Team:Paris Bettencourt/Project/Sabotage
From 2013.igem.org

Background
One of the main concerns about tuberculosis today is the emergence of antibiotic resistant strain
Results
- Construction and characterization of phagemids coding for small RNA targeting antibiotic resistance proteins
- successful conversion of antibiotic resistant population of E. coli to a sensitive state
BioBricks
Aims
Our objective is to suppress the antibiotic resistance of a given bacterial population making it sensitive to treatments with antibiotics.
Skip to Design
Skip to Results
Skip to Perspectives
Introduction
As resistance against antibiotics are growing and pharmaceutical pipelines are drying up, we decided to investigate a new strategy based on specific silencing of the genes responsible for resistance through bioengineered stealth bacteriophages. The silencing of the genes is obtained with a simple and modular system of tailormade small RNAs and the spreading of this construct in a bacterial population is based on an autonomous phagemid/helper phage system. We demonstrated the validity of this trojan horse strategy by converting back to a sensitive state populations of bacteria initially resistant to antibiotics like chloramphenicol or kanamycin.
 Figure 1. The Trojan Horse Strategy : Infecting the cells with a phage bearing a sRNA silencing the expression of antibiotic resistant genes converting it back to a sensitive state.
Figure 1. The Trojan Horse Strategy : Infecting the cells with a phage bearing a sRNA silencing the expression of antibiotic resistant genes converting it back to a sensitive state.
Design
Design a silencing device
Most resistance against antibiotics come from specific proteins that bacteria can produce such as efflux pump that extract the antibiotics from the cell or enzymes that quickly metabolize the antibiotic molecules to make them inactive. We choose a straight forward strategy to force bacteria to stop producing those proteins by targeting the mRNA coding for them with tailormade small RNAs. In order to design our silencing device (small RNA) we used the protocol described in Na et al 2013 and inserted a 24bp sequence complementary to the RBS area of the target genes inside the scaffold sequence that allow the stabilization of the hybridization through recruitment of the protein Hfq. We designed three sRNA respectively targeting Kanamycine resistance gene located on a pCOLA Duet Vector, Chloramphenicol resistance on pACYC Duet Vector and lac Z on E. coli's chromosome.

Figure 2. Structure of the Chloramphenicol resistance silencing module
Design a genetic element that spread in a population
In nature, non-lytic filamentous bacteriophages are quite good at spreading genetic elements in bacterial populations and we thus immediately thought about them as vectors of choice. However being infected by a phage represent a huge burden for an individual bacteria which we thought would be detrimental for our construct to be able to maintain itself long enough in a population for this system to be clinically relevant.
Therefore we choose to use a phagemid/helper system which is composed of two mobile interacting elements. The « heavy » elements of this system is an M13 phage which genome contains an altered packaging signal thus reducing the probability of packaging in a protein capside and escaping from the cell. Its role is to produce capside proteins which will in fact be used by a « light » element called phagemid which is a normal plasmid harboring a packaging signal.
As a result a cell containing both elements will produce and secrete a lot of phagemid encapsulated plus some helper phages from time to time. We expect such a system to infect with light elements the majority of cells in a population and to be able to spread and maintain itself thanks to a small number of coinfected cells harboring both light (phagemid) and heavy (helper phage) elements transforming them into phage-producing cells.
Our silencing device is loaded on the light element in order to spread it efficiently. As the post-transcriptional regulation we are using only rely on RNA and doesn’t require synthesis of any protein, its cost for the cell will be very low. Moreover producing protective proteins against antibiotics is costly for the cell and lower its fitness in an antibiotic free environment, we thus expect our silencing device to be a temporary relief for the infected cell which should avoid early counterselection dynamics
Design of a sequential killing strategy
To perform our proof of concept experiments, our targeted genes were different antibiotic resistance genes on commercial plasmids that were transformed in MG1655 cells. This model of plasmid bearing resistance was chosen because it was more close to real life situation. Indeed resitance genes are mostly located on plasmids.
The experimental set up consisted of a sequential strategy ; a phage infection followed by a treatment with antibiotics. The phage infection leads to the silencing of the antibiotic-resistance genes. After the silencing is complete, antibiotics are used to kill the cells to prove the efficiency of the silencing and the possibility of using this strategy as a therapy.
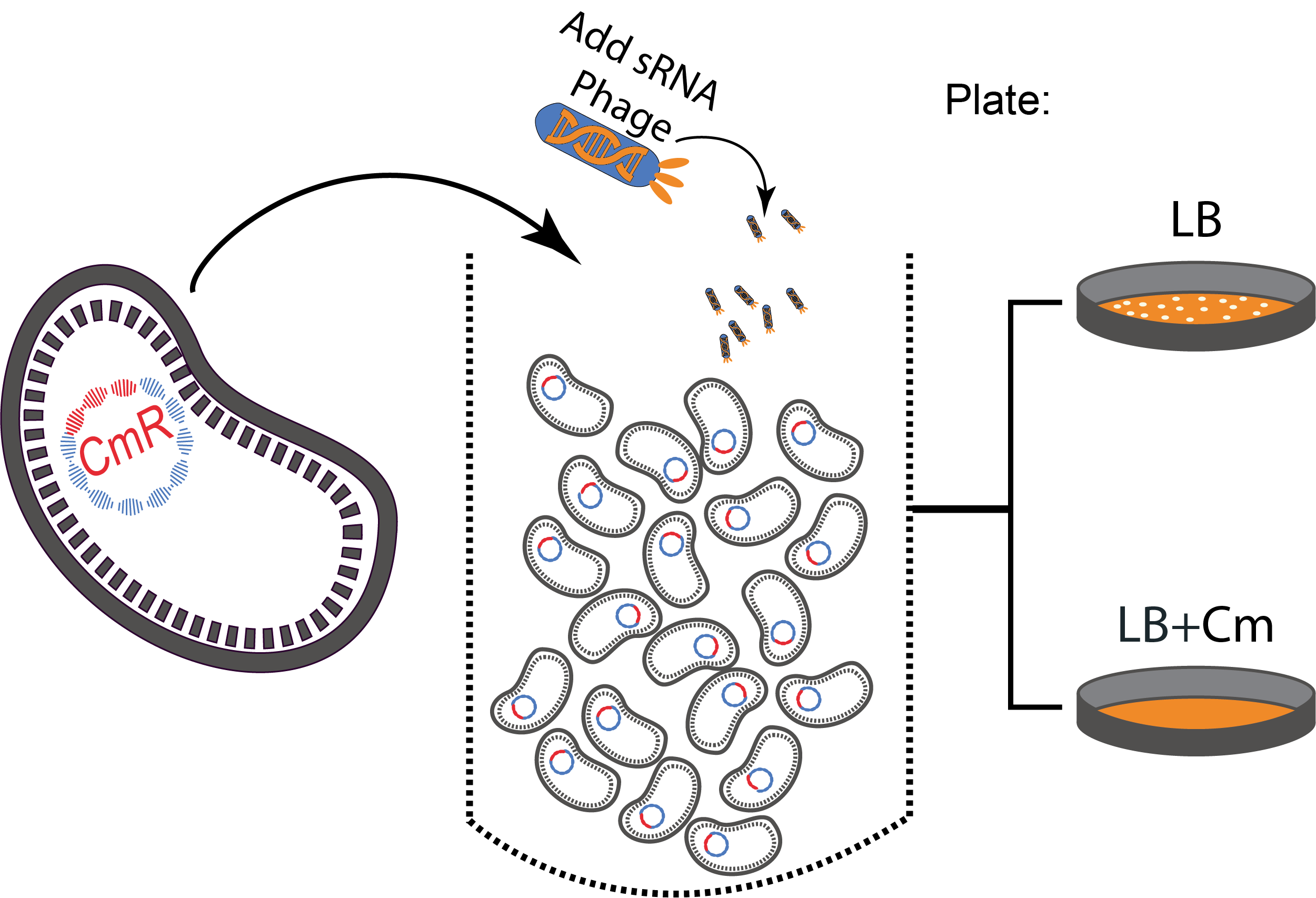 Figure 3. A sequential killing strategy :
Figure 3. A sequential killing strategy : Characterization of the phagemid system
We received the phagemid Helper system used in this project from Monica Ortiz. it consists of a helper plasmid, M13K07 bearing Kanamycine resistance and a phagemid, Litmus-28 bearing GFP and a Ampiciline resistance. Using those different markers, we characterized the system by infecting the cells with a ratio of phage of 1 to 100 and plating appropriate dilutions on different medias (LB,Kan, Amp, Kan/Amp). We show that the phagemid is spread a thousand fold more than the helper plasmid. There are as many cells infected with the helper as cells infected by both helper and phagemid, that is to say, phages producing cells. THose results show the validity of using this system to spread a genetic element in a bacterial population. Most of the cells bear the desired genetic element and the spread is carried on by a minor quantity of cells producing the vector.
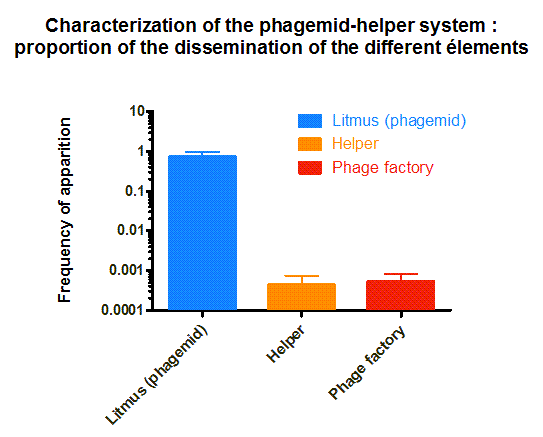 Figure 4. Characterization of the phagemid helper system
Figure 4. Characterization of the phagemid helper system
The critical role of using a low cost element
 Figure 5. Characterization of the phagemid helper system
Figure 5. Characterization of the phagemid helper system
Making a Chloramphenicol resistant E. Coli population sensitive to Chloramphenicol
We used the a phagemid bearing the sRNA anti-Cm we designed to infect a population of chloramphenicol resistant cells. By applying the sequential killing strategy described in Figure 3, we efficiently killed 99,1% of the bacterial population (chloramphenicol concentration 500ug/ul). We tested this strategy with different concentrations of chloramphenicol. We show that the rise of chloramphenicol concentration directly lead to drastic reduction of the survival of the infected cells.
In order to show the general application of our system, we then applied the same strategy on MG1665 transformed with pCOLA Duet. Those kanamycine resistant cells were infected with phages bearing our sRNA anti Kan. 99,87% of the population was killed (standard deviation 0,23 %) at 500mg/mL of kanamycine.
 Figure 6. Killing Chloramphenicol resistant E. Coli population with Chloramphenicol
Figure 6. Killing Chloramphenicol resistant E. Coli population with Chloramphenicol
Charaterization of the origin of resistance
One of the main limit of our system could be the development of resistances. Indeed, there are at least two ways to become resistant to our system: by avoiding infection or by developing a resistance to the silencing. In order to get a better idea of what is the main cause of resistance we screened the survivor colonies for GFP expression. GFP positive colonies correspond to infected cells with malfunctioning silencing. We show that 70% of resistance is caused by a resistance towards the sRNA. This resistance could be due to a mismatch between the sRNA sequence and the target sequence resulting from a single point mutation.

Figure 7. Origin of resistance to the sequential killing
Conclusion and perspectives
Several sRNA on the same phagemid : Although our silencing small RNAs were assembled in phagemids with Gibson assembly, the Biobricks cutting sites included in our BBG adapter method provide full modularity. In our case a direct application is the assembly of multiple silencing modules (promotor+sRNA+terminator) on a single phagemid vector. As the main clinical problem with bacteria is multi or even pan-resistance this modularity provide an interesting strategy to explore as it allows simultaneous downregulation of multiple target genes.
Building a library of sRNA : As our silencing strategy rely on base pair complementarity, a main concern from the clinical point of view would be the fast emergence of resistances to silencing through single point mutations. Indeed even if the majority of mutations that would decrease the efficiency of the silencing would also lead to an impaired or non functionnal protective protein, some neutral mutations will still be made possible by the redundancy of the genetic code. To tackle this issue an interesting solution would be to build a library of small RNA containing single point mutation on those particular neutral positions and load it on our phagemids. By being one move ahead of evolution we expect that such an anticipating strategy would considerably slow down the emergence of resistance to silencing.
Possible applications : As most bacteria species have their own phages, our strategy could be adapted to a broad range of diseases. Tuberculosis of course is particularly concerned because the number of panresistance cases is exploding in hospitals but potential targets could also be Staphylococcus aureus infections or even dangerous strains of E. coli as suggested in our model.
Literature
Na D, Yoo SM, Chung H, Park H, Park JH, Lee SY. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nature Biotechnology(2013).
Monica E Ortiz and Drew Endy. Engineered cell-cell communication via DNA messaging. Journal of Biological Engineering (2012).
Seung Min Yoo, Dokyun Na and Sang Yup Lee. Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nature Protocols 8, 1694–1707 (2013).
Woodford N, Wareham DW; UK Antibacterial Antisense Study Group. Tackling antibiotic resistance: a dose of common antisense? The Journal of Antimicrobial Chemotherapy. 2009 Feb;63(2):225-9.
 "
"







 +33 1 44 41 25 22/25
+33 1 44 41 25 22/25


