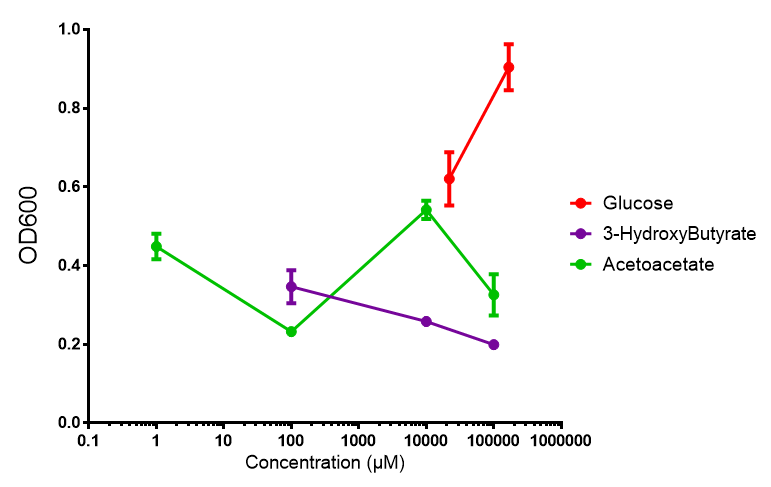
3HB
Acetoacetate
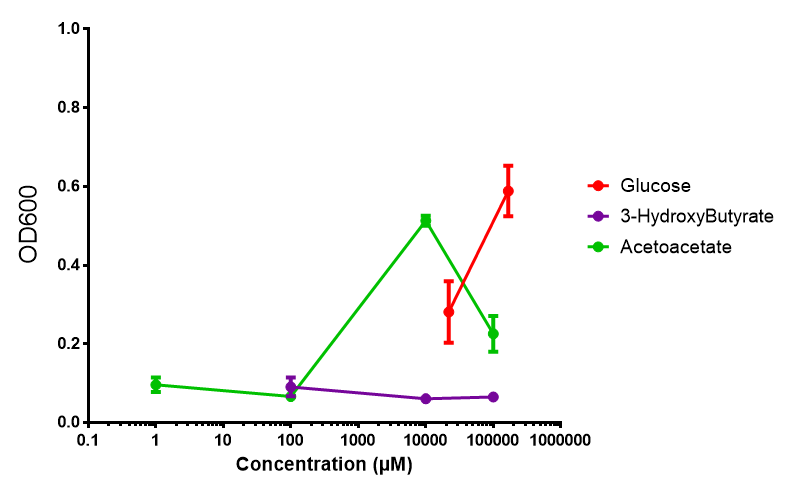
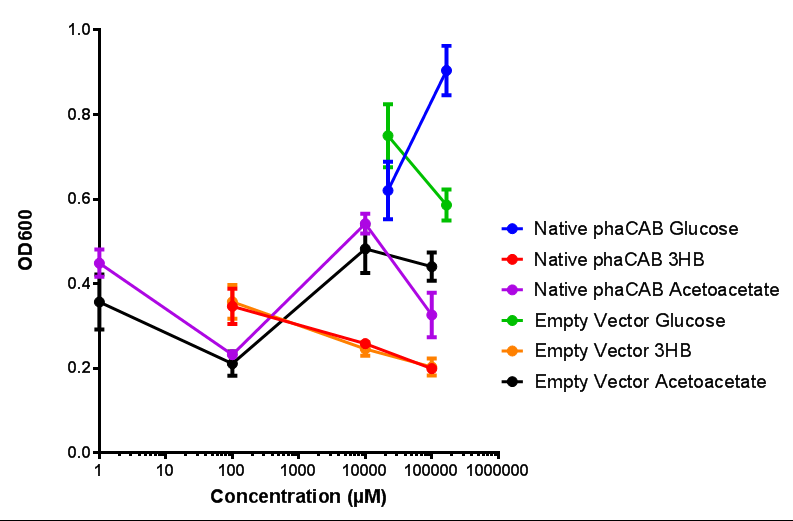
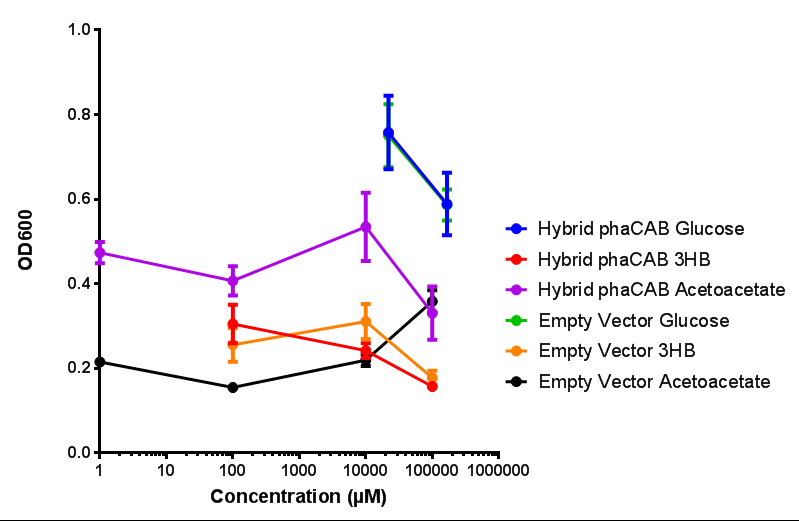
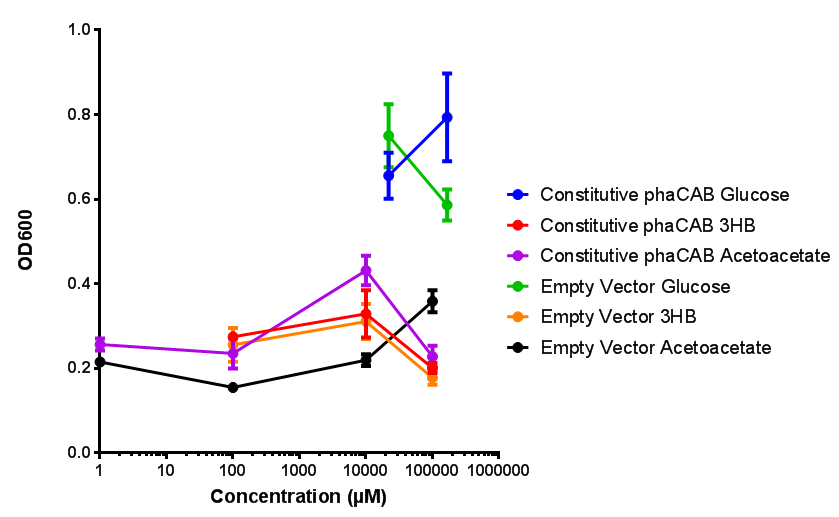
Compiled sole carbon growth
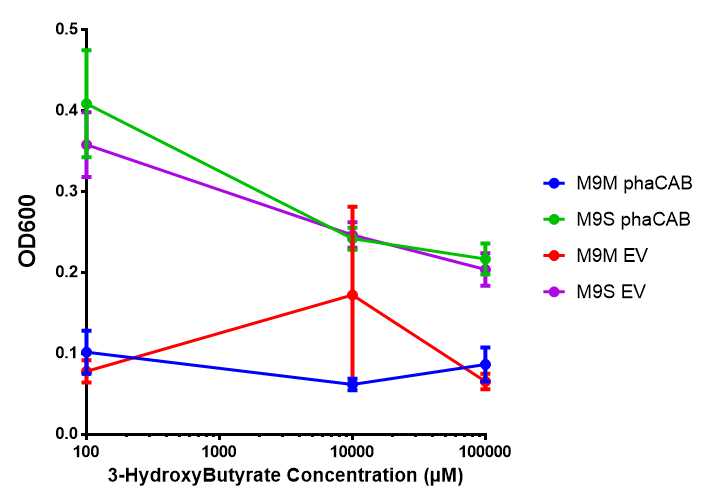
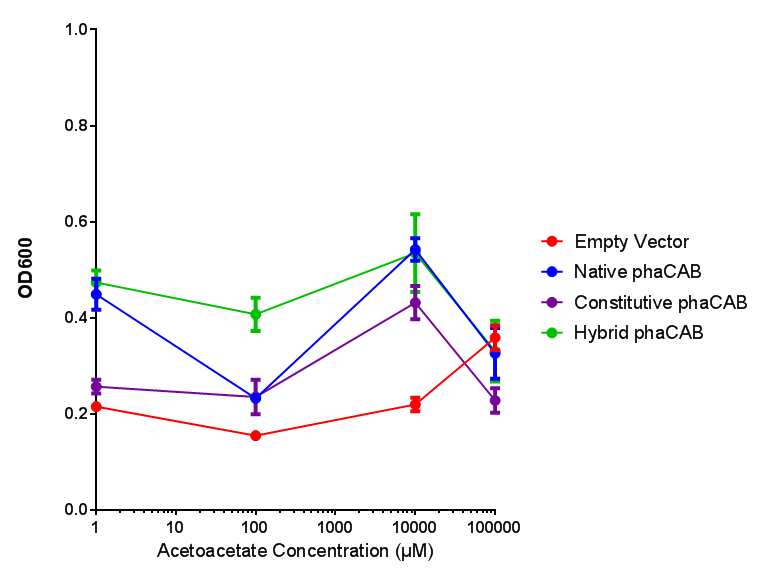
phaCAB on M9 minimal and supplemented media
phaCAB with novel promoter
 "
"