Team:Edinburgh/Project/Results/Aggregation Results
From 2013.igem.org
Hristianita (Talk | contribs) |
Hristianita (Talk | contribs) m (moved Team:Edinburgh/Aggregation Results to Team:Edinburgh/Project/Results/Aggregation Results: Organising) |
||
| (4 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
<div class='content'> | <div class='content'> | ||
| - | <h3> Aggregation – metal induced biofilm formation in '' | + | <h3> Aggregation – metal induced biofilm formation in ''B. subtilis'' </h3> |
| - | As most used laboratory strain of '' | + | As the most used laboratory strain of ''Bacillus subtilis'' - the 168 - is not able to form biofilms, we have obtained a sample of native ''‘‘B. subtilis’’'' capable to do that. Figure 1. presents the difference in on-plate appearance of the two starins. |
[[File:AggregationResults1.png]] | [[File:AggregationResults1.png]] | ||
| - | '''Figure 1.''' Native strain of '' | + | '''Figure 1.''' Native strain of ''B. subtilis'' (left) formed a biofilm layer (top of the left plate) whereas ''B. subtilis'' 168 (right) is not capable to do that. |
| Line 19: | Line 19: | ||
'''Figure 2'''. SinR PCR product of an expected size - 333bp | '''Figure 2'''. SinR PCR product of an expected size - 333bp | ||
| - | In order to perform deletion of native SinI and SinR in '' | + | In order to perform deletion of native SinI and SinR in ''B. subtilis'' genome their flanking regions were amplified from gDNA (Figure 3) and used for GenBrick assembly together with metal repressible SinR. |
[[File:AggregationResults3png.png]] | [[File:AggregationResults3png.png]] | ||
Latest revision as of 10:38, 4 October 2013
Aggregation – metal induced biofilm formation in B. subtilis
As the most used laboratory strain of Bacillus subtilis - the 168 - is not able to form biofilms, we have obtained a sample of native ‘‘B. subtilis’’ capable to do that. Figure 1. presents the difference in on-plate appearance of the two starins.
Figure 1. Native strain of B. subtilis (left) formed a biofilm layer (top of the left plate) whereas B. subtilis 168 (right) is not capable to do that.
The first task to obtain a control over the biofilm formation pathway was to clone the SinR transcription factor – a master regulator of that process. This was performed (Figure 2) and the gene was deposited in the Registry of Standard Biological Parts as BBa_K1122000.
Figure 2. SinR PCR product of an expected size - 333bp
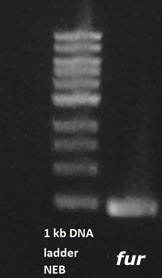
In order to perform deletion of native SinI and SinR in B. subtilis genome their flanking regions were amplified from gDNA (Figure 3) and used for GenBrick assembly together with metal repressible SinR.
Figure 3. SinR flanking region has a PCR product of an expected size.

| 
| | | | 
|
| This iGEM team has been funded by the MSD Scottish Life Sciences Fund. The opinions expressed by this iGEM team are those of the team members and do not necessarily represent those of MSD | |||||
 "
"