| Abstract
There are many applications of the game theory in some aspects of our life. Each individual has two kinds of choices--to betray or stay silent, and the choice you make would determine your fate. To betray the other side, you may risk being revenged. While staying silent, companion's betrayal may hurt you deeply. As for our project, we work out a new way to imitate the game theory by constructing a community of two E. Coli bacteria. Here we use the growth rate of each species to represent its fate. The effect of one's silent or betrayal on the other species' fate is acted through intercellular signal molecules of two quorum sensing systems. Each signal molecule regulates the expression of toxic genes in the other species and reduces its growth rate. We characterize the consequence of each strategy by quantitatively measure the growth rates of each species in the community
Background
Project Details
our design
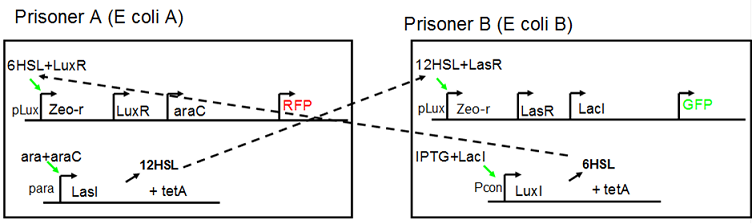
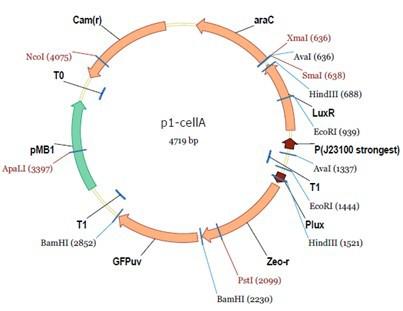
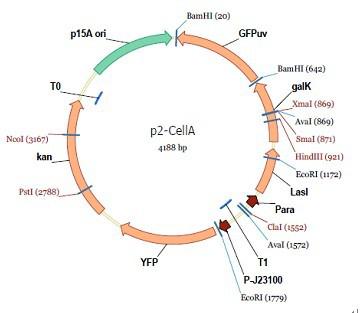 Cell A:
Constant expressions:
LuxR(with C6HSL, activate pLux, express Zeo-r): promoter J23100 and rbs B0034
araC(with arabinose, activate LasI, tetA): promoter J23100 and rbs B0034
mCherry (red fluorescent protein labeled cell A): promoter pCon, rbs
Adjustable expressions:
Zeo-r, adjusted by C6HSL, produced by LuxI from cell B, depends on cell B density and IPTG concentration
LasI: adjusted by arabinose, LasI produces C12HSL, control cell B
tetA: adjusted by arabinose
Cell A:
Constant expressions:
LuxR(with C6HSL, activate pLux, express Zeo-r): promoter J23100 and rbs B0034
araC(with arabinose, activate LasI, tetA): promoter J23100 and rbs B0034
mCherry (red fluorescent protein labeled cell A): promoter pCon, rbs
Adjustable expressions:
Zeo-r, adjusted by C6HSL, produced by LuxI from cell B, depends on cell B density and IPTG concentration
LasI: adjusted by arabinose, LasI produces C12HSL, control cell B
tetA: adjusted by arabinose
Arabinose, Zeocin, Nickel concentration are controlled by you
C12SHL concentration: controlled by cell A concentration and arabinose
Nickel either kill cell or slow cell growth: tetA increase sensitivity of cell B to Nickel:
zeocin either kill cell or slow cell growth:Zeo-r decreases sensitivity of cell B to zeocin
-asv: amino acid sequence that increase response time of LuxI, tetA and Zeo-r to regulation
pMB1 and p15A: control plasmid replication in E coli
Kan and Cam(r): with antibiotics Kanamycin and chloramphenicol, prevent lost of plasmids
Experiment process
Results
project results
modeling results
Biobricks checking
process
results
We chose four biobricks to detect after search . They are xylose、KNO3、glucose and 12HSL.</p>
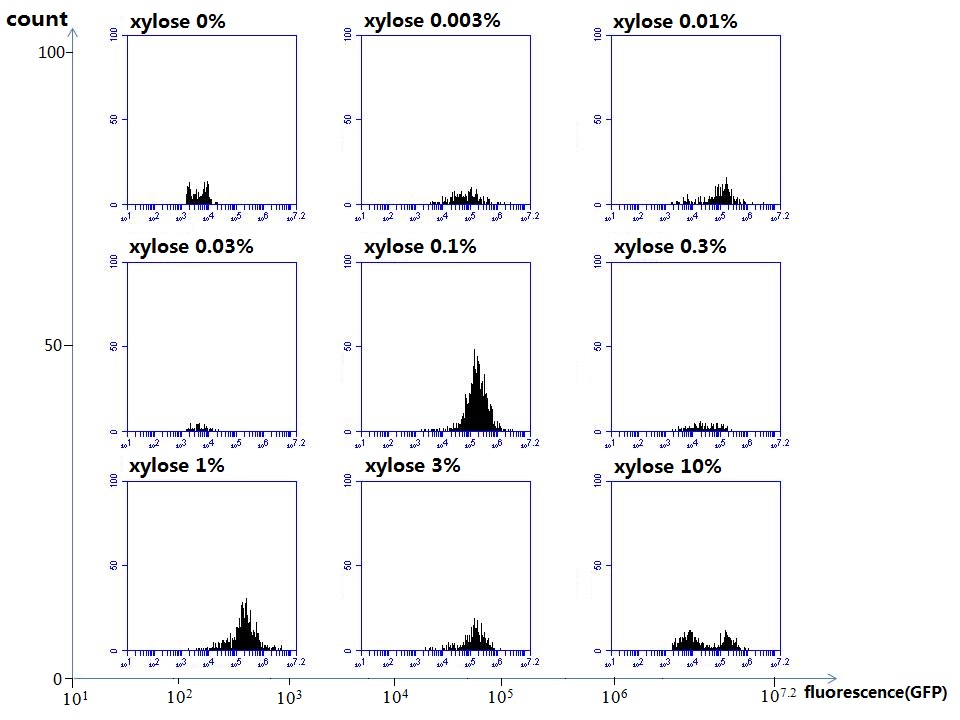
xylose
 KNO3
KNO3
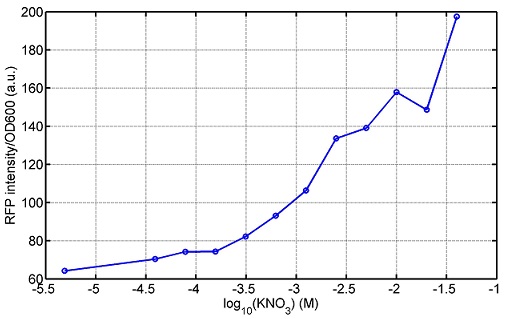
Compared with the results of the efficiency of the xylose inducible promoter detected by the HKT, our consequence differed from them in some aspects, or better. We had used the FCM to detect it. There was an obvious tendency that the efficiency of the promoter increased as the xylose concentration gathered up. What was amazing was that the fluorescence we got was 10 times brighter than the result the HKT got, which appeared to be great.
But when the concentration of the xylose was 0.03% and 0.3%, we got an unexpected result that the expression of the GFP fell after rise. What’s more. While the concentration of the xylose was 10%, the expression of the GFP decreased, which corresponded with the result of the HKT.
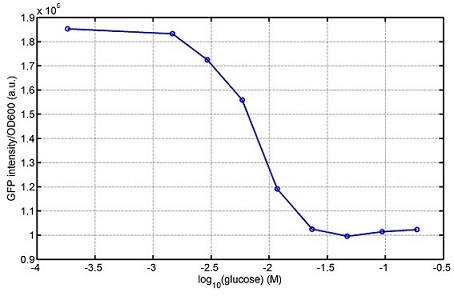 lalala
lalala
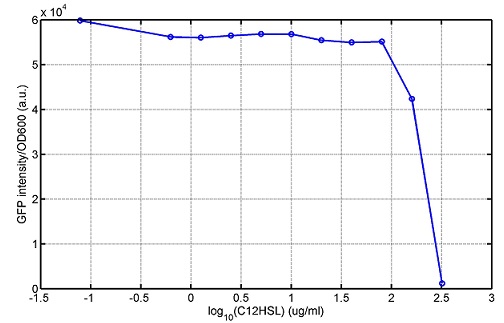
|