The second part is a biosensor of the hydrolysis product PNP. As there is no promoter that can react to parathion, we hope that the sensing of this product can reflect the existence of parathion.But there is no such a PNP-sensing part in the iGEM registry, so we start building up the sensor from scratch. We target a PNP-oxidation gene cluster isolated from Rhodococcus sp.

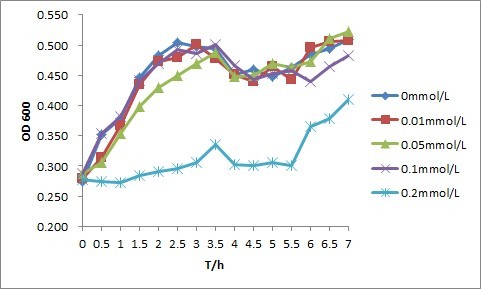
PNP are toxic to cells. Since our experiments are mostly under the condition of PNP, we test the tolerance of E.coli DH5-α under various concentration of PNP before other experiments.

According to the previous study of Takeo, nphA1 and nphA2 encode enzymes involved in PNP-degradation pathway and nphR encodes an araC/XylS-type regulatory protein. The transcription of nphA1 is only induced in the presence of both PNP and NphR is constitutively expressed. We find the promoter region which can react to PNP in nphA1 and utilized NphR to construct the biosensor.

We construct NphR into pSB1A3 which keeps it constitutively expressed, and the promoter region with GFP reporter into pSB3K3. Also, we choose strong RBS BBa_B0034. After construction, we transform these two vectors into the E coli. to test it sensing ability.
We do a series of experiments about our sensor's transcription efficiency. We add PNP to liquid medium and incubated with our transgenic E.coli for several hours. And then we measure the fluorescence intensity with excitation wavelength at 470nm and emission wavelength at 509nm. We also measure absorption at 600nm which reflects E.coli's growth.
According to the experimental results, we find that our sensor does work. It shows that GFP transcription efficiency has a positive correlation relationship with time at the presence of PNP, though it will take a few hours before we can detect the changes. What's more, we think transcription efficiency may also have a positive correlation relationship with PNP concentration but at a very low level. The sensor will be in optimal situation when the concentration of PNP is 0.05mmol/L.

Is nphR needed? To ensure that our conclusion is reliable,we design another experiment which NphR is deficient. As the result shows, NphR is necessary.


We will do further work to examine the promoter's inducing curve by a more specified gradient of PNP concentration. However, there still have some problems. The biggest problem is that the GFP intensity only rises at low level. Maybe we should use other bacteria or look for the more precise promoter region.
In fact,in another experiment about the tolerance of PNP, we have drawn the conclusion that E.coli growth more fast on the condition that the concentration of PNP is 0.05mmol/L. At this concentration, the sensor is most sensitive. Maybe PNP plays a role as a carbon resource in low concentration but a toxin in high concentration. No concrete research results are served about that. Anyway, the discovery of the optimal concentration may be a useful guideline of our product. To deal with those samples with high PNP concentration, we can dilute the certain samples reasonably,enhancing the efficiency of the sensor.
 "
"

