Team:TU-Munich/Modeling/Protein Predictions
From 2013.igem.org
(→Analysis of Receptor Sequences) |
(→Analysis of Receptor Sequences) |
||
| Line 19: | Line 19: | ||
==Analysis of Receptor Sequences== | ==Analysis of Receptor Sequences== | ||
<div class="box-center"> | <div class="box-center"> | ||
| - | For several purposes in our project we needed a synthetic receptor that enables us to express protein-domains at the cellular or extracellular side of the cell membrane. As a template we investigated several different plant-receptors form the well understood dicotyledon ''Arabidopsis thaliana'' and the moss we are currently using as our chassis ''Physcomitrella patens''. The ''A. taliana''-receptors have the advantage that their transgenic expression has successfully been demonstrated (Ref.) whereas the ''P. patens''-receptors bear less risk that they do not function in the evolutionary distant moss.<br> | + | For several purposes in our project we needed a synthetic receptor that enables us to express protein-domains at the cellular or extracellular side of the cell membrane. As a template we investigated several different plant-receptors form the well understood dicotyledon ''Arabidopsis thaliana'' and the moss we are currently using as our chassis ''Physcomitrella patens''. The ''A. taliana''-receptors have the advantage that their transgenic expression has successfully been demonstrated (Ref.) whereas the ''P. patens''-receptors bear less risk that they do not function in the evolutionary distant moss (Ref).<br> |
As there were many different availible receptors that we could use as a template for our synthetic receptor we applied bioinformatical methods to evaluate the suitability of thes receptors. From this work the three examples ERF, FLS2 and SERK are shown (see Table 1). | As there were many different availible receptors that we could use as a template for our synthetic receptor we applied bioinformatical methods to evaluate the suitability of thes receptors. From this work the three examples ERF, FLS2 and SERK are shown (see Table 1). | ||
Revision as of 20:14, 2 July 2013
Prediction of Protein Structures
Search for Homologous Structures using HHprd
Prediction of Possible Structure using iTasser
Analysis of Receptor Sequences
For several purposes in our project we needed a synthetic receptor that enables us to express protein-domains at the cellular or extracellular side of the cell membrane. As a template we investigated several different plant-receptors form the well understood dicotyledon Arabidopsis thaliana and the moss we are currently using as our chassis Physcomitrella patens. The A. taliana-receptors have the advantage that their transgenic expression has successfully been demonstrated (Ref.) whereas the P. patens-receptors bear less risk that they do not function in the evolutionary distant moss (Ref).
As there were many different availible receptors that we could use as a template for our synthetic receptor we applied bioinformatical methods to evaluate the suitability of thes receptors. From this work the three examples ERF, FLS2 and SERK are shown (see Table 1).
Table 1:
Examined Receptors | |||
| Receptor | Organism | Length (aa) | Sequence reference |
| ERF | A. thaliana | 1031 | [http://www.ncbi.nlm.nih.gov/protein/NP_197548.1 NP_197548.1] |
| FLS2 | A. thaliana | 1173 | [http://www.ncbi.nlm.nih.gov/protein/NP_199445.1 NP_199445.1] |
| SERK | P. patens | 625 | [http://www.ncbi.nlm.nih.gov/protein/XP_001759122.1 XP_001759122.1] |
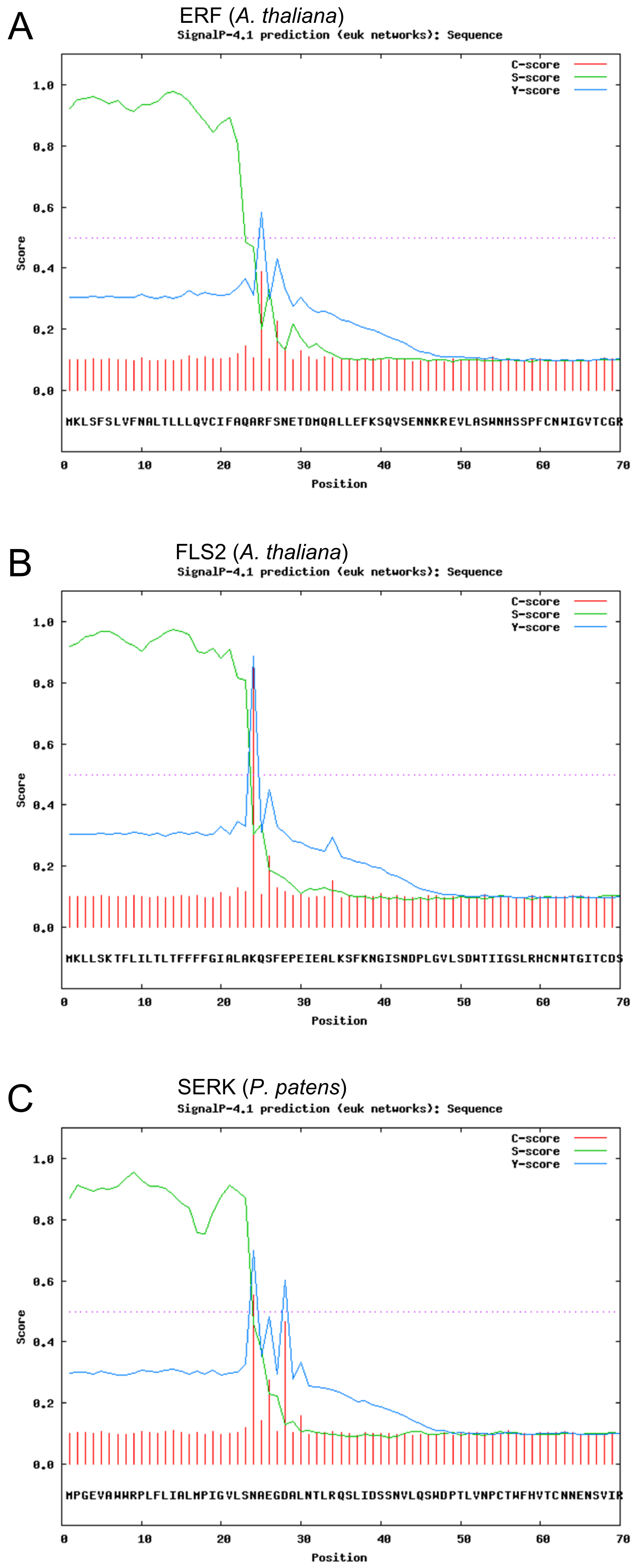
Prediction of Signal Peptides
Introduction: [http://www.cbs.dtu.dk/services/SignalP SignalP 4.1 Server]
Results:
Discussion:
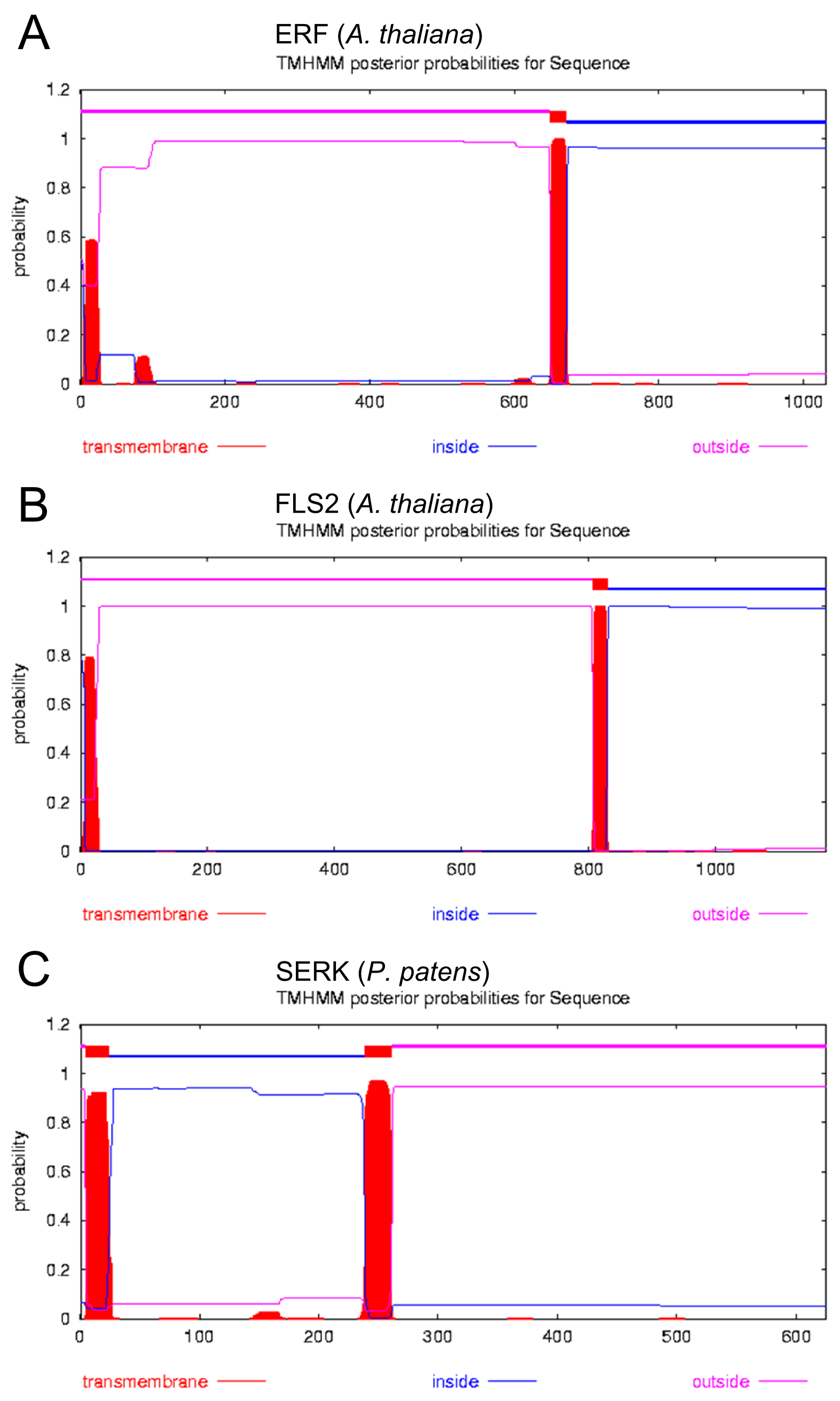
Prediction of Transmembrane Regions
Introduction: [http://www.cbs.dtu.dk/services/TMHMM TMHMM]
Results:
Discussion:
Prediction of Cellular Localization
Introduction: [http://wolfpsort.org Wolfpsort]
Results:
Discussion:
Choice of the SERK Receptor
Analysis of Protein-Ligand Interaction
Structure based prediction using PDBePISA
References:
http://www.ncbi.nlm.nih.gov/pubmed/6327079 Edens et al., 1984
- http://www.ncbi.nlm.nih.gov/pubmed/6327079 Edens et al., 1984 Edens, L., Bom, I., Ledeboer, A. M., Maat, J., Toonen, M. Y., Visser, C., and Verrips, C. T. (1984). Synthesis and processing of the plant protein thaumatin in yeast. Cell, 37(2):629–33.
$(document).ready(function(){
// put the footer in the right place
$("#footer-box").prepend($("#social-footer"));
// implement image preloading
var images = new Array()
function preload() {
for (i = 0; i < preload.arguments.length; i++) {
images[i] = new Image()
images[i].src = preload.arguments[i]
}
}
// preload menu backgrounds
preload( " ",
",
"", "
", "
", "
", "
" );
// preload team pictures
if ( $("div#teamfield").length > 0 ) {
preload( " ",
",
"", "
", "
", "
", "
", "
", "
", "
", "
", // Katrin "
", "
", "
", "
", "
", "
", "
", "
", "
", // Rosario "
", "
", "
", "
", "
", "
", "
", "
", "
", // Fabian "
", "
", "
", "
", "
", "
", "
", "
", "
", // Andreas "
", "
", "
", "
", "
", "
", "
", "
", "
", // Louise "
", "
", "
", "
", "
", "
", "
", "
", "
", // Johanna "
", "
", "
", "
", "
", "
", "
", "
", "
", // Meike "
", "
", "
", "
", "
", "
", "
", "
", "
", // Volker "
", "
", "
", "
", "
", "
", "
", "
", "
", // Polte "
", "
", "
", "
", "
", "
", "
", "
", "
", // Leonie "
", "
", "
", "
", "
", "
", "
", "
", "
", // Philipp "
", "
", "
", "
", "
", "
", "
", "
", "
", // Jeff "
", "
", "
", "
", "
", "
", "
", "
", "
", // Chris "
", "
", "
", "
", "
", "
", "
", "
", "
", // Flo "
", "
", "
", "
", "
", "
", "
", "
" );
}
// Slideshows
$('.bxslider').bxSlider({
responsive: false, auto: true, autoHover: true
});
$('.bxgallery').bxSlider({
slideMargin: 10, minSlides: 3, maxSlides: 3, moveSlides: 1, slideWidth: 5000
});
$("ul.bxgallery img").slimbox({ loop: true }, function(el) { url = el.src; url = url.substring(0, url.lastIndexOf('/')); url = url.replace('/thumb/', '/'); // description = $(el).parents("div.thumbinner").children("div.thumbcaption").text(); return [url, ]; }, function(el) { return (this == el) || (this.parentNode.parentNode && (this.parentNode.parentNode == el.parentNode.parentNode)); });
// Counter and Countdown
function render_counter(c) { i = 0; iid = window.setInterval(function(){ if ( (c-i) > (c/200) ) { $('span#counter').html(i); i += Math.round(c/200); } else { $('span#counter').html(c); window.clearInterval(iid); } }, 10); }
if ($('span#counter').length > 0) { $.ajax({ url: "https://2013.igem.org/Special:PopularPages", success: function( html ) { dom = $.parseHTML(html); visitors = $(dom).find('a[title="Team:TU-Munich"]').parent().text(); visitors = visitors.substring(visitors.indexOf('(')+1); visitors = visitors.substring(0, visitors.indexOf(' ')); visitors = visitors.replace(',', ); render_counter(visitors); }, error: function( xhr, status ) { render_counter(4700); } }); }
if ($('span#countdown) { clock = window.setInterval(function(){ jetzt = new Date(); time_left = Date.UTC(2013, 9, 5, 4, 0, 0) - Date.UTC(jetzt.getUTCFullYear(), jetzt.getUTCMonth(), jetzt.getUTCDate(), jetzt.getUTCHours(), jetzt.getUTCMinutes(), jetzt.getUTCSeconds()); left_sec = (time_left/1000)%60; left_sec = (left_sec < 10) ? "0" + left_sec : left_sec; left_min = Math.floor(time_left/60000)%60; left_min = (left_min < 10) ? "0" + left_min : left_min; left_h = Math.floor(time_left/3600000)%24; left_h = (left_h < 10) ? "0" + left_h : left_h; left_d = Math.floor(time_left/86400000); left_d = (left_d == 1) ? left_d + " day" : left_d + " days"; $('span#countdown').html(left_d + " " + left_h + ":" + left_min + ":" + left_sec); }, 1000); }
// Animate teamfield
if ( $("div#teamfield").length > 0 ) {
var $members = $("div#teamfield a");
$("body").mousemove(function(event){ for (i=0; i<$members.length; i++) {
if ( $members.eq(i).offset().left > event.pageX ) {
if ( $members.eq(i).offset().top > event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("top-left");
} else if ( $members.eq(i).offset().top <= event.pageY && ( $members.eq(i).offset().top + $members.eq(i).height() ) >= event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("left");
} else if ( ( $members.eq(i).offset().top + $members.eq(i).height() ) < event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("bottom-left");
}
} else if ( $members.eq(i).offset().left <= event.pageX && ( $members.eq(i).offset().left + $members.eq(i).width() ) >= event.pageX ) {
if ( $members.eq(i).offset().top > event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("top");
} else if ( $members.eq(i).offset().top <= event.pageY && ( $members.eq(i).offset().top + $members.eq(i).height() ) >= event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("front");
} else if ( ( $members.eq(i).offset().top + $members.eq(i).height() ) < event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("bottom");
}
} else if ( ( $members.eq(i).offset().left + $members.eq(i).width() ) < event.pageX ) {
if ( $members.eq(i).offset().top > event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("top-right");
} else if ( $members.eq(i).offset().top <= event.pageY && ( $members.eq(i).offset().top + $members.eq(i).height() ) >= event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("right");
} else if ( ( $members.eq(i).offset().top + $members.eq(i).height() ) < event.pageY ) {
$members.eq(i).removeClass();
$members.eq(i).addClass("bottom-right");
}
}
} });
}
});
 "
"





AutoAnnotator:
Follow us:
Address:
iGEM Team TU-Munich
Emil-Erlenmeyer-Forum 5
85354 Freising, Germany
Email: igem@wzw.tum.de
Phone: +49 8161 71-4351