Team:Imperial College/data
From 2013.igem.org
| (38 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Imperial_College/Templates:header}} | {{:Team:Imperial_College/Templates:header}} | ||
| + | <h1>Data</h1> | ||
| + | <h2>Media characterisation</h2> | ||
| + | <h3>Growth assays with different experimental media</h3> | ||
| + | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
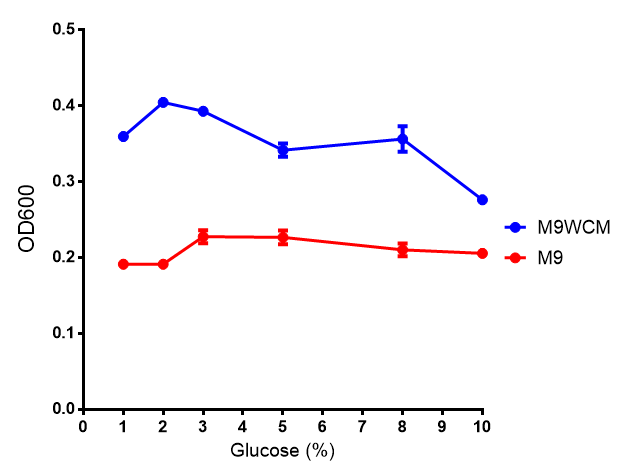
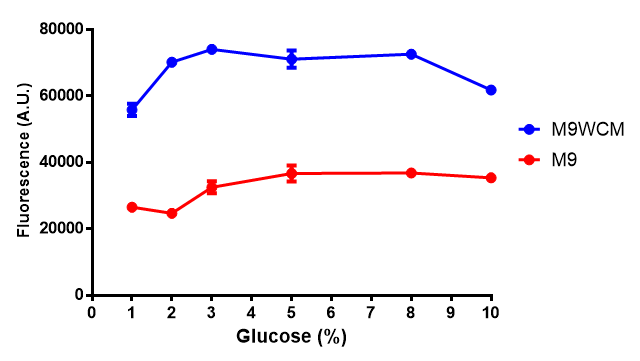
| + | |[[File:LB_M9.png|thumbnail|left|400px|<b>Figure 1: The medias used for our growth assays using the MG1655 strain.</b> phaCAB is an operon involved in synthesis of P3HB while EV represents an empty vector. SR refers to stress response cells containing a red pigment. M9M is minimal media, while M9S is supplemented minimal media, as described [https://2013.igem.org/Team:Imperial_College/Protocols#M9_minimal_and_supplemented_media here]. WCM refers to waste conditioned media, which is made from sterile filtrated SRF waste, see [https://2013.igem.org/Team:Imperial_College/Protocols#Waste_Conditioned_Media_.28WCM.29 here]. Error bars are SEM, n=4.]] | ||
| + | |[[File:Media8888.jpg|thumbnail|right|400px|Fig]] | ||
| + | |} | ||
| - | |||
| - | |||
| - | |||
| - | |||
<br> | <br> | ||
| - | < | + | <h2 class="clear">Waste Assays</h2> |
| + | These assays were designed to test whether our chassis could grow either directly with waste and/or in waste conditioned media. Waste conditioned media is a filter sterilised version of the waste media and was designed for several reasons; we were unsure whether mixed waste would be toxic to Ecoli and hence a less concentrated version may be more suitable and secondly large chunks of waste would prevent accurate OD600 measurements and therefore we decided to filter out the largest chunks. | ||
| + | |||
| + | |||
| + | |||
| + | <h3>Waste media </h3> | ||
| + | |||
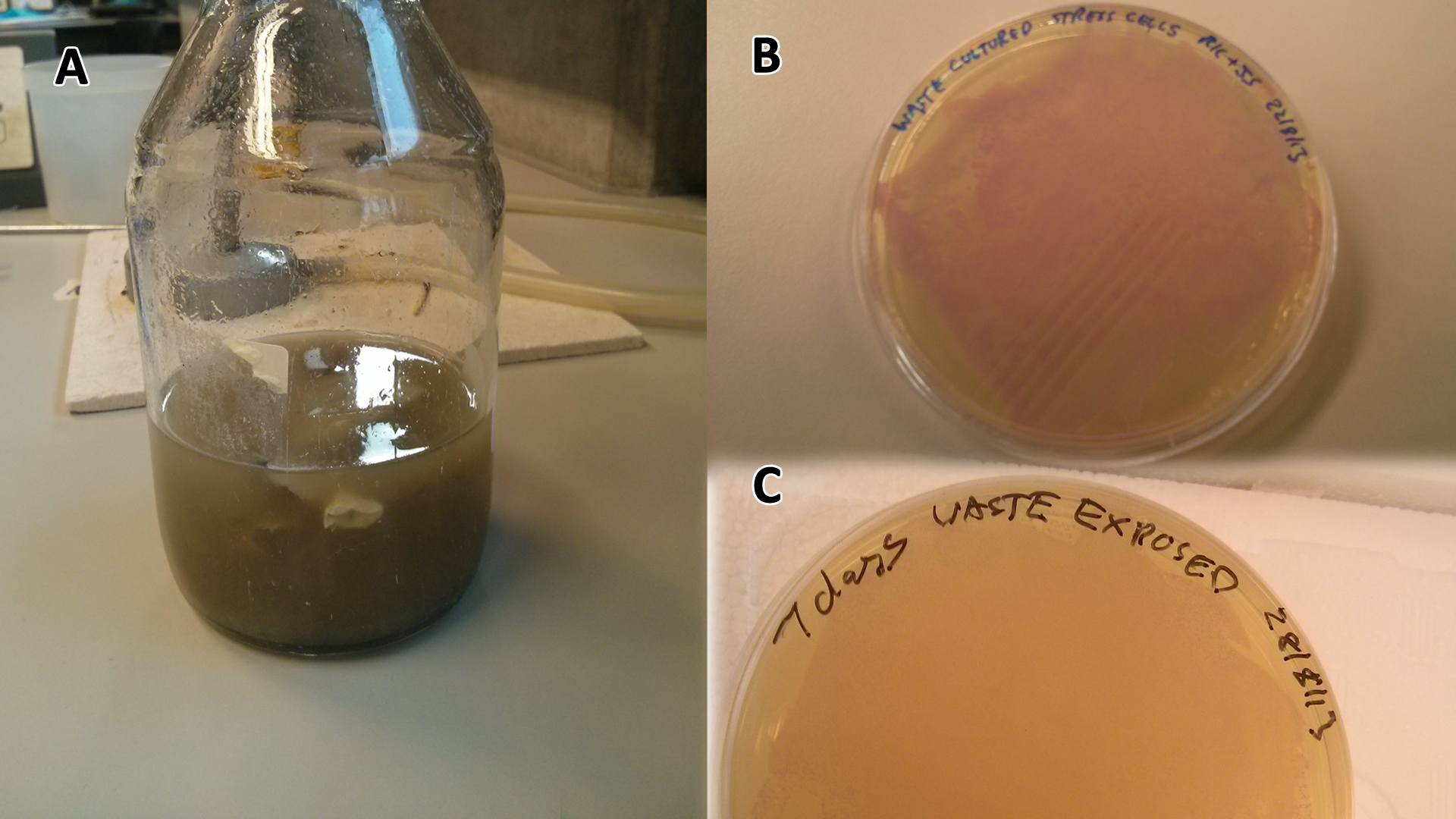
| + | We can see from this experiment whether our bacteria can grow solely on the mixed waste. | ||
| - | |||
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
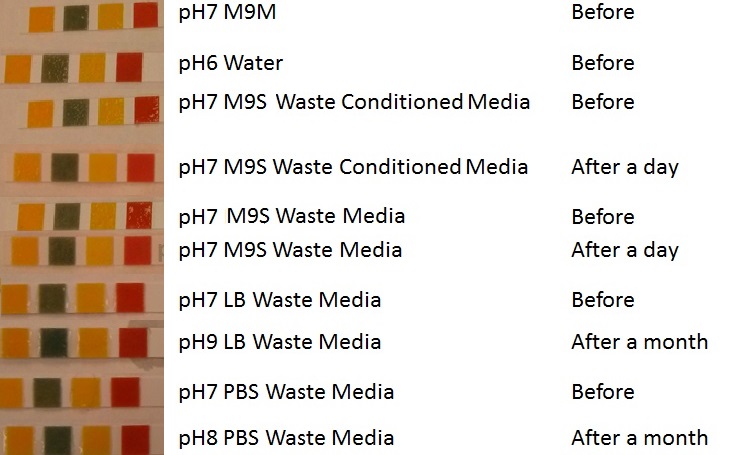
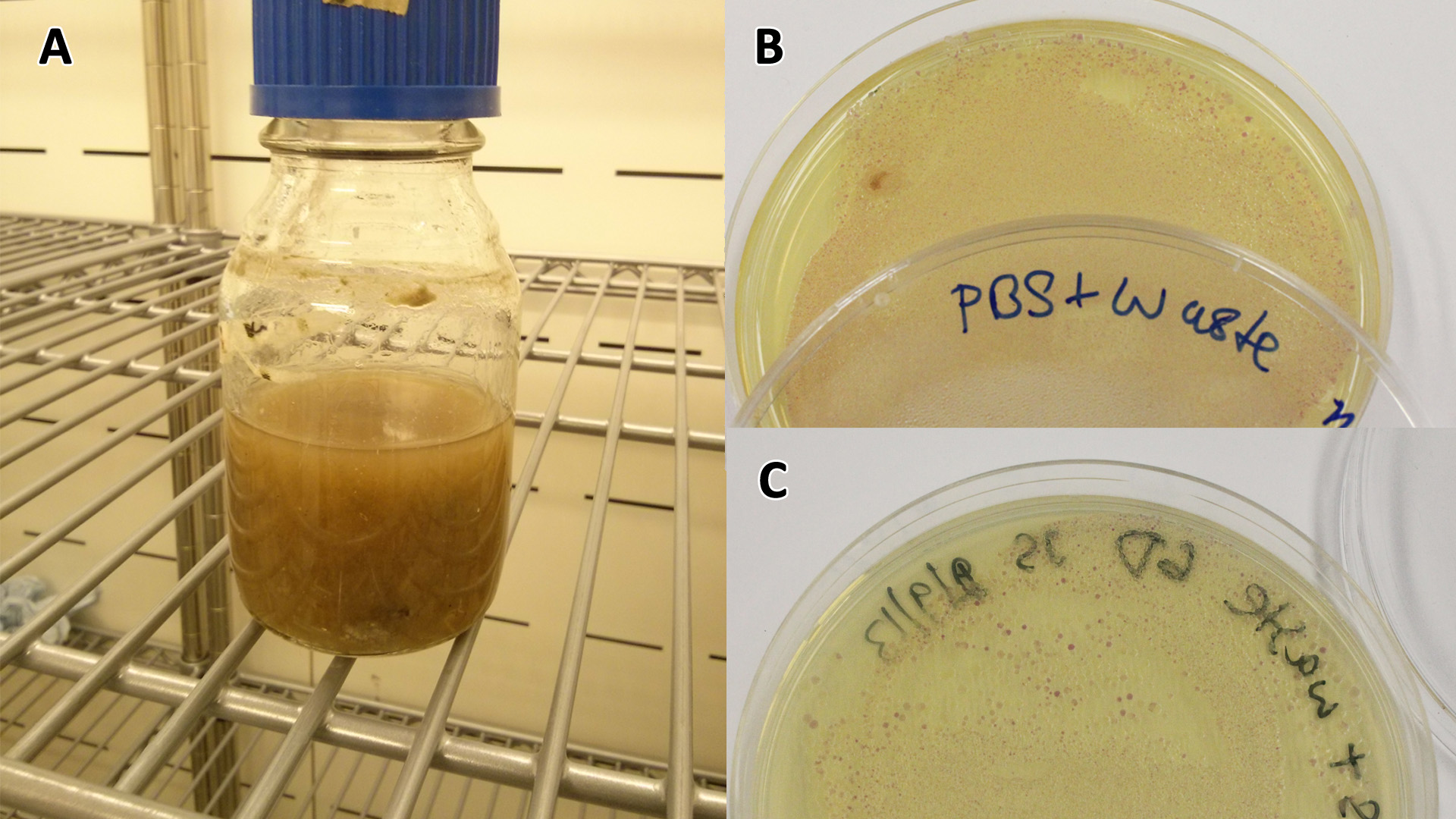
| - | |[[File:Waste_cocktail.png|thumbnail|center|400px|<b>(A)</b> | + | |[[File:Waste_cocktail.png|thumbnail|center|400px|<b>(A)</b> Waste media<b>(B)</b> Ecoli containing mCherry stress biosensor (BBa_K639003) were grown in mixed waste <b>(A)</b> over 3 days, then streaked in a qualitative assay to check for growth. <b>(C)</b> mCherry stress biosensor (BBa_K639003) transformed Ecoli were streaked again after 7 days growth in SRF.]] |
| - | |[[File:PBS_Plus_Waste.jpg|thumbnail|center|400px|<b>(A)</b> | + | |[[File:PBS_Plus_Waste.jpg|thumbnail|center|400px|<b>(A)</b> waste media made up in PBS (phosphate buffered saline). <b>(B)</b> E coli expressing mCherry stress biosensor (BBa_K639003) grown in waste media <b>(A)</b> over 3 days, then streaked onto an antibiotic containing plate to qualitatively assess whether the E. coli had survived. <b>(C)</b> Streaked again after 6 days growth in SRF.]] |
|} | |} | ||
| + | |||
| + | Conclusion MG1655 can survive solely on mixed waste. | ||
| - | < | + | <h3>Waste conditioned media</h3> |
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
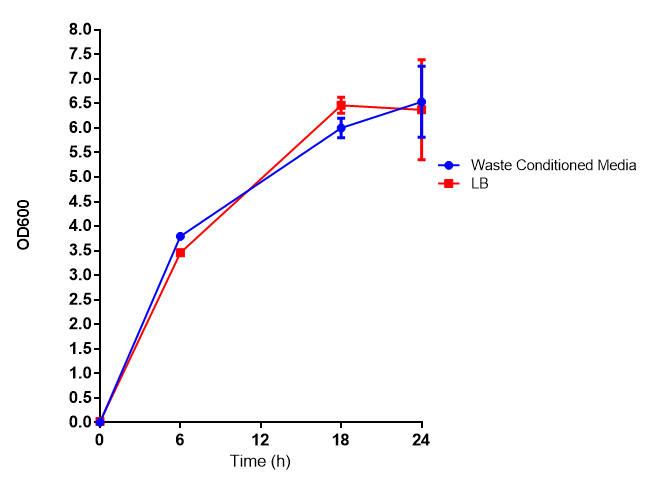
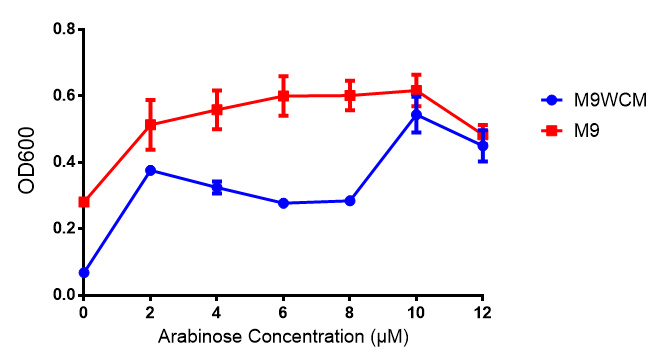
|[[File:WCM_media.png|thumbnail|right|400px|Growth curve of our [http://parts.igem.org/Part:BBa_K639003 mCherry] MG1655 bacteria. MG1655 were grown with LB media and sterile filtrated WCM at 37ºC. Error bars represents SE of the mean, n=4]] | |[[File:WCM_media.png|thumbnail|right|400px|Growth curve of our [http://parts.igem.org/Part:BBa_K639003 mCherry] MG1655 bacteria. MG1655 were grown with LB media and sterile filtrated WCM at 37ºC. Error bars represents SE of the mean, n=4]] | ||
| - | |[[File:Very_stressed_ecoli.jpg|thumbnail|left|400px| | + | |[[File:Very_stressed_ecoli.jpg|thumbnail|left|400px|mCherry stress biosensor (BBa_K639003) transformed MG1655 were grown with [https://2013.igem.org/Team:Imperial_College/Protocols LB-WCM] for 48 hours.]] |
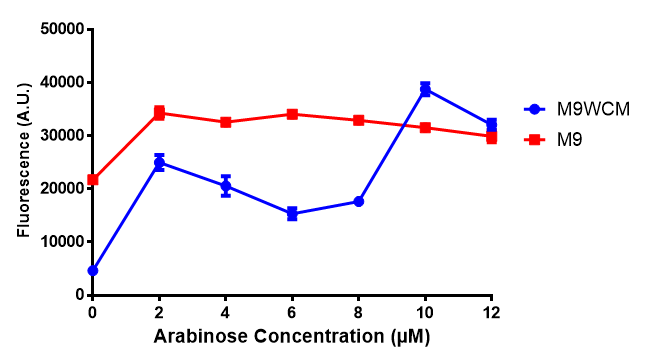
| + | [[File:WCM2.png|thumbnail|center|400px|<b>Figure 1: MG1655 in WCM with plasmids EV and SR.</b> There is no growth inhibition when comparing the empty vector with the mCherry stress biosensor (BBa_K639003) vector in each media. WCM contains LB but trends in a simliar manner to M9S, suggesting that the chassis most likely due to toxins present within it from the mixed waste. Error bars are SEM, n=4.]] | ||
|} | |} | ||
| - | |||
| + | <h2 class="clear">Characterisation of our Biobricks</h2> | ||
| - | < | + | {| class="wikitable" style="margin: 1em auto 1em auto;" |
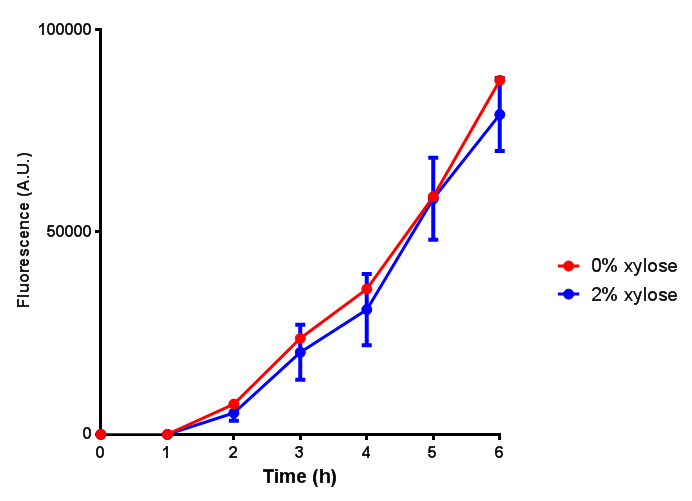
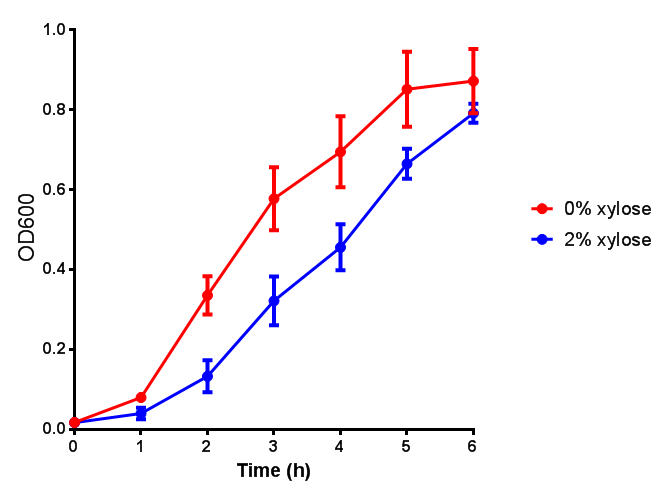
| + | | [[File:Bdh2_fluor.png|thumbnail|right|400px|<b>Bdh2 induction to gauge growth effect.</b> Error bars are SEM, n=4.]] | ||
| + | | [[File:Bdh2.png|thumbnail|right|400px|]] | ||
| + | |} | ||
| + | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| + | | [[File:CLE_fluor.png|thumbnail|right|400px|<b>CLE induction to gauge growth effect.</b> Error bars are SEM, n=4.]] | ||
| + | | [[File:CLE.png|thumbnail|right|400px|]] | ||
| + | |} | ||
| + | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| + | | [[File:ESTCS2_fluor.png|thumbnail|right|400px|<b>ESTCS2 induction to gauge growth effect.</b> Error bars are SEM, n=4.|]] | ||
| + | | [[File:ESTCS2.png|thumbnail|right|400px|]] | ||
| + | |} | ||
| + | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| + | |[[File:PulA.png|thumbnail|left|400px|<b>PulA induction to gauge growth effect.</b> Error bars are SEM, n=4.]] | ||
| + | |[[File:PueB.png|thumbnail|right|400px|<b>PueB induction to gauge growth effect.</b> Error bars are SEM, n=4.]] | ||
| + | |} | ||
| + | |||
| + | <h3 style="clear: left">Empty Vector Control</h3> | ||
| + | [[File:EV.png]] | ||
| - | |||
| - | < | + | <h2>Characterisation of existing Biobricks</h2> |
| - | < | + | <h3>phaCAB biobrick characterisation</h3> |
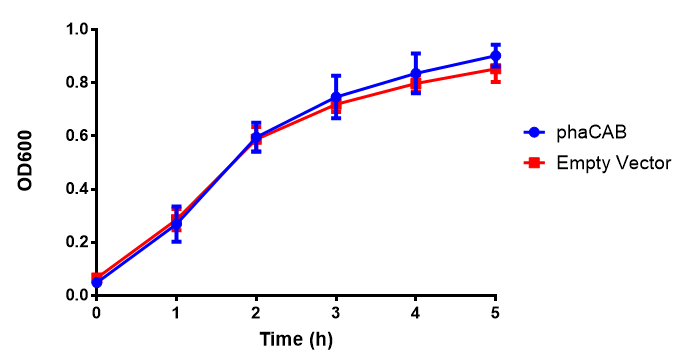
| - | < | + | <h4 class="clear">LB</h4> |
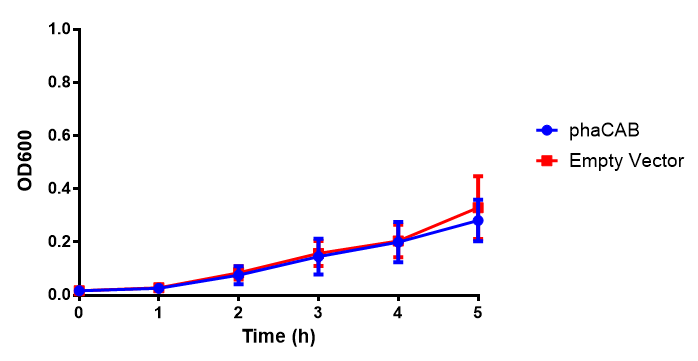
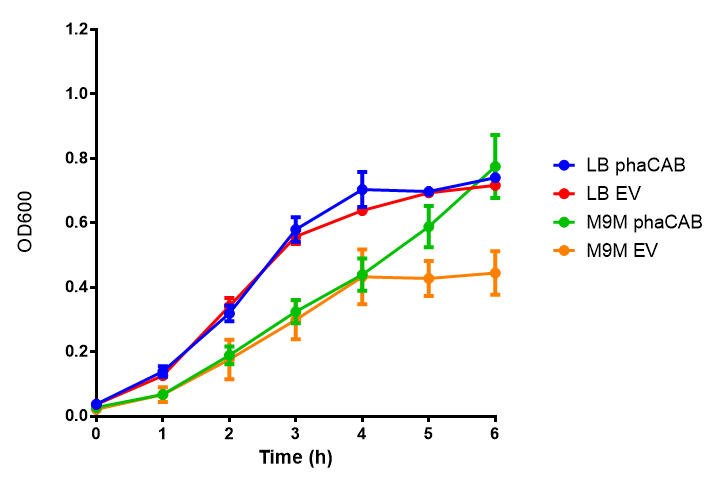
[[File:LB.png|thumbnail|center|500px|<b>Figure 1: MG1655 in LB with plasmids EV and phaCAB.</b> There is no growth inhibition when comparing the empty vector with the phaCAB vector in each media. LB shows the strongest growth curve with minimal latency. Error bars are SEM, n=4.]] | [[File:LB.png|thumbnail|center|500px|<b>Figure 1: MG1655 in LB with plasmids EV and phaCAB.</b> There is no growth inhibition when comparing the empty vector with the phaCAB vector in each media. LB shows the strongest growth curve with minimal latency. Error bars are SEM, n=4.]] | ||
| - | < | + | <h4 class="clear">M9 Minimal</h4> |
[[File:M9M.png|thumbnail|center|500px|<b>Figure 1: MG1655 in M9M with plasmids EV and phaCAB.</b> There is no growth inhibition when comparing the empty vector with the phaCAB vector in each media. M9M shows the least growth growth of all the medias as it has low carbon and amino acid content. Error bars are SEM, n=4.]] | [[File:M9M.png|thumbnail|center|500px|<b>Figure 1: MG1655 in M9M with plasmids EV and phaCAB.</b> There is no growth inhibition when comparing the empty vector with the phaCAB vector in each media. M9M shows the least growth growth of all the medias as it has low carbon and amino acid content. Error bars are SEM, n=4.]] | ||
| - | < | + | <h4 class="clear">M9 Supplemented</h4> |
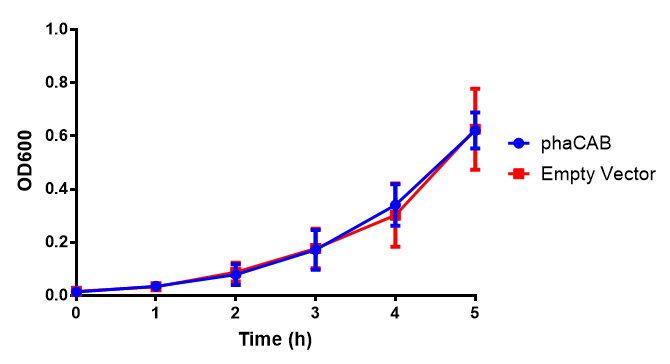
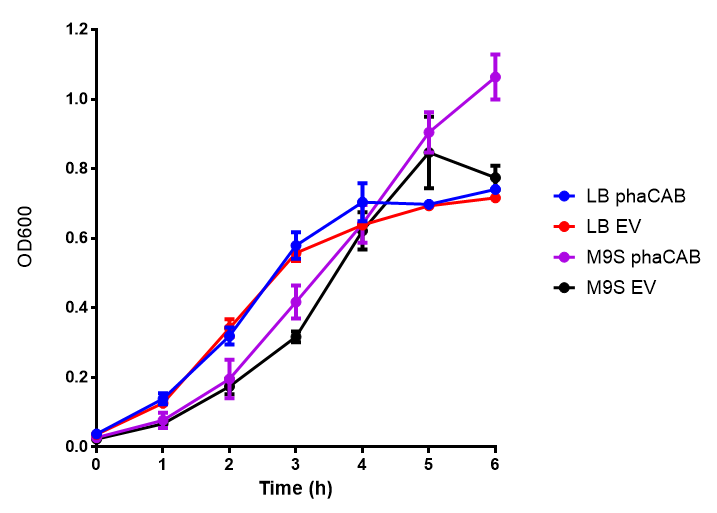
[[File:M9S.png|thumbnail|center|500px|<b>Figure 1: MG1655 in M9S with plasmids EV and phaCAB.</b> There is no growth inhibition when comparing the empty vector with the phaCAB vector in each media. M9S shows a lag phase in growth but quickly increases due to increased amino acid content, nearly reaching LB after 5h. Error bars are SEM, n=4.]] | [[File:M9S.png|thumbnail|center|500px|<b>Figure 1: MG1655 in M9S with plasmids EV and phaCAB.</b> There is no growth inhibition when comparing the empty vector with the phaCAB vector in each media. M9S shows a lag phase in growth but quickly increases due to increased amino acid content, nearly reaching LB after 5h. Error bars are SEM, n=4.]] | ||
| - | < | + | <h3>Stress biosensor characterisation BBa_K639003</h3> |
| - | < | + | <h4>IPTG induction assay</h4> |
Originally we intended on using [http://parts.igem.org/Part:BBa_K639003 BBa_K639003] to detect whether our cells were stressed when placed in various toxic byproducts. However, as the data below shows, this biobrick is very leaky. As such, we are using the stress sensor as a marker for cell growth and also to show that the cells had been successfully transformed with the correct chloramphenicol resistance. | Originally we intended on using [http://parts.igem.org/Part:BBa_K639003 BBa_K639003] to detect whether our cells were stressed when placed in various toxic byproducts. However, as the data below shows, this biobrick is very leaky. As such, we are using the stress sensor as a marker for cell growth and also to show that the cells had been successfully transformed with the correct chloramphenicol resistance. | ||
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| Line 57: | Line 86: | ||
|} | |} | ||
| + | [[File:StressResponse.jpg|400px|thumb|left|BBa_K639003 transformed into E coli. strain MG1655. Pink colonies are visible, which relate to 'leaky' RFP production]] | ||
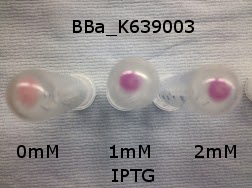
| - | < | + | [[File:K639003IPTG.jpeg|500px|thumb|left|BBa_K639003 transformed into E coli. strain MG1655. Cells were grown at 37oC in 4ml LB with 0, 1 or 2mM IPTG. At 6 hours post IPTG induction, cells were spun down.]] |
| + | |||
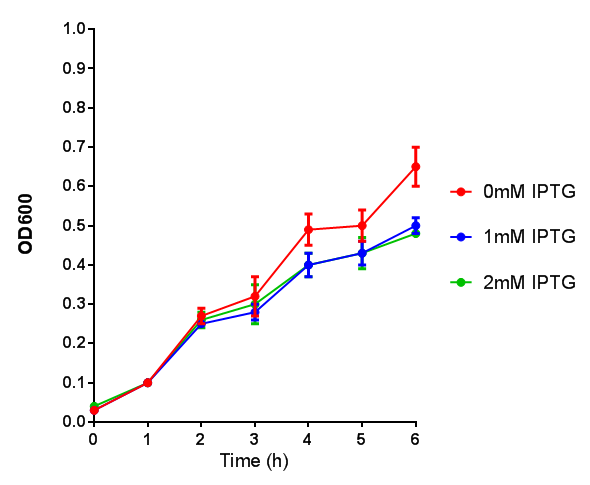
| + | [[File:BBa K639003OD600.jpg|500px|thumb|left|Growth curve. BBa_K639003 transformed into E coli. strain MG1655. Cells were grown at 37oC in 4ml LB with 0, 1 or 2mM IPTG]] | ||
| + | |||
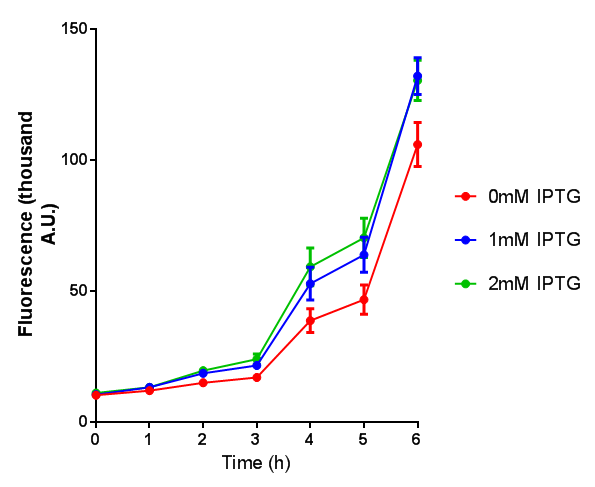
| + | [[File:BBa K639003FL.jpg|500px|thumb|left| FL. BBa_K639003 transformed into E coli. strain MG1655. Cells were grown at 37oC in 4ml LB with 0, 1 or 2mM IPTG]] | ||
| + | |||
| + | |||
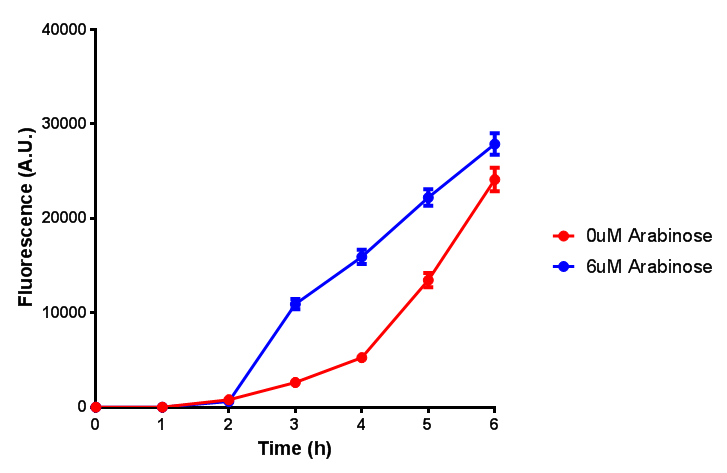
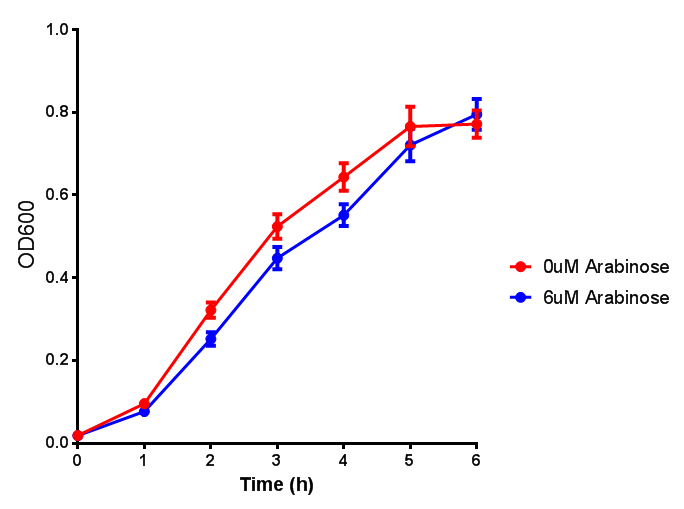
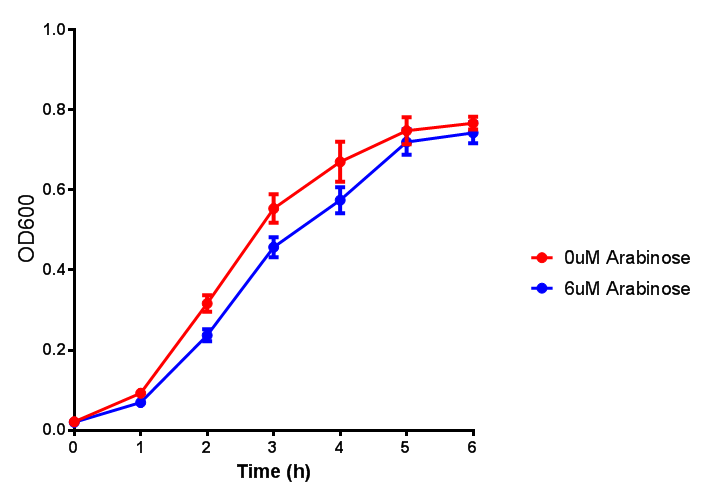
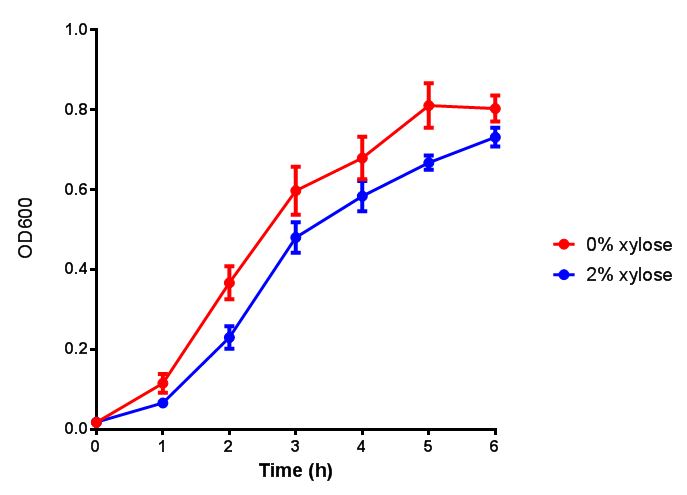
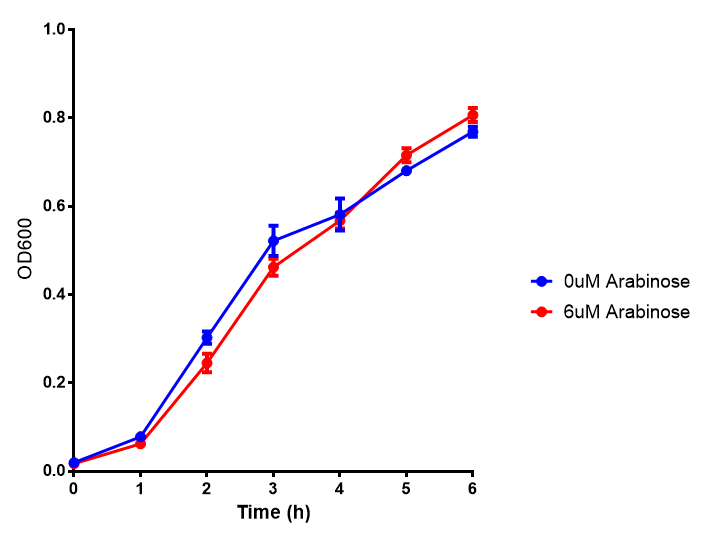
| + | <h3>pBAD characterisation</h3> | ||
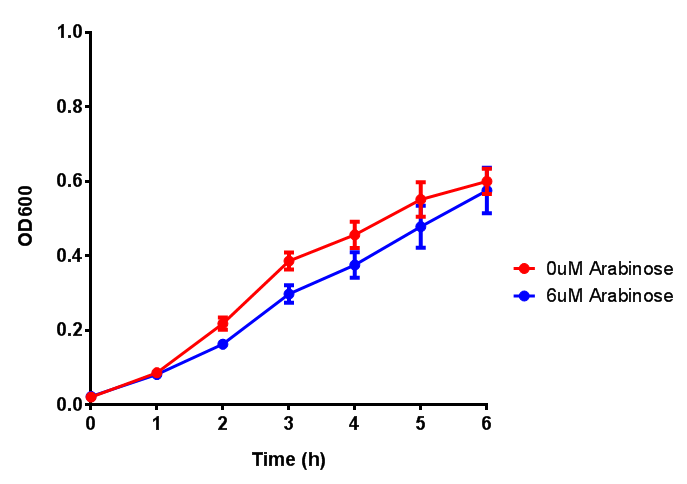
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| Line 71: | Line 108: | ||
| - | < | + | <h3>Glucose </h3> |
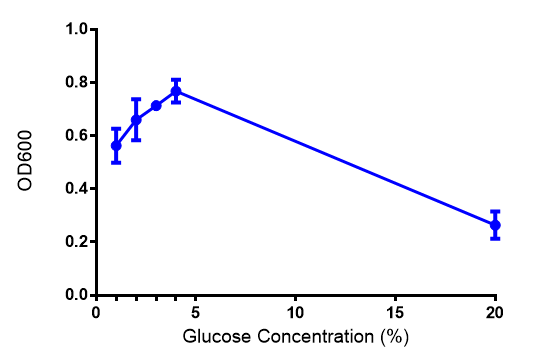
| - | [[File:Glucose_graph.png|thumbnail|center|600px|Cell growth of | + | [[File:Glucose_graph.png|thumbnail|center|600px|Cell growth of phaCAB <i>E. coli</i> at 4 concentrations of glucose. Optimum growth is at 2-4% glucose at 37ºC. Error bars represents SE of the mean, n=4]] |
| - | + | ||
| - | + | ||
| - | + | ||
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| Line 83: | Line 117: | ||
ANOVA analysis shows that... | ANOVA analysis shows that... | ||
| + | <h3>Plastic Toxicity Assays</h3> | ||
| - | < | + | <h4>L-lactic Acid </h4> |
| - | + | ||
| - | + | ||
| - | + | ||
[[File:L-LActic_Acid.png|thumbnail|center|600px|Cell growth of MG1655 on 5mM L-Lactic Acid. Error bars represents SE of the mean, n=4.]] | [[File:L-LActic_Acid.png|thumbnail|center|600px|Cell growth of MG1655 on 5mM L-Lactic Acid. Error bars represents SE of the mean, n=4.]] | ||
<h3>Ethylene glycol </h3> | <h3>Ethylene glycol </h3> | ||
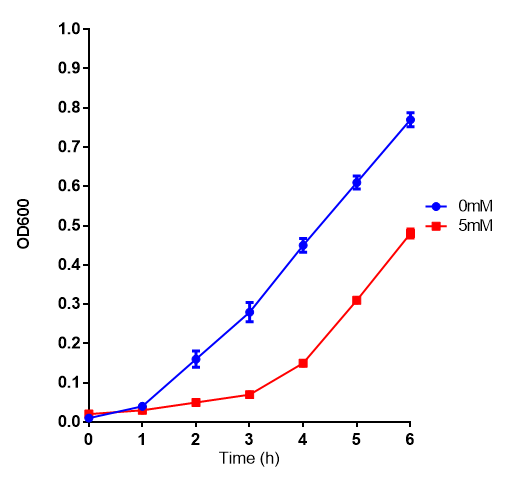
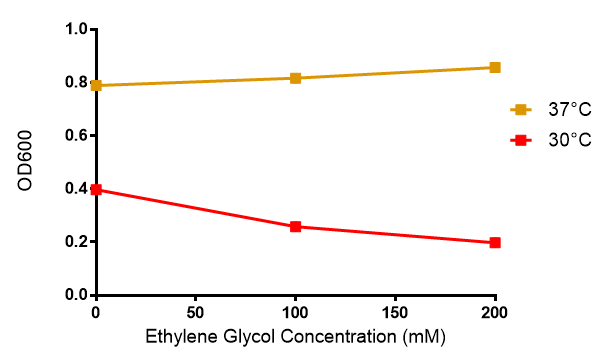
[[File:EG_growth.png|thumbnail|center|600px|Cell growth of MG1655 in ethylene glycol, a byproduct of polyurethane degradation. Cells were grown in 0mM, 100mM or 200mM Ethylene Glycol at 30ºC. Error bars represents SE of the mean, n=4]] | [[File:EG_growth.png|thumbnail|center|600px|Cell growth of MG1655 in ethylene glycol, a byproduct of polyurethane degradation. Cells were grown in 0mM, 100mM or 200mM Ethylene Glycol at 30ºC. Error bars represents SE of the mean, n=4]] | ||
| - | < | + | |
| + | Reduced growth at 30oC likely due to decreased efficiency of MG1655 ethylene glycol break down enzymes. These enzymes (see UC Davis 2012) are endogenously expressed and detoxify Ethylene Glycol. | ||
| + | |||
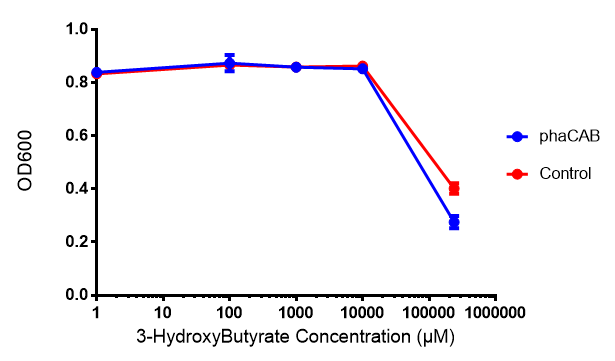
| + | <h4>3-hydroxybutyrate (3HB) </h4> | ||
[[File:3HB666.png]] | [[File:3HB666.png]] | ||
| - | < | + | <h4>Acetoacetate </h4> |
[[File:AA777.png]] | [[File:AA777.png]] | ||
| - | < | + | <h4>Poly(3-hydroxybutyrate) P(3HB) </h4> |
[[File:31ug_P3HB.png]] | [[File:31ug_P3HB.png]] | ||
| - | < | + | <h4>Poly(lactic acid) (PLA) </h4> |
[[File:PLA.png]] | [[File:PLA.png]] | ||
| - | < | + | <h3>Sole carbon source</h3> |
| + | <h4>3HB</h4> | ||
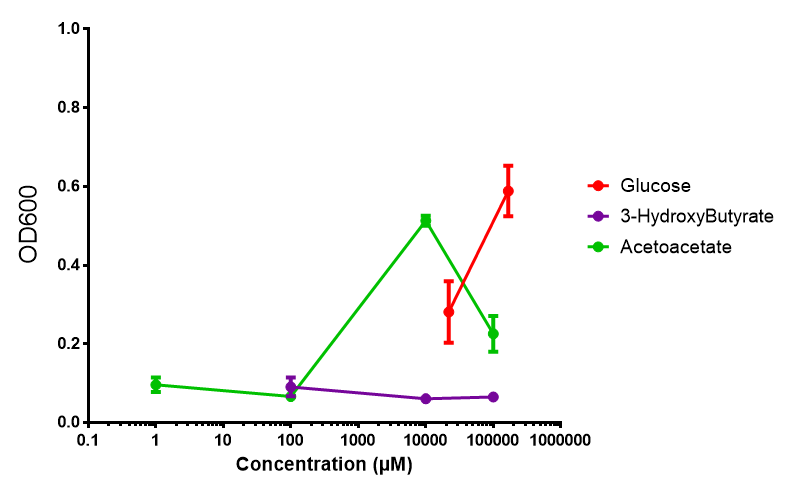
| + | [[File:3HB_sole_carbon_source.png]] | ||
| + | <h4>Acetoacetate</h4> | ||
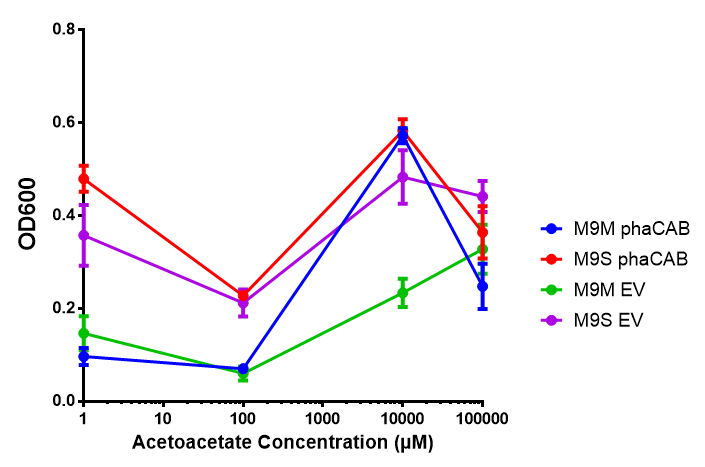
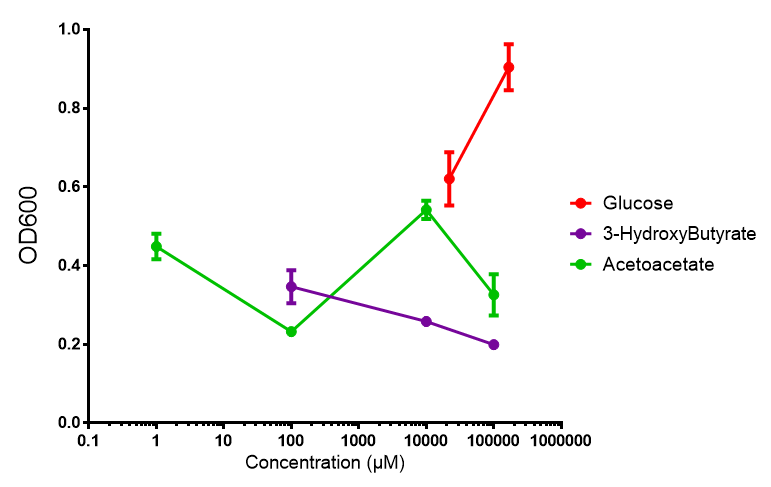
| + | [[File:AA_sole_carbon_source.png]] | ||
| + | [[File:M9M_phaCAB.png]] | ||
| + | [[File:M9S_phaCAB.png]] | ||
| - | < | + | <h2>Western blots</h2> |
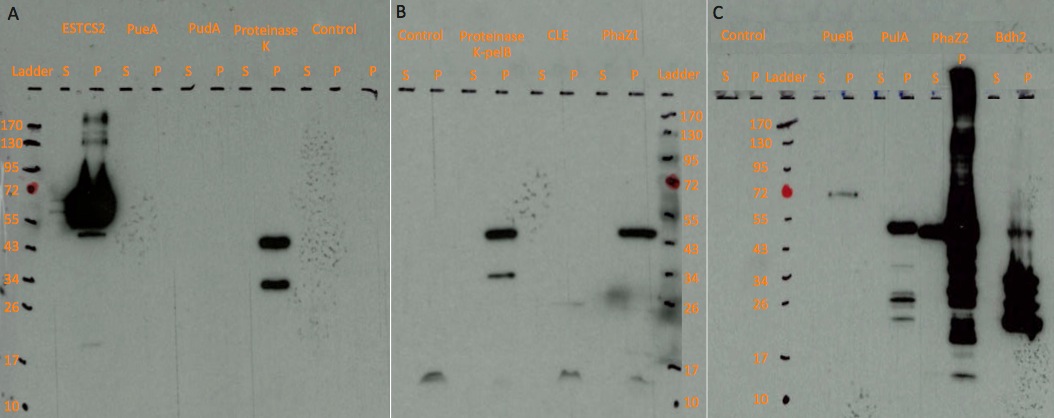
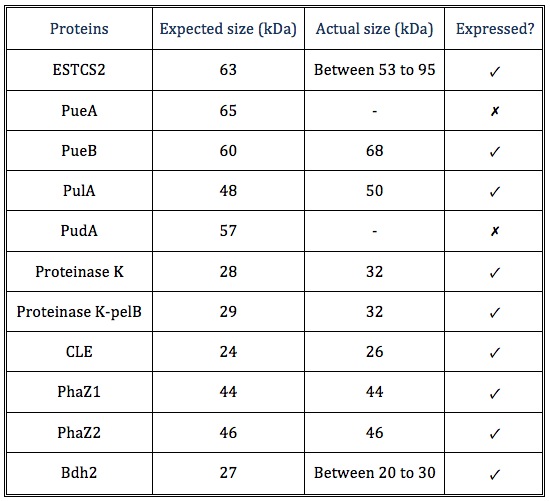
| + | [[File:Western_blot.png|thumbnail|center|950px|Figure: Western blot results on three chemiluminescent-compatible X-ray films. For each protein, we ran both lysed pellet(P) and supernatant(S) sample on gel, in order to test protein expression and secretion respectively.]] | ||
| + | [[File:Western blot protein sizes.png|center]] | ||
| + | <h2>Enzyme Kinetics</h2> | ||
| + | <h2>PHB production</h2> | ||
| + | <h3 class="clear">Nile red staining</h3> | ||
| + | [[File:Nilered.JPG|thumbnail|right|400px|<b>left</b> control cells <b>right phaCAB transformed MG1655</b>]] <br><br><br> | ||
| + | O/N cutures of MG1655 transformed with either control or phaCAB plasmid were spread onto LB-agar plates with 3% glucose and Nile red staining. The staining indicates the production of P3HB. | ||
| - | |||
| - | + | <h3 class="clear">Purification of P3HB</h3> | |
| - | <h3> | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
{| class="wikitable" style="margin: 1em auto 1em auto;" | {| class="wikitable" style="margin: 1em auto 1em auto;" | ||
| - | | [[File: | + | |[[File:IMG_2191.JPG|thumbnail|left|200px|P(3HB) purified from phaCAB transformed MG1655 that were grown in LB with 3% glucose]] |
| - | | [[File: | + | |[[File:IMG_2191.JPG|thumbnail|right|200px|P(3HB) purified from phaCAB transformed MG1655 that were grown in LB with 3% glucose]] |
| + | |[[File:Plasticfrommixedwaste.jpg|thumbnail|center|250px|P(3HB) purified from phaCAB transformed MG1655 that were grown in M9M mixed waste.]] | ||
|} | |} | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
{{:Team:Imperial_College/Templates:footer}} | {{:Team:Imperial_College/Templates:footer}} | ||
Latest revision as of 13:26, 25 September 2013
Contents
|
Data
Media characterisation
Growth assays with different experimental media
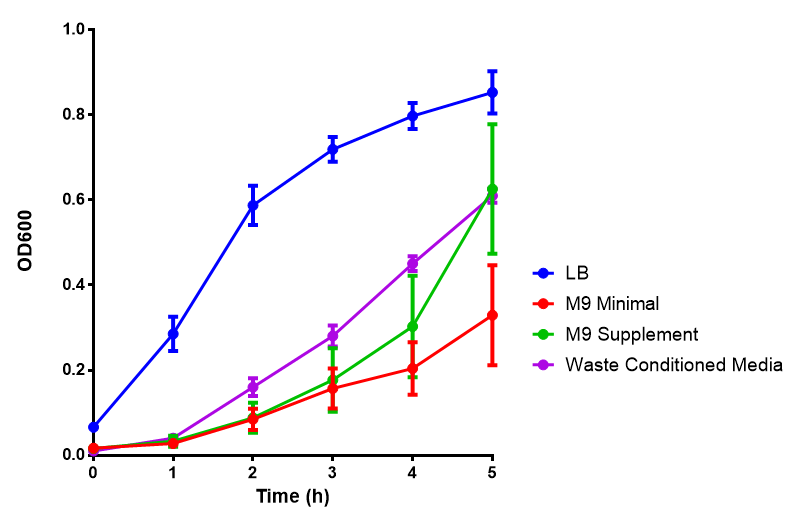 Figure 1: The medias used for our growth assays using the MG1655 strain. phaCAB is an operon involved in synthesis of P3HB while EV represents an empty vector. SR refers to stress response cells containing a red pigment. M9M is minimal media, while M9S is supplemented minimal media, as described here. WCM refers to waste conditioned media, which is made from sterile filtrated SRF waste, see here. Error bars are SEM, n=4. |
Waste Assays
These assays were designed to test whether our chassis could grow either directly with waste and/or in waste conditioned media. Waste conditioned media is a filter sterilised version of the waste media and was designed for several reasons; we were unsure whether mixed waste would be toxic to Ecoli and hence a less concentrated version may be more suitable and secondly large chunks of waste would prevent accurate OD600 measurements and therefore we decided to filter out the largest chunks.
Waste media
We can see from this experiment whether our bacteria can grow solely on the mixed waste.
Conclusion MG1655 can survive solely on mixed waste.
Waste conditioned media
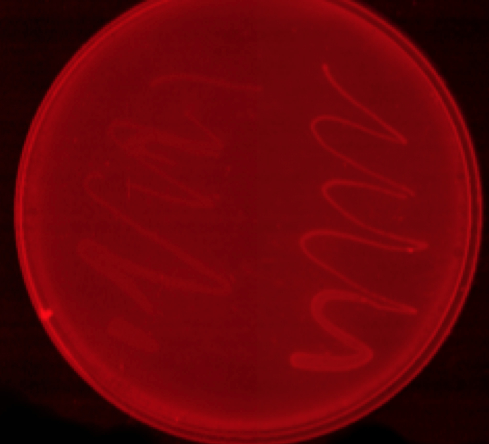
 mCherry stress biosensor (BBa_K639003) transformed MG1655 were grown with LB-WCM for 48 hours. 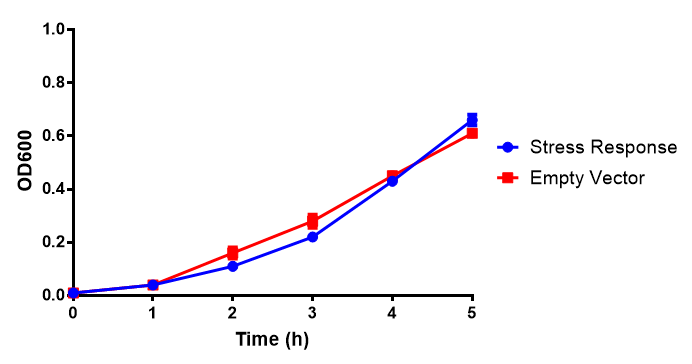 Figure 1: MG1655 in WCM with plasmids EV and SR. There is no growth inhibition when comparing the empty vector with the mCherry stress biosensor (BBa_K639003) vector in each media. WCM contains LB but trends in a simliar manner to M9S, suggesting that the chassis most likely due to toxins present within it from the mixed waste. Error bars are SEM, n=4. |
Characterisation of our Biobricks
Empty Vector Control
Characterisation of existing Biobricks
phaCAB biobrick characterisation
LB
M9 Minimal
M9 Supplemented
Stress biosensor characterisation BBa_K639003
IPTG induction assay
Originally we intended on using [http://parts.igem.org/Part:BBa_K639003 BBa_K639003] to detect whether our cells were stressed when placed in various toxic byproducts. However, as the data below shows, this biobrick is very leaky. As such, we are using the stress sensor as a marker for cell growth and also to show that the cells had been successfully transformed with the correct chloramphenicol resistance.
pBAD characterisation
Glucose
ANOVA analysis shows that...
Plastic Toxicity Assays
L-lactic Acid
Ethylene glycol
Reduced growth at 30oC likely due to decreased efficiency of MG1655 ethylene glycol break down enzymes. These enzymes (see UC Davis 2012) are endogenously expressed and detoxify Ethylene Glycol.
3-hydroxybutyrate (3HB)
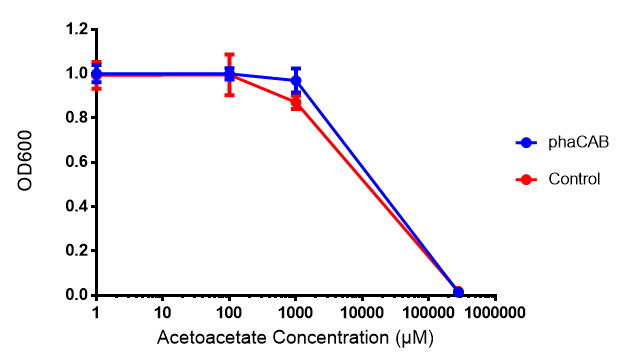
Acetoacetate
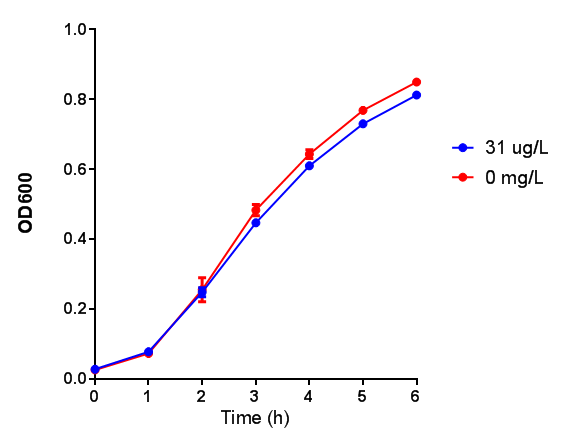
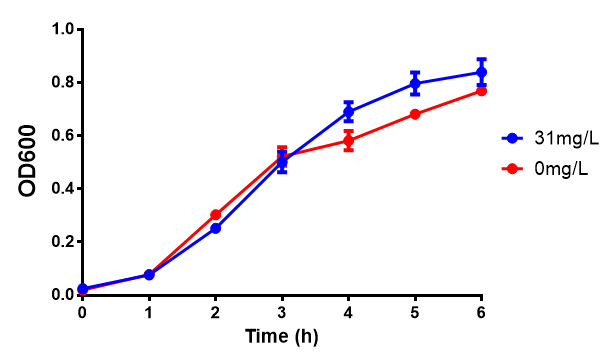
Poly(3-hydroxybutyrate) P(3HB)
Poly(lactic acid) (PLA)
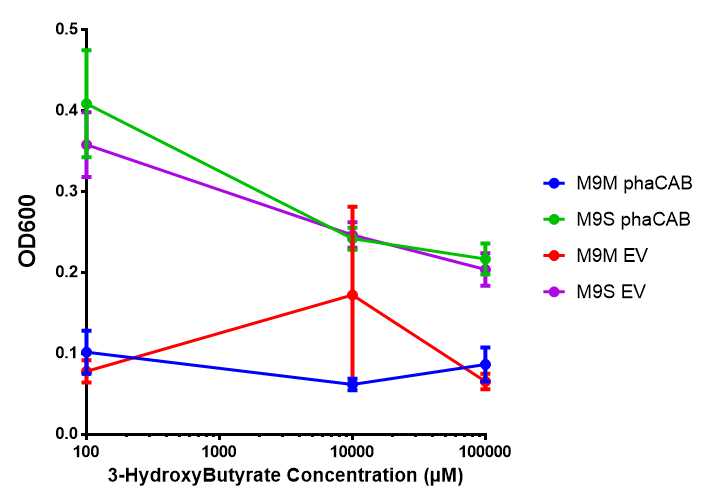
Sole carbon source
3HB
Acetoacetate
Western blots
Enzyme Kinetics
PHB production
Nile red staining
O/N cutures of MG1655 transformed with either control or phaCAB plasmid were spread onto LB-agar plates with 3% glucose and Nile red staining. The staining indicates the production of P3HB.
Purification of P3HB
 "
"