Team:NYMU-Taipei/Project/Inhibition/Prohibition
From 2013.igem.org
JosephHuang (Talk | contribs) |
|||
| (59 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:NYMU-Taipei/Header}} | {{:Team:NYMU-Taipei/Header}} | ||
| - | = | + | =Prohibiting Sprouting= |
| + | [[File:NYMU Taipei Nosema lifecycle.png|right|thumb|350px|ScientificBeekeeping.com http://scientificbeekeeping.com/the-nosema-twins-part-1/]] | ||
==Introduction== | ==Introduction== | ||
| - | + | In order to stop the invasion of ''Nosema ceranae'', the first step we take is to fully understand its life cycle which completes by bees. The picture beside indicates the relation between''Nosema ceranae'' stage and the place it grows. Through this picture, we can plan different ways to block the invasion of ''N. ceranae'' at each stage and also analyze the advantages and drawbacks of each ways, as well as to decide which stage is the best and most attainable for us to weaken the transmission ability of ''Nosema ceranae''. | |
| + | |||
==Background== | ==Background== | ||
===Stop it at the very begining!!!=== | ===Stop it at the very begining!!!=== | ||
| - | + | After ''N. ceranae'' invades epithelial cells of a bee's midgut, it will soon reproduce in two days and new generation appears. Since its lifecycle is so short that there is no time for biological methods to react and inhibit the development of the pathogen, we choose to inhibit it before entering bees' epithelial cells. When ''N. ceranae'''s spore moves to bees' midgut, it will start to germinate and develop a structure called polar filament. This structure will then bind to epithelial cells of bees' midgut and poke a hole at the membrane of an epithelial cell, causing the bee eventually dead. In order to inhibit polar filament's development, we find out that in other researches shows that there is a protein called PTP1, a protein on the surface of the polar filament, is highly relevant to the binding of the polar filament and epithelial cells. It is also highly believed that obstructing the function of PTP1 can stop the invasion of ''N. ceranae'' into the epithelial cell of the bee's midgut. So we decide to suppress the growth of the polar filament by obstruct the function of PTP1. In the research that is done by other scientist, they have found out that in the producing process of PTP1, a mannose polymer will bind to the protein. We initially decided to use mannosidase as our weapon to stop the invasion and even cloned the gene from ''E. coli'' ([http://parts.igem.org/Part:BBa_K1104100 BBa_K1104100]), however, we have considered the efficiency of how fast the enzyme can degrade the polymer, the long base pairs of mannosidase, also due to the difficulty to test its function, we changed our weapon into FimH, which is a mannose binding protein. In this way, at the appearance of the polar filament our protein can immediately block PTP1 and stop the invasion of ''N. ceranae''. | |
| - | + | ||
| - | [ | + | |
| - | + | ||
| + | ===FimH=== | ||
| + | FimH protein, a protein expressed in ''Escherichia coli'' K-12 substrain MG1655, has a mannose-binding domain from the 22nd to 179th amino acid. FimH protein is originally located on the tip of the fimbria of the ''E. coli'', which will adhere to the mannose on the host's cell membrane, allowing the bacteria to colonize various host tissues. In the sprouting of ''Nosema ceranae'', the O-glycosylation of polar tube protein 1 plays a role in the function of the polar tube, as this type of glycosylation can increase the stability of proteins protecting them from degradation and can create a “sticky” polar tube that adheres to mannose receptors. Therefore, we managed to use FimH to bind to the mannose polymer in order to stop the adhesion between the polar tube and the epithelial cell of the bee's midgut.<br> | ||
| + | |||
==Circuit Design== | ==Circuit Design== | ||
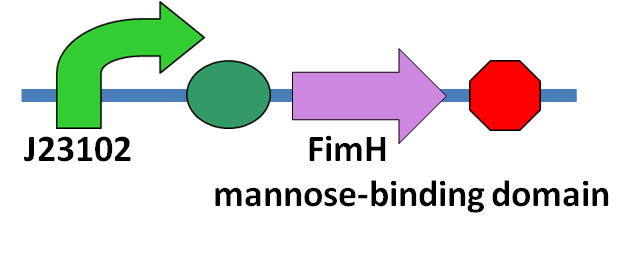
| - | + | [[File:NYMU_FimH circuit.png|200px|thumb|left|'''Circuit Design''' <br>A constitutive promoter, BBa_J23102, following the coding sequence of the FimH protein mannose-binding domain.]] | |
| - | + | Our main goal in this part is to stop the invasion of the ''N. ceranae'' from entering the epithelial cell, so the mannose binding protein we produce should constitutively produce in the bee's midgut. The circuit we construct will have a constitutive promoter, BBa_J23102, following the coding sequence of the FimH protein mannose-binding domain.The circuit is shown on the left. <br> | |
| + | <br><br> | ||
| + | ==Experiment== | ||
| + | To check whether FimH really works in our ''Bee. coli'', we fused RFP and FimH protein mannose-binding domain([http://parts.igem.org/Part:BBa_K1104101 BBa_K1104101]), so that when FimH binds to the mannose polymer, we can easily observe through seeing the red fluorescence.We will call this fusion protein RFP-FimH on the following text.<br> | ||
| + | [[File:NYMU_RFP-FimH circuit.png|right|thumb|200px|'''Circuit for RFP-FimH functional test''' <br>The circuit is constructed with a LacI regulated promoter and a RFP-FimH fusion protein coding sequence at the downstream.]] | ||
| + | ===Circuit Design=== | ||
| + | <br> | ||
| + | Through blunt-end ligation, we insert the 22nd to 179th amino acid of FimH protein gene in front the stop codon of the RFP coding sequence, in order to form a protein fusion. The promoter for this circuit is the LacI regulated promoter.We use this promoter as a constitutive promoter, since there is no LacI protein expressed in the bacteria. The picture on the right is the circuit design for our experiment.([http://parts.igem.org/Part:BBa_K1104102 BBa_K1104102]) <br> | ||
| + | [[File:NYMU RFP-FimH(XP).png|left|thumb|300px|'''Check for the Length of the RFP-FimH''' <br>The electrophoresis gel shows the result after the digestion. The central well is the 100 bp marker.The two well on both sides of the marker is the digestion sample of RFP-FimH from different colony. The lower band is the part with the promoter and the RFP-FimH coding sequence, which has the length of 1,485 nt. The upper band is the backbone, pSB1A2, we use.]] | ||
| + | [[File:NYMU_RFP-FimH cell.jpg|right|thumb|200px|'''Fluorescence Check''' <br>Both eppendorf are the liquid culture of the ''E. coli'' with RFP-FimH coding sequence cloned. We can obviously see the red light fluorescence emitted, showing that the fusion protein have successfully been produced and the RFP is still well-function.]] | ||
| - | + | To proof that the protein fusion actually express in the ''E. coli'' we have done digestion on XbaI and PstI cutting site to proof that the length of our biobrick is right. So there is only the coding sequence of RFP-FimH instead of RFP only. The picture on the left is the result of the electrophoresis after the digestion. Secondly, through observing the fluorescence emitted by the ''E. coli'' we can be sure that we have produced the protein that we want, which is the RFP-FimH fusion protein. The result is as the picture on the right.<br> | |
| + | |||
| + | |||
| - | === | + | <br><br><br> |
| - | + | ===RFP-FimH functional test=== | |
| + | After cloning RFP-FimH gene into ''E. coli'', it is a must to examine whether our part is properly combined and well-functional. Therefore we decide to extract RFP-FimH fusion protein from the cytoplasm of the ''E. coli'' and test its function by observing whether it will bind to the mannose. We originally want to test directly on the ''N. ceranae'' polar tube, however, we found it hard to cultivate under no matter sugar water or mosquito cell line, so we changed our material into ''E. coli'' to be the material that carries the mannose to conduct the functional test. | ||
====Experimental process is listed below==== | ====Experimental process is listed below==== | ||
| - | 1. | + | <br> |
| - | 2. | + | 1.Sonicate the ''E. coli'' that can produce RFP-FimH fusion protein <br> |
| - | 3. | + | 2.Centrifuge under 13200rpm for 1 min. and keep the supernatant <br> |
| - | 4. | + | 3.Cultivate ''Escherichia coli'' K-12 substr MG1655 for 2 hours <br> |
| - | 5. | + | 4.Take 100λ of liquid <br> |
| - | 6. | + | 5.Centrifuge under 13200rpm for 2 min.<br> |
| - | + | 6.Discard the supernatant and add in 100λ of the protein extraction <br> | |
| - | + | 7.Fully mix the ''E. coli'' with the protein extraction <br> | |
| - | + | 8.Put the mixture under 4℃ for 30 min. <br> | |
| - | + | 9.Centrifuge the mixture under 13200rpm for 1.5 min. <br> | |
| - | {| | + | 10.Observe the pellet under the fluorescence microscope and see if there is fluorescence emitted <br><br> |
| + | experiment group: | ||
| + | {|class="wikitable" | ||
|- | |- | ||
| - | + | ! | |
| - | | | + | |control group |
| - | + | |experimental group | |
|- | |- | ||
| - | | | + | !protein added |
| + | |RFP | ||
| + | |RFP-FimH | ||
|} | |} | ||
| - | |||
| - | |||
| + | [[File:NYMU_RFP-FimH_function.jpg|right|thumb|300px|'''RFP-FimH Functional Test Result''' <br>The eppendorf on the right is the control group with RFP protein added in the ''E. coli'' and 1x PBS mix. On the left is the experimental group with RFP-FimH fusion protein added. We can see that the eppendorf on the left shows a red color at the pellet, while it shows white on the right eppendorf. This indicates that the fusion protein we produce can actually binds to the mannose, which is the target that we want to block on ''N. ceranae''.]] | ||
| + | ====Experiment data==== | ||
| + | We anticipate that the group adding the RFP-FimH will express red fluorescence and the group adding RFP will not express red fluorescence in the pellet. Since the protein with FimH will bind to the mannose on the ''E. coli'', so it will express red fluorescence. The picture on the right shows that the group with FimH has a red fluorescence. Which means RFP-FimH that we cloned really works, and can hopefully block the mannose polymer on the PTP1 and stop the invasion of ''N. ceranae'' into the epithelial cell. | ||
| + | <br><br><br><br><br> | ||
==Reference== | ==Reference== | ||
| - | 1:[http://ecocyc.org/ECOLI/NEW-IMAGE?type=ENZYME&object= | + | <br> |
| - | + | 1. EcoCyc: Encyclopedia of Escherichia coli K-12 Genes and Metabolism [http://ecocyc.org/ECOLI/NEW-IMAGE?type=ENZYME&object=EG10315-MONOMER#/ FimH] <br> | |
| - | 2 | + | 2. Xu, Yanji, Takvorian, Peter M., Cali, Ann, Orr, George, & Weiss, Louis M. (n.d.). Glycosylation of the Major Polar Tube Protein of Encephalitozoon hellem, a Microsporidian Parasite That Infects Humans. American Society for Microbiology. <br> |
| - | + | 3. Xu, Yanji, Takvorian, Peter M., Cali, Ann, Orr, George, & Weiss, Louis M. (n.d.). ''Glycosylation of the Major Polar Tube Protein of Encephalitozoon hellem, a Microsporidian Parasite That Infects Humans.'' American Society for Microbiology. <br> | |
| - | 3 | + | 4. XU, Y., & WEISS, L. (August 01, 2005). The microsporidian polar tube: A highly specialised invasion organelle. ''International Journal for Parasitology'', 35, 9, 941-953. <br> |
| - | + | 5. Gisder, Sebastian, Hedtke, Kati, Möckel, Nadine, Frielitz, Marie-Charlotte, Linde, Andreas, & Genersch, Elke. (n.d.). Five-Year Cohort Study of Nosema spp. in Germany: Does Climate Shape Virulence and Assertiveness of Nosema ceranae?▿ †. American Society for Microbiology (ASM. <br> | |
| - | + | 6. Development of insect cell lines for the growth of Deformed Wing Virus and N. ceranae , http://beediseases.org.uk/sub-projects/cell-lines, 2013/10/29 <br> | |
| + | 7. Nosema ceranae The Inside Story, Bee Health, February 20, 2012 <br> | ||
| + | 8. Bouzahzah, B., & Weiss, L. M. (January 01, 2010). Glycosylation of the major polar tube protein of Encephalitozoon cuniculi. Parasitology Research, 107, 3, 761-4. <br> | ||
{{:Team:NYMU-Taipei/Footer}} | {{:Team:NYMU-Taipei/Footer}} | ||
Latest revision as of 03:47, 29 October 2013


Contents |
Prohibiting Sprouting
Introduction
In order to stop the invasion of Nosema ceranae, the first step we take is to fully understand its life cycle which completes by bees. The picture beside indicates the relation betweenNosema ceranae stage and the place it grows. Through this picture, we can plan different ways to block the invasion of N. ceranae at each stage and also analyze the advantages and drawbacks of each ways, as well as to decide which stage is the best and most attainable for us to weaken the transmission ability of Nosema ceranae.
Background
Stop it at the very begining!!!
After N. ceranae invades epithelial cells of a bee's midgut, it will soon reproduce in two days and new generation appears. Since its lifecycle is so short that there is no time for biological methods to react and inhibit the development of the pathogen, we choose to inhibit it before entering bees' epithelial cells. When N. ceranae's spore moves to bees' midgut, it will start to germinate and develop a structure called polar filament. This structure will then bind to epithelial cells of bees' midgut and poke a hole at the membrane of an epithelial cell, causing the bee eventually dead. In order to inhibit polar filament's development, we find out that in other researches shows that there is a protein called PTP1, a protein on the surface of the polar filament, is highly relevant to the binding of the polar filament and epithelial cells. It is also highly believed that obstructing the function of PTP1 can stop the invasion of N. ceranae into the epithelial cell of the bee's midgut. So we decide to suppress the growth of the polar filament by obstruct the function of PTP1. In the research that is done by other scientist, they have found out that in the producing process of PTP1, a mannose polymer will bind to the protein. We initially decided to use mannosidase as our weapon to stop the invasion and even cloned the gene from E. coli ([http://parts.igem.org/Part:BBa_K1104100 BBa_K1104100]), however, we have considered the efficiency of how fast the enzyme can degrade the polymer, the long base pairs of mannosidase, also due to the difficulty to test its function, we changed our weapon into FimH, which is a mannose binding protein. In this way, at the appearance of the polar filament our protein can immediately block PTP1 and stop the invasion of N. ceranae.
FimH
FimH protein, a protein expressed in Escherichia coli K-12 substrain MG1655, has a mannose-binding domain from the 22nd to 179th amino acid. FimH protein is originally located on the tip of the fimbria of the E. coli, which will adhere to the mannose on the host's cell membrane, allowing the bacteria to colonize various host tissues. In the sprouting of Nosema ceranae, the O-glycosylation of polar tube protein 1 plays a role in the function of the polar tube, as this type of glycosylation can increase the stability of proteins protecting them from degradation and can create a “sticky” polar tube that adheres to mannose receptors. Therefore, we managed to use FimH to bind to the mannose polymer in order to stop the adhesion between the polar tube and the epithelial cell of the bee's midgut.
Circuit Design
Our main goal in this part is to stop the invasion of the N. ceranae from entering the epithelial cell, so the mannose binding protein we produce should constitutively produce in the bee's midgut. The circuit we construct will have a constitutive promoter, BBa_J23102, following the coding sequence of the FimH protein mannose-binding domain.The circuit is shown on the left.
Experiment
To check whether FimH really works in our Bee. coli, we fused RFP and FimH protein mannose-binding domain([http://parts.igem.org/Part:BBa_K1104101 BBa_K1104101]), so that when FimH binds to the mannose polymer, we can easily observe through seeing the red fluorescence.We will call this fusion protein RFP-FimH on the following text.
Circuit Design
Through blunt-end ligation, we insert the 22nd to 179th amino acid of FimH protein gene in front the stop codon of the RFP coding sequence, in order to form a protein fusion. The promoter for this circuit is the LacI regulated promoter.We use this promoter as a constitutive promoter, since there is no LacI protein expressed in the bacteria. The picture on the right is the circuit design for our experiment.([http://parts.igem.org/Part:BBa_K1104102 BBa_K1104102])
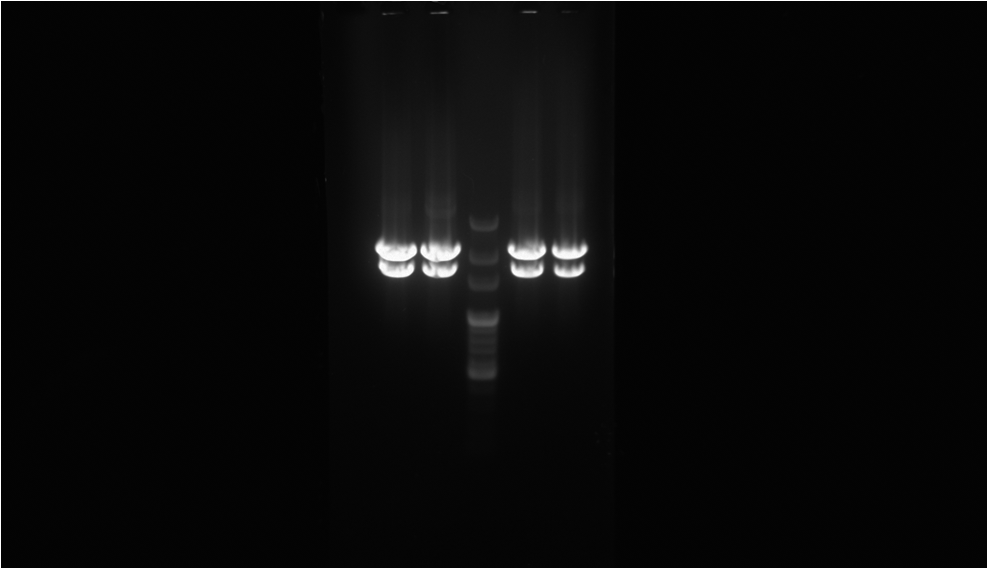
The electrophoresis gel shows the result after the digestion. The central well is the 100 bp marker.The two well on both sides of the marker is the digestion sample of RFP-FimH from different colony. The lower band is the part with the promoter and the RFP-FimH coding sequence, which has the length of 1,485 nt. The upper band is the backbone, pSB1A2, we use.
To proof that the protein fusion actually express in the E. coli we have done digestion on XbaI and PstI cutting site to proof that the length of our biobrick is right. So there is only the coding sequence of RFP-FimH instead of RFP only. The picture on the left is the result of the electrophoresis after the digestion. Secondly, through observing the fluorescence emitted by the E. coli we can be sure that we have produced the protein that we want, which is the RFP-FimH fusion protein. The result is as the picture on the right.
RFP-FimH functional test
After cloning RFP-FimH gene into E. coli, it is a must to examine whether our part is properly combined and well-functional. Therefore we decide to extract RFP-FimH fusion protein from the cytoplasm of the E. coli and test its function by observing whether it will bind to the mannose. We originally want to test directly on the N. ceranae polar tube, however, we found it hard to cultivate under no matter sugar water or mosquito cell line, so we changed our material into E. coli to be the material that carries the mannose to conduct the functional test.
Experimental process is listed below
1.Sonicate the E. coli that can produce RFP-FimH fusion protein
2.Centrifuge under 13200rpm for 1 min. and keep the supernatant
3.Cultivate Escherichia coli K-12 substr MG1655 for 2 hours
4.Take 100λ of liquid
5.Centrifuge under 13200rpm for 2 min.
6.Discard the supernatant and add in 100λ of the protein extraction
7.Fully mix the E. coli with the protein extraction
8.Put the mixture under 4℃ for 30 min.
9.Centrifuge the mixture under 13200rpm for 1.5 min.
10.Observe the pellet under the fluorescence microscope and see if there is fluorescence emitted
experiment group:
| control group | experimental group | |
| protein added | RFP | RFP-FimH |

The eppendorf on the right is the control group with RFP protein added in the E. coli and 1x PBS mix. On the left is the experimental group with RFP-FimH fusion protein added. We can see that the eppendorf on the left shows a red color at the pellet, while it shows white on the right eppendorf. This indicates that the fusion protein we produce can actually binds to the mannose, which is the target that we want to block on N. ceranae.
Experiment data
We anticipate that the group adding the RFP-FimH will express red fluorescence and the group adding RFP will not express red fluorescence in the pellet. Since the protein with FimH will bind to the mannose on the E. coli, so it will express red fluorescence. The picture on the right shows that the group with FimH has a red fluorescence. Which means RFP-FimH that we cloned really works, and can hopefully block the mannose polymer on the PTP1 and stop the invasion of N. ceranae into the epithelial cell.
Reference
1. EcoCyc: Encyclopedia of Escherichia coli K-12 Genes and Metabolism [http://ecocyc.org/ECOLI/NEW-IMAGE?type=ENZYME&object=EG10315-MONOMER#/ FimH]
2. Xu, Yanji, Takvorian, Peter M., Cali, Ann, Orr, George, & Weiss, Louis M. (n.d.). Glycosylation of the Major Polar Tube Protein of Encephalitozoon hellem, a Microsporidian Parasite That Infects Humans. American Society for Microbiology.
3. Xu, Yanji, Takvorian, Peter M., Cali, Ann, Orr, George, & Weiss, Louis M. (n.d.). Glycosylation of the Major Polar Tube Protein of Encephalitozoon hellem, a Microsporidian Parasite That Infects Humans. American Society for Microbiology.
4. XU, Y., & WEISS, L. (August 01, 2005). The microsporidian polar tube: A highly specialised invasion organelle. International Journal for Parasitology, 35, 9, 941-953.
5. Gisder, Sebastian, Hedtke, Kati, Möckel, Nadine, Frielitz, Marie-Charlotte, Linde, Andreas, & Genersch, Elke. (n.d.). Five-Year Cohort Study of Nosema spp. in Germany: Does Climate Shape Virulence and Assertiveness of Nosema ceranae?▿ †. American Society for Microbiology (ASM.
6. Development of insect cell lines for the growth of Deformed Wing Virus and N. ceranae , http://beediseases.org.uk/sub-projects/cell-lines, 2013/10/29
7. Nosema ceranae The Inside Story, Bee Health, February 20, 2012
8. Bouzahzah, B., & Weiss, L. M. (January 01, 2010). Glycosylation of the major polar tube protein of Encephalitozoon cuniculi. Parasitology Research, 107, 3, 761-4.
 "
"