Team:Newcastle/Parts/HBsu-fp
From 2013.igem.org
| Line 162: | Line 162: | ||
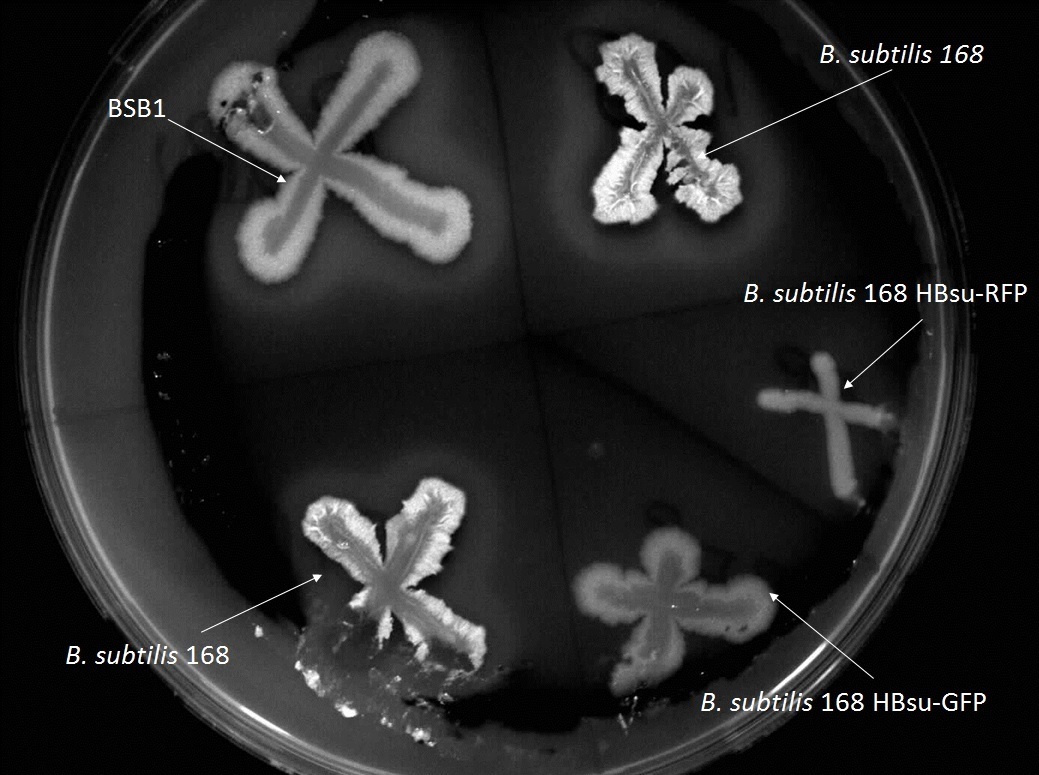
''Figure 1. Starch plate showing alpha amylase activity. The ''B. subtilis'' 168 HBsu-GFP and HBsu-RFP colonies have no zones of clearance indicating that our HBsu-sfGFP and HBsu-RFP have integrated into the B. subtilis chromosome via the ''amyE'' region stopping starch digestion.'' | ''Figure 1. Starch plate showing alpha amylase activity. The ''B. subtilis'' 168 HBsu-GFP and HBsu-RFP colonies have no zones of clearance indicating that our HBsu-sfGFP and HBsu-RFP have integrated into the B. subtilis chromosome via the ''amyE'' region stopping starch digestion.'' | ||
| + | <html> | ||
| + | <style type="text/css"> | ||
| + | #body img{ | ||
| + | margin-bottom:3px; | ||
| + | } | ||
| + | table{ | ||
| + | border:2px solid black; | ||
| + | margin-left:50px; | ||
| + | margin-bottom: 50px; | ||
| + | width: 650px; | ||
| + | } | ||
| + | .italic{ | ||
| + | font-style: italic; | ||
| + | } | ||
| + | |||
| + | </style> | ||
| + | <body> | ||
| + | <table border="0"> | ||
| + | <tr> | ||
| + | <td><img src="https://static.igem.org/mediawiki/2013/b/b5/BareCillus_0.1mM_IPTG_10mins.jpg" alt="Pulpit rock" width="304" height="228"><div class="italic">Plate 1: XL1B + 1:1 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml)</div></td> | ||
| + | <td><img src="https://static.igem.org/mediawiki/2013/8/8b/BareCillus_0.1mM_IPTG_30mins.jpg" alt="Pulpit rock" width="304" height="228"><div class="italic">Plate 2: XL1B + 1:3 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml)</div></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><img src="https://static.igem.org/mediawiki/2013/1/12/BareCillus_0.1mM_IPTG_60mins.jpg" alt="Pulpit rock" width="304" height="228"><div class="italic">Plate 3: XL1B + 1:5 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml)</div></td> | ||
| + | <td><img src="https://static.igem.org/mediawiki/2013/1/1a/BareCillus_0.1mM_IPTG_120mins.jpg" alt="Pulpit rock" width="304" height="228"><div class="italic">Plate 4: XL1B + 1:1 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml)</div></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td><img src="https://static.igem.org/mediawiki/2013/f/fb/BareCillus_0.1mM_IPTG_240mins.jpg" alt="Pulpit rock" width="304" height="228"><div class="italic">Plate 5: XL1B + 1:3 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml)</div></td> | ||
| + | <td></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </body> | ||
| + | </html> | ||
| + | |||
[[Image:BareCillus 0.1mM IPTG 10mins.jpg|300px]] | [[Image:BareCillus 0.1mM IPTG 10mins.jpg|300px]] | ||
[[Image:BareCillus 0.1mM IPTG 30mins.jpg|300px]] | [[Image:BareCillus 0.1mM IPTG 30mins.jpg|300px]] | ||
Revision as of 16:41, 21 September 2013

Contents |
HBsu-xFP BioBrick
Purpose and Justification
Bacillus subtilis contains HBsu, a non-specific DNA binding protein which is a homologue to eukaryotic histones. This protein is encoded by the gene hbs gene which is 263bp in length and involved not only in DNA binding but also SRP, DNA repair, HR, and pre-secretory protein translocation (Kouji, et al., 1999). It binds to DNA by forming homodimer.
We conjugated the hbs gene with red fluorescent protein (RFP/GFP) in order to fluorescently tag the bacterial chromosome. We produced BBa_K1185001 which codes for a HBsu-sfGFP conjugate and BBa_K1185002 coding for a HBsu-RFP conjugate. Each bacterium was transformed with only one or the other. The expression of these BioBricks was regulated through an IPTG induce promoter (Pspac).
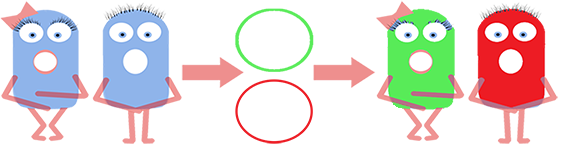
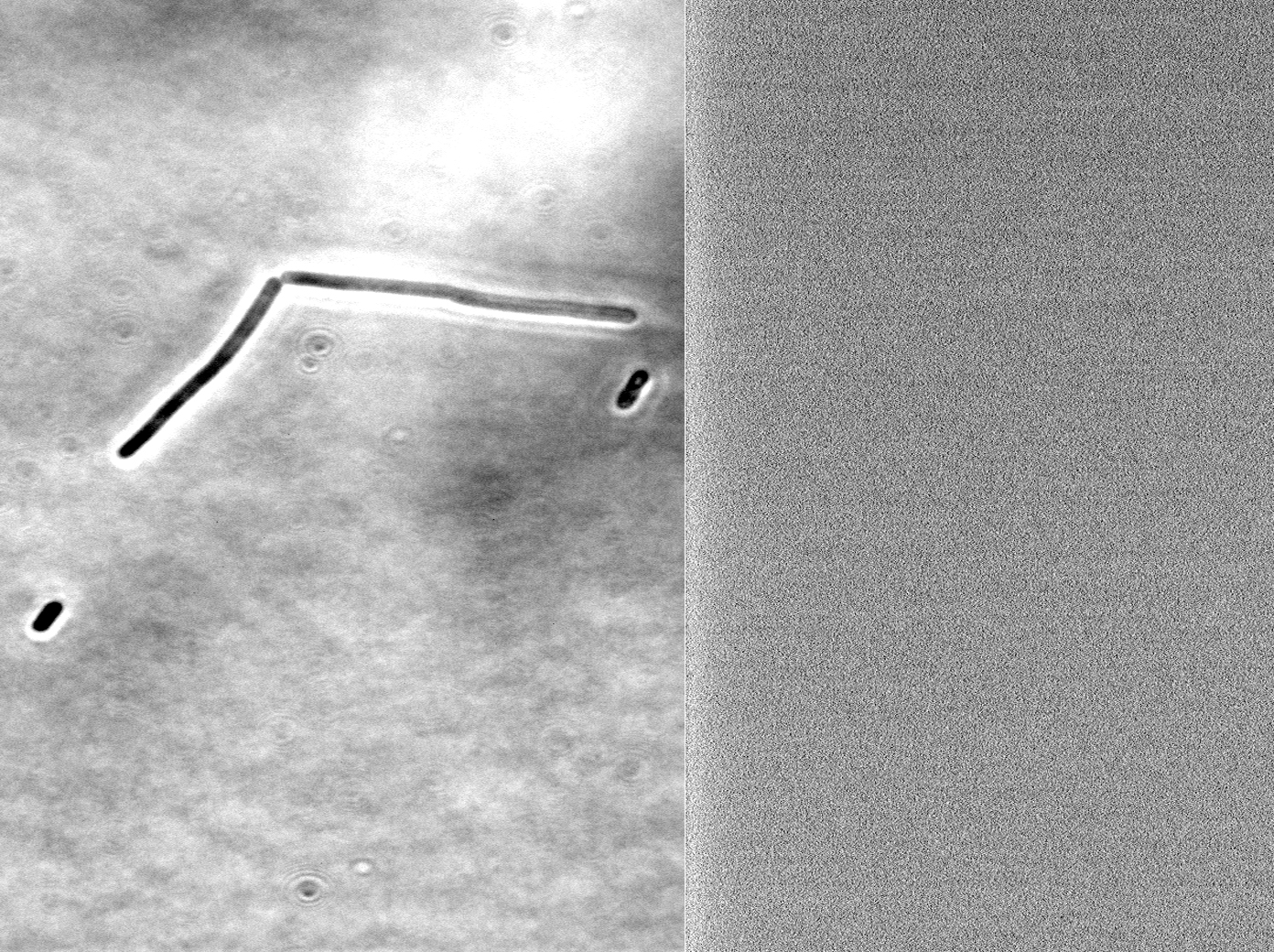
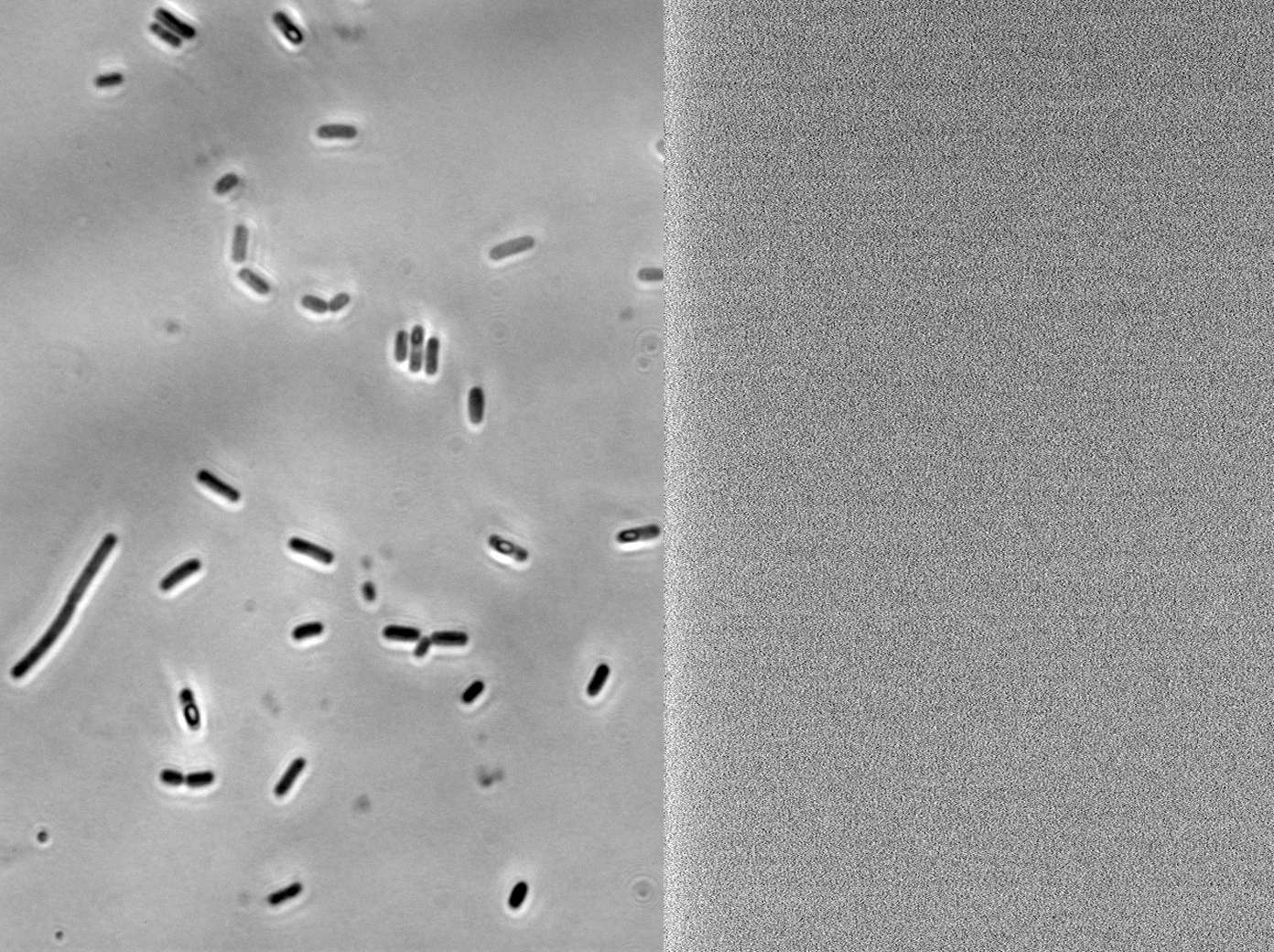
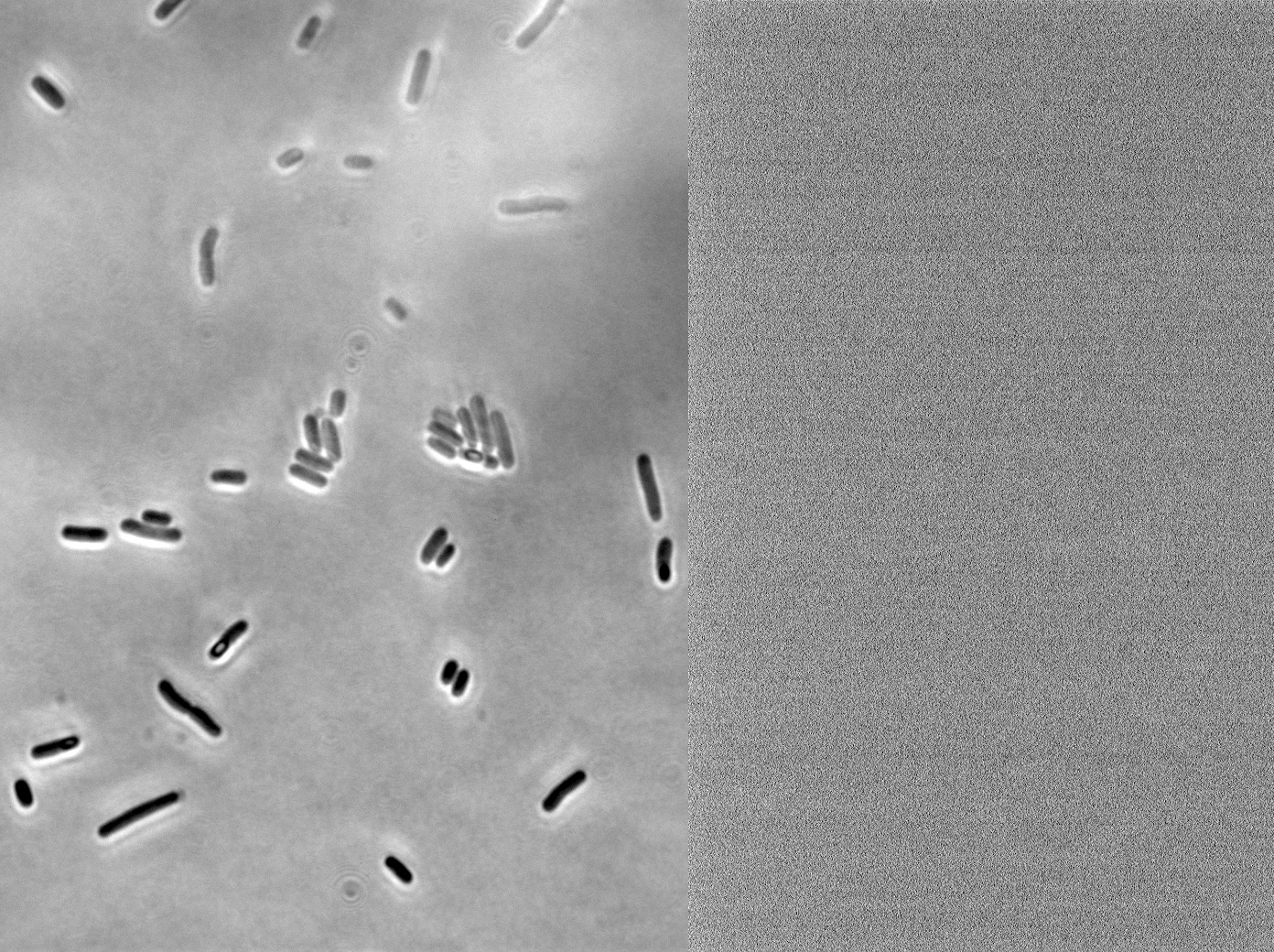
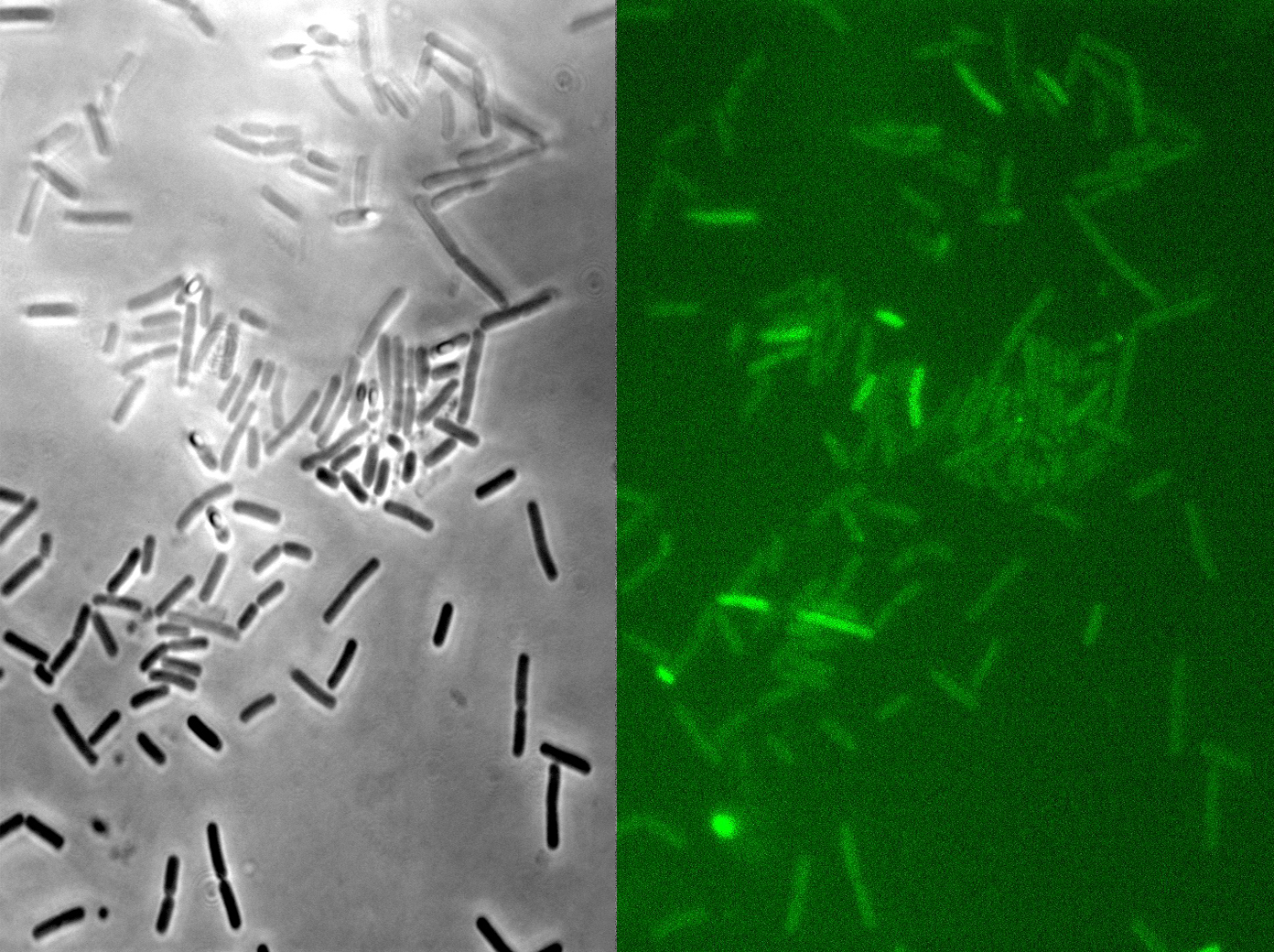
We fused one bacteria containing HBsu-RFP and another containing HBsu-sfGFP, and viewed the genomic shuffling via fluorescent microscopy. The formation of a colour mosaic allows us to confidently claim that recombination has occurred. Genomic shuffling has importance as it can be used to evolve and improve a cells phenotype.
Design
As part of our genome shuffling sub-project we required one set of bacteria coding for a protein that binded to DNA and fluoresced green and another set which coded for a protein that binded to DNA and fluoresced red. Our HBsu-xFP BioBricks code for HBsu: a B. subtilis histone-like protein which binds indiscriminately to DNA. So that we could observe the distribution of this protein we attached an amino acid linker sequence and a sfGFP/RFP (depending on the BioBrick) coding region to the 3' end of the HBsu coding region. Our first step was to plan our two DNA constructs using gene designer:
Figure 1. The BioBrick BBa_K1185001 consists of superfolded green fluorescent protein (sfGFP) ,BBa_E0040, conjugated to a HBsu protein.
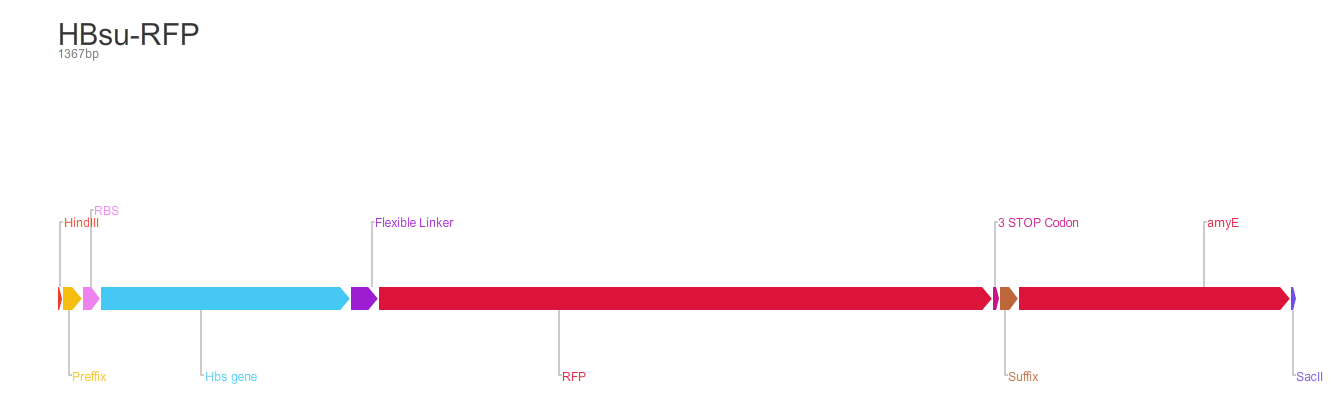
Figure 2. BioBrick BBa_K1185002 contains red fluorescent protein(RFP) ,BBa_E1010, attached to HBsu.
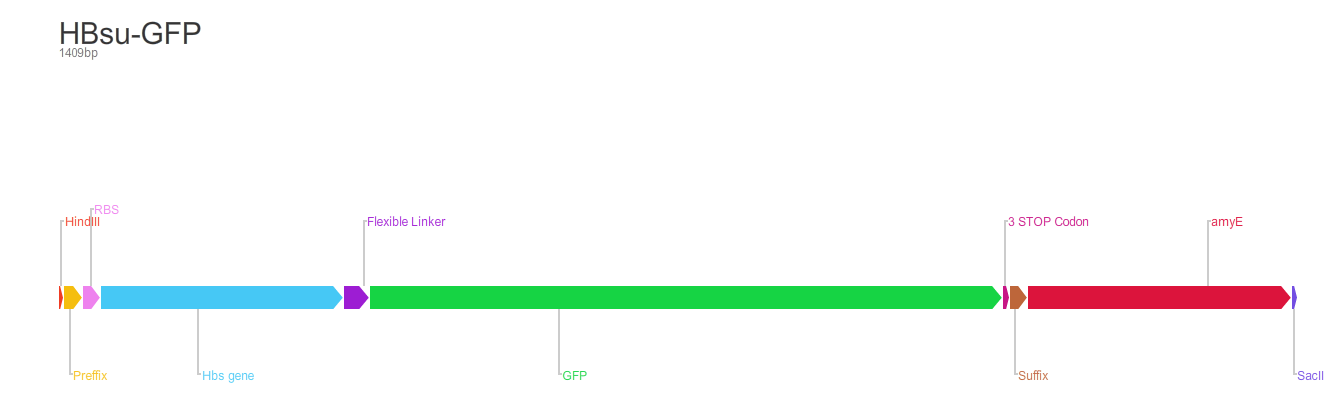
From the 5' end, the Design consisted of a HindIII restriction site, followed by a BioBick prefix sequence, a ribosome binding site(RBS), the HBsu gene, flexible linker sequence, sfGFP/RFP, ~300bp amyE homology region and a SacII restriction site. The HBsu, flexible linker sequence and sfGFP/RFP code for a fluorescently labelled HBsu. The ~300bp amyE homology region allows the single cross over and integration of plasmids into the genome. We did not include a promoter in our BioBrick as the pMutin4 plasmid we used as a vector contained a Pspac promoter 5' to its multiple cloning site. The HindIII and SacII restriction site on the 5' and 3'end of construct were included to allow integration into a pMutin4 multiple cloning site.
Modelling
Construction
The BioBrick BBa_K1185001 consists of superfolded green fluorescent protein (sfGFP) ,BBa_E0040, conjugated to a HBsu protein:
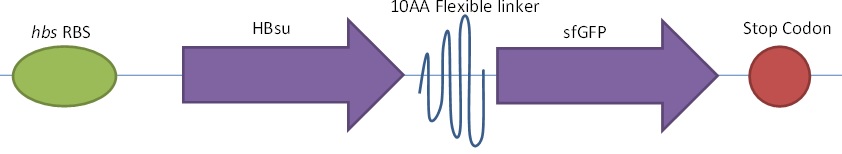
Whereas BioBrick BBa_K1185002 contains red fluorescent protein(RFP) ,BBa_E1010, attached to HBsu:
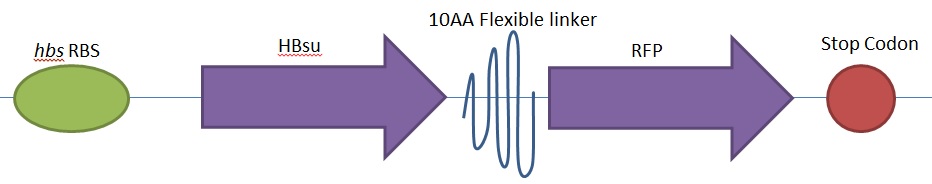
The constructs were synthesised by DNA synthesis company DNA 2.0. The components within the synthesised sequence were as follows: -
Parts:
- HinIII restriction site - for integration into a pMutin4 multiple cloning site.
- BioBrick prefix.
- Ribosome binding site(RBS) - naturally occuring HBsu RBS
- HBsu gene - binds to the B.subtilis DNA.
- amino acid linker sequence - joins HBsu and xFP.
- sfGFP/RFP - fluorescently marks the protein.
- Stop codon.
- BioBrick suffix.
- ~300bp amyE homology region - allows the single cross over and integration of plasmids into the B.subtilis genome.
- SacII restriction site - for integration into a pMutin4 multiple cloning site.
The synthesised construct was then inserted into the pMutin4 plasmid (Fig.3) to be used in our project. We then cloned out the BioBricks which are flanked by the prefix and suffix from the pMutin 4 plasmid and cloned it into the pSB1C3 plasmid in order to be sent off to the iGEM registry.
Cloning and Integration
pMutin4 - used as a B.subtilis integration vector
 |
 Figure 1. Plasmid map of Pmutin4 without insert |
 Figure 2. Plasmid map of our HBsu-sfGFP |
 Figure 3. Plasmid map of our HBsu-RFP |
Plasmids (pMutin4) containing the HBsu-RFP and HBsu-sfGFP BioBricks were returned from DNA 2.0 were amplified in E.coli. The E. coli competent cell preparation and E. coli transformation protocols were used to transform these plasmids into E. coli cells for cloning. Transformant cells were incubated in order to clone the transformed plasmids. Then the cloned plasmids were extracted and transformed into B. subtilis. Integration of the BioBrick from the vector plasmids to the host chromosomes was facilitated through homologous recombination with crossover events within the amyE homology region. The pMutin4 plasmid contains a ery+ resistance marker for B.subtilis and amp+ for E.coli which we used to select for transformed cells. pMutin4 also contains lacI, lacZ and a Pspac promoter which is an IPTG induced promoter which regulated the transcription of the BioBricks.
Figure 4. This is the final product of the homologous recombination.
pSB1C3 - for submission to the parts registry
In order to submit the HBsu-RFP and HBsu-sfGFP BioBricks to the iGEM repository they needed to be cloned out of pMutin4 and into psB1C3 plasmids. We cut our BioBricks and the linearised psB1C3 plasmid from iGEM using EcoRI and PstI. We then purified the two BioBricks using the gel extraction protocol.
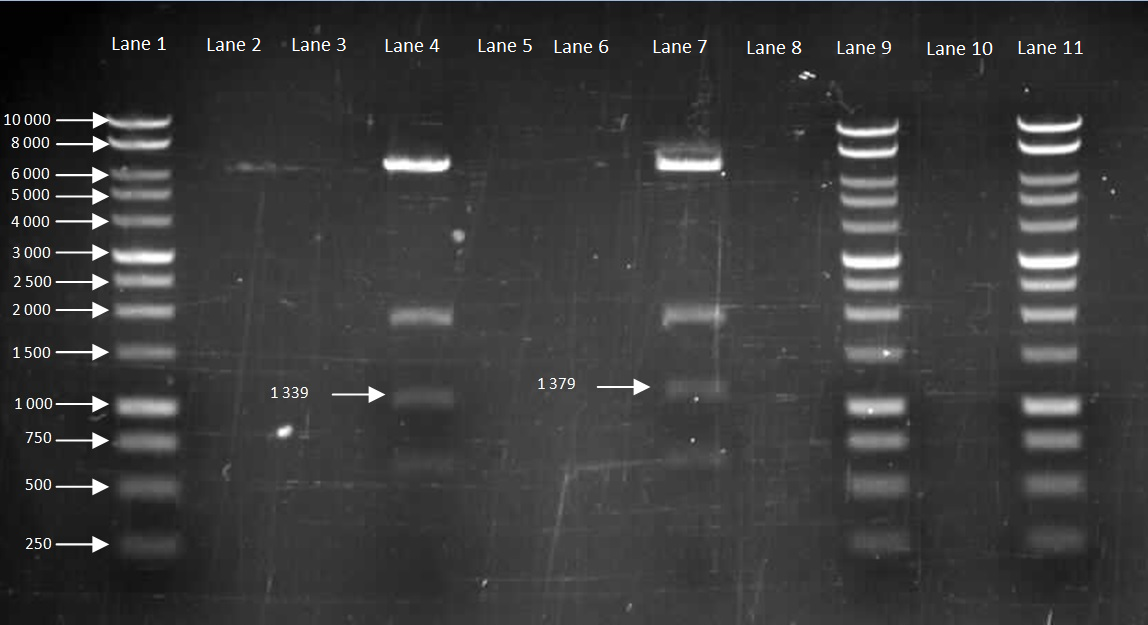
Figure 5. Gel to confirm size of HBsu-GFP and HBsu-RFP in pMutin4. Lane 1: 10kB DNA Ladder, Lane 2-3: Empty, Lane 4: HBsu-RFP, Lane 5-6: Empty, Lane 7: Hbsu-GFP, Lane 8:Empty, Lane 9: 10kB DNA Ladder, Lane 10: Empty and Lane 11: 10kB DNA Ladder. HBsu-RFP is labelled as '1339' bp in length and HBsu-sfGFP is labelled as '1379' bp in length.
 Figure 6. After we have confirmed the size, we ran a restriction digest and cut out the fragment of interest of HBsu-RFP and HBsu-sfGFP using the transilluminator and performed the gel extraction protocol. Lane 1: 10kB DNA Ladder, Lane 2-4: Empty, Lane 5: HBsu-RFP, Lane 6-7: Empty, Lane 8: 10kB DNA Ladder, Lane 9-11:Empty and Lane 12: Hbsu-GFP.
Figure 6. After we have confirmed the size, we ran a restriction digest and cut out the fragment of interest of HBsu-RFP and HBsu-sfGFP using the transilluminator and performed the gel extraction protocol. Lane 1: 10kB DNA Ladder, Lane 2-4: Empty, Lane 5: HBsu-RFP, Lane 6-7: Empty, Lane 8: 10kB DNA Ladder, Lane 9-11:Empty and Lane 12: Hbsu-GFP.
After we seperated our BioBricks we checked the nanodrop reading to find out the concentration of DNA which turned out to be for 20ng/ul HBsu-RFP and 17 ng/ul for HBsu-GFP). We were then able to ligate the BioBricks into the EcoR1 and Pst1 cut pSB1C3 backbone using T4 ligase. We transformed the pSB1C3 into E.coli XL1-B and innoculated the following plates:
 Plate 1: XL1B + 1:1 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 2: XL1B + 1:3 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 3: XL1B + 1:5 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 4: XL1B + 1:1 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 5: XL1B + 1:3 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 6: XL1B + 1:5 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 7: XL1B + No insert pSB1C3 on LB + Cm (5ug/ml) |
Plates 1 to 6 had colonies showing our cloning and transformation had worked. Plate 7 had no colonies as it was a negative control. |
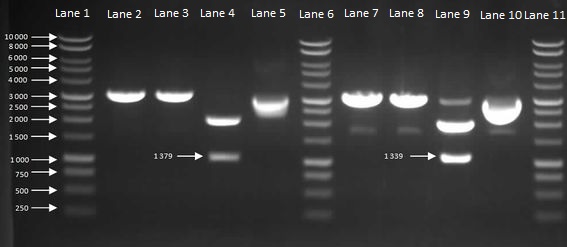
We then inoculated one colony from each sample in to LB and let it grow overnight in order to harvest the plasmids. With the amplified plasmids we could then check that it is the correct size and confirm that cloning had worked (Figure 7). The HBsu-GFP and HBsu-RFP insert was confirmed to be the correct size of 1379bp and 1339bp respectively.
Figure 7: Gel to Confirm Size of HBsu-GFP and HBsu-RFP insert size. Lane 1: 1kB Ladder, Lane 2: HBsu-GFP + PstI, Lane 3: Hbsu-GFP + EcoRI, Lane 4: HBsu-GFP + PstI and EcoRI, Lane 5: uncut product HBsu-sfGFP, Lane 6: 1kB Ladder, Lane 7: HBsu-RFP + Pst1, Lane 8: HBsu-RFP + EcoRI, Lane 9: HBsu-RFP + EcoRI and Pst1, Lane 10: uncut HBsu-RFP and Lane 11: 1kB Ladder.
As Figure 7 shows, the cloning had worked so we were able to send off our HBsu-sfGFP (BBa_K1185001) and HBsu-RFP (BBa_K1185002) BioBricks to the parts registry in the Massachusetts Institute of Technology.
 Figure 8. Plasmid map of BBa_K1185001 in pSB1C3.
Figure 8. Plasmid map of BBa_K1185001 in pSB1C3.
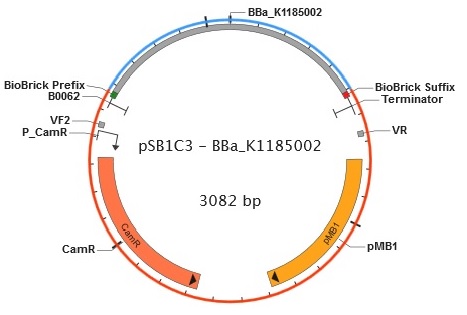 Figure 9. Plasmid map of BBa_K1185002 in pSB1C3.
Figure 9. Plasmid map of BBa_K1185002 in pSB1C3.
Testing and Characterisation
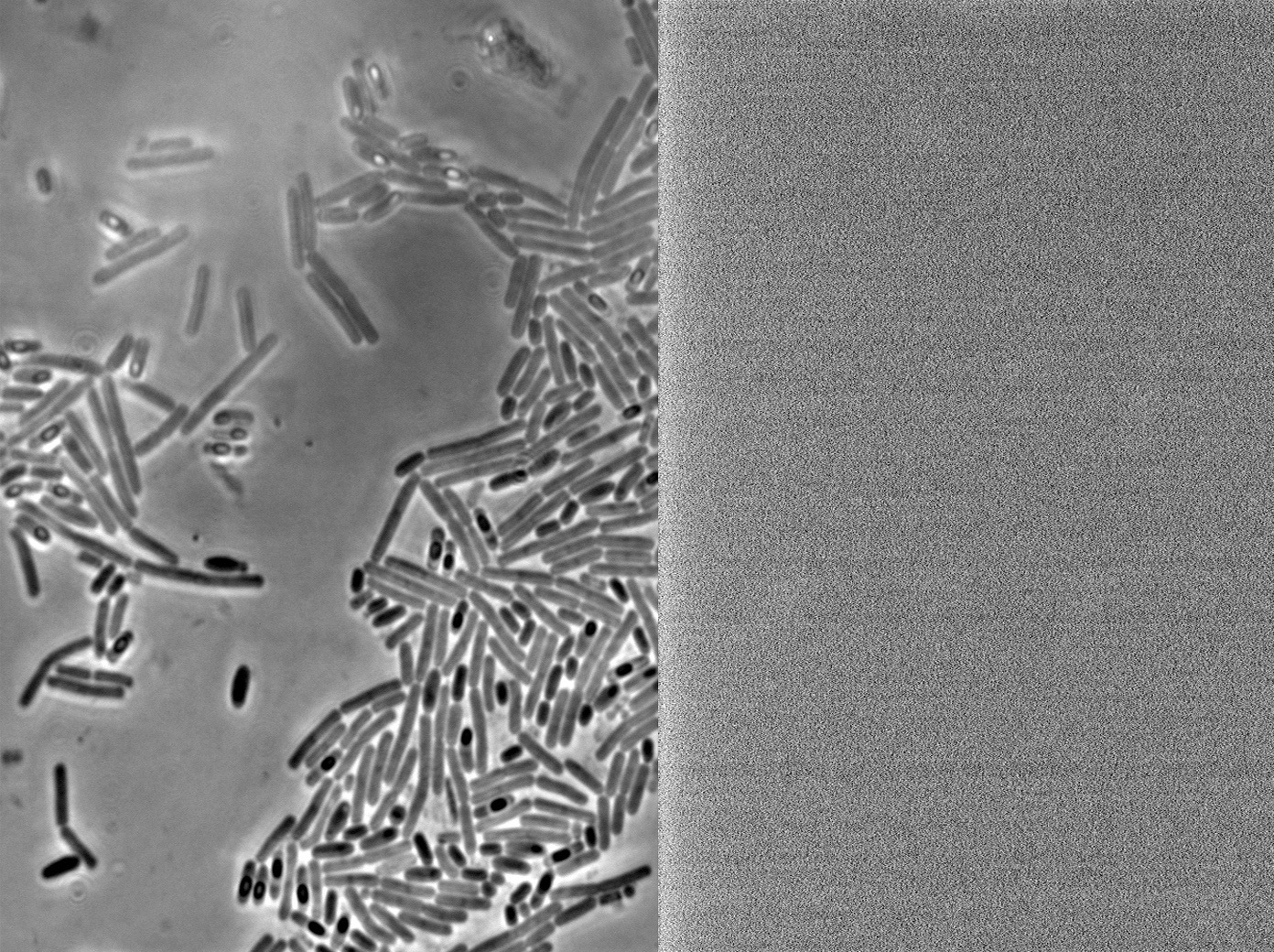
Figure 1. Starch plate showing alpha amylase activity. The B. subtilis 168 HBsu-GFP and HBsu-RFP colonies have no zones of clearance indicating that our HBsu-sfGFP and HBsu-RFP have integrated into the B. subtilis chromosome via the amyE region stopping starch digestion.
 Plate 1: XL1B + 1:1 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 2: XL1B + 1:3 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 3: XL1B + 1:5 HBsu-GFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 4: XL1B + 1:1 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml) |
 Plate 5: XL1B + 1:3 HBsu-RFP pSB1C3 on LB + Cm (5ug/ml) |

 [[Image:|300px]]
[[Image:|300px]]
[[Image:|300px]]
[[Image:|300px]]
[[Image:|300px]]
[[Image:|300px]]
References
 "
"