Team:Linkoping Sweden/Project
From 2013.igem.org














Contents |
The basics
The idea is basically to first grow E.Coli that has been genetically modified to produce an antibody with the enzyme luciferase attached to its constant part. At first the antibody will be constructed to adhere to an egg antigen, but our long time goal is to be able to switch to whichever antigen desired. Luciferase is an enzyme that cleaves luciferin in the presence of ATP, giving rise to a long lasting green light by the principle of luminescence. Luminescence is superior to fluorescence due to the fact that it will not fade, amongst other things.
Project Video
Antibody fusion protein
The first step in our project will be to create a fusion protein consisting of a luciferase linked together with a single domain antibody from a cameloid, VHH. The idea behind using these antibodies is the fact that they consist only of a single domain. This makes it possible for them to be expressed lesser organisms, such as E. coli, something that would be very hard for conventional antibodies found in other mammals.
Our ulterior motive behind cloning an antibody in E. coli is that it allows us to engineer the protein after our preferences. What we want to do is connect the VHH antibody to a luciferase protein via a linker. If the translation of the fusion protein is successful, the luciferase will be active when luciferin and ATP is added to the solution.
Another factor to consider is the binding affinity that the antibody part of the fusion protein has for its antigen. This is probably the most critical part of our experiment. The challenge lies in trying to make the antibody regain most of its affinity, while being connected to another protein via a linker.
Antigen fusion protein
With the antibody affinity for the antigen maintained, we will have a fusion protein which successfully binds to an antigen and that emits light when luciferin and ATP is added. However, the protein should emit light regardless if it has an antigen bound or not. As such, this alone would not be sufficient to determine the presence of an antigen.
The second stage of our project will be to engineer a similar fusion protein as the first one, but this time it will consist of the antigen linked together with Red Fluorescent Protein (RFP). If successful, our two fusion proteins will have affinity for each other.
The idea behind using RFP is that it absorbs light at the same wavelengths where luciferase emits them, causing a quenching effect through fluorescence resonance energy transfer (FRET). This FRET will only occur where a fusion antigen has bound to a fusion antibody as fusion antigens and fusion antibodies normally would be too far away for FRET to be observed.
We expect more fusion antigens to be bound to fusion proteins when no free antigen is present in the solution as there will be less competition for the antibodies. This will in turn lead to a difference in emission spectra between a blank containing no free antigens and a sample containing free antigen. Our idea can be summarized as follows.
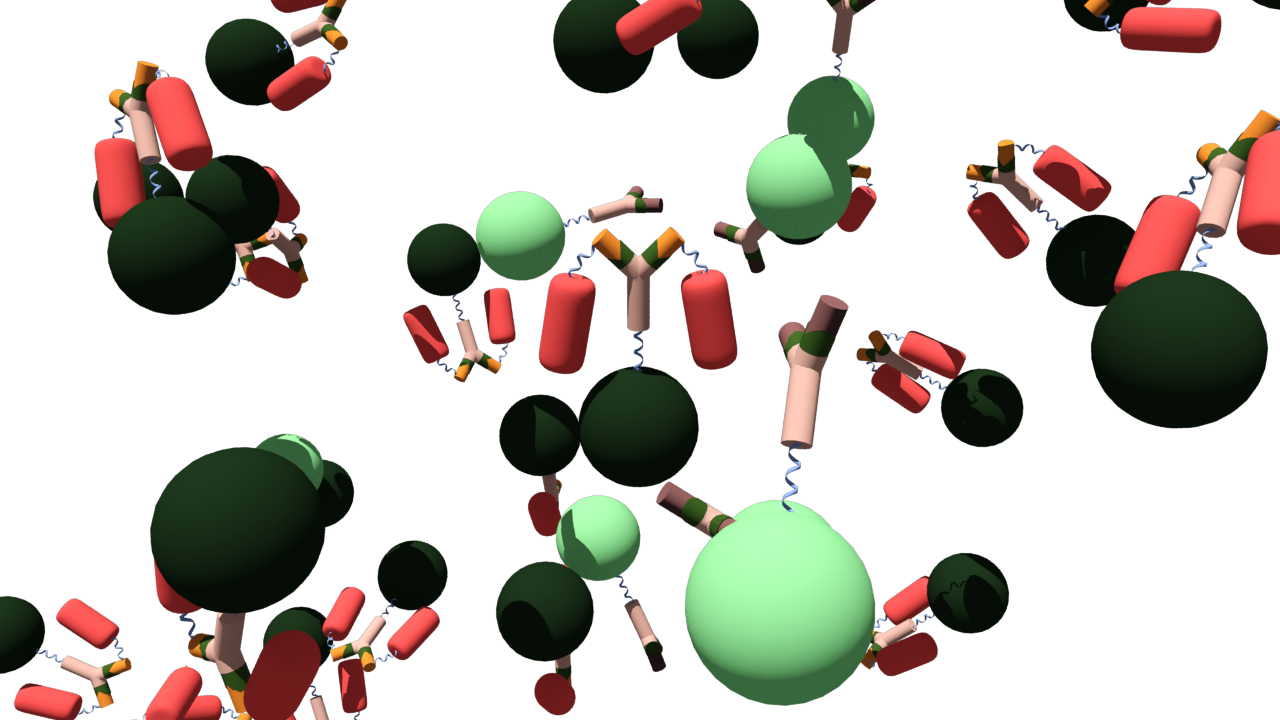
As the antigen (brown) binds to the antibody-Luciferase fusion protein (beige and green) green luminescence can be seen. If the false antigen-RFP fusion protein (yellow and red) binds to the antibody-Luciferase fusion protein, luciferase is quenched and the FRET mechanism causes fluorescence of the RFP.
Ref: Branchini et al. Analytical Biochemistry 414 (2011) 239–245. Sequential bioluminescence resonance energy transfer–fluorescence resonance energy transfer-based ratiometric protease assays with fusion proteins of firefly luciferase and red fluorescent protein.






 "
"