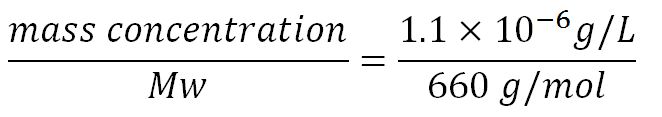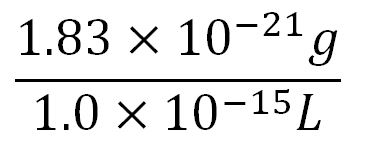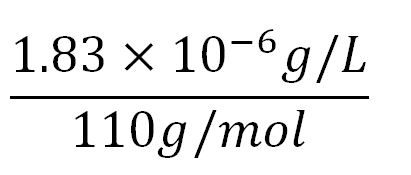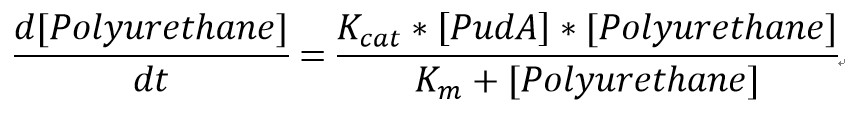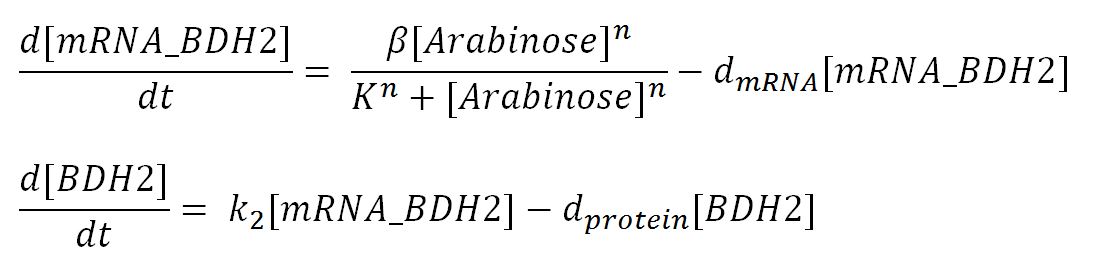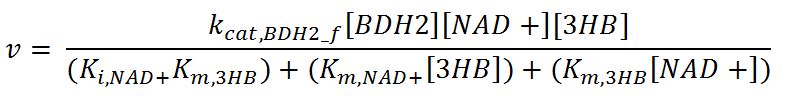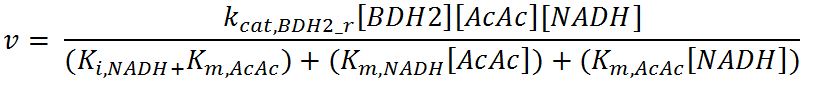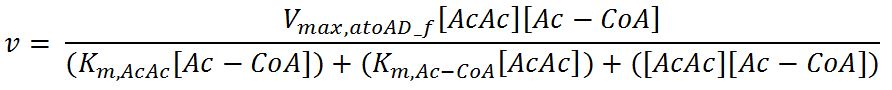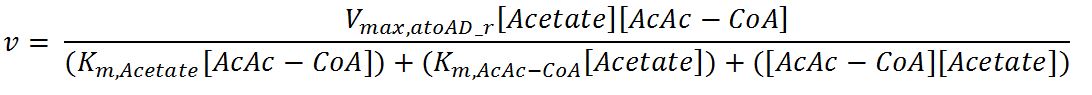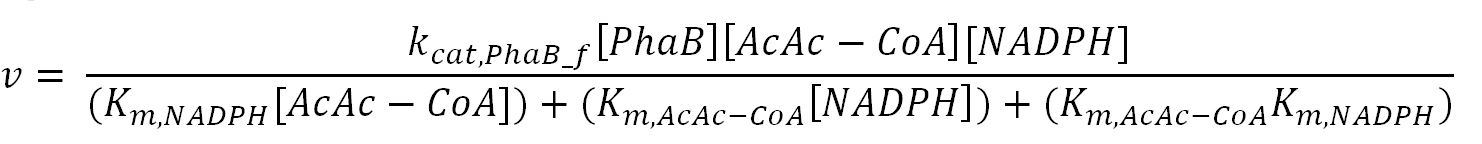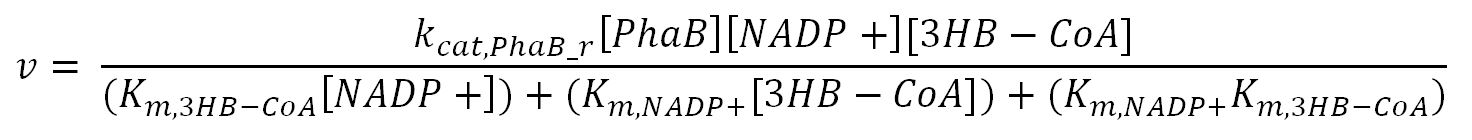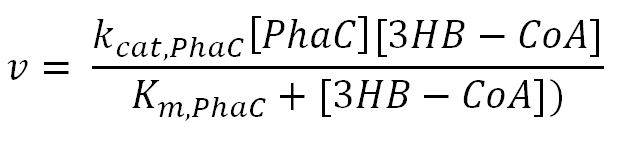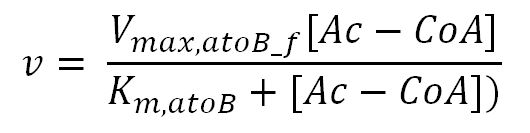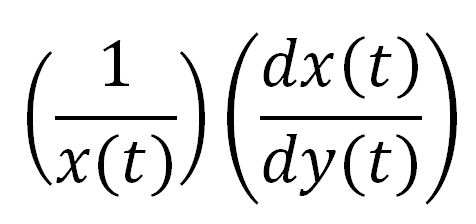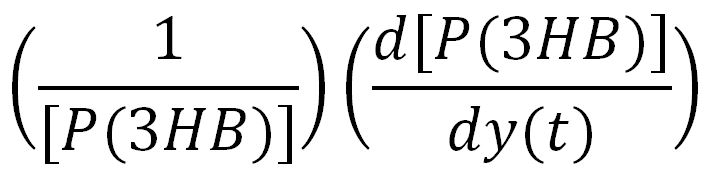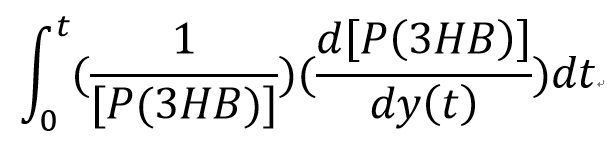Team:Imperial College/Waste Degradation: SRF
From 2013.igem.org
| Line 1,532: | Line 1,532: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | + | <p align="justify"> | |
| + | It should also be noted that acetate, acetoacetate and acetyl-CoA cannot be predicted by the metabolic model. Therefore, their initial concentrations are rough estimates (value similar to other metabolites) such that a reasonable P(3HB) output can be produced. | ||
<html> | <html> | ||
</div> | </div> | ||
Revision as of 02:53, 5 October 2013
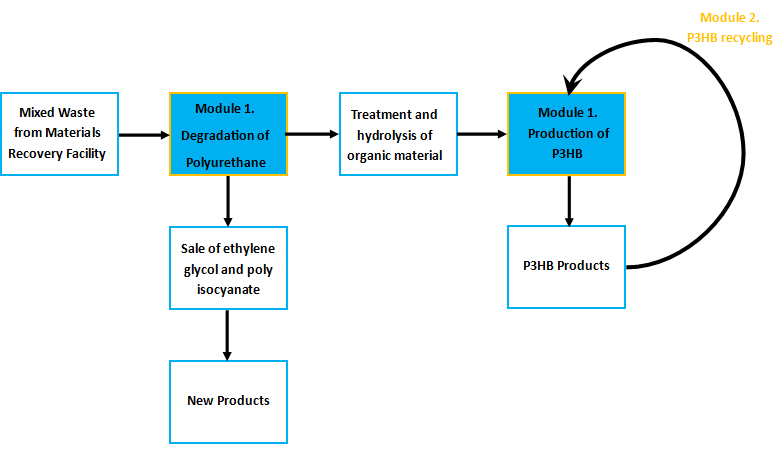
Module 1: Resource-full Waste
Non-recyclable waste is sourced from a recycling centre, placed in a bioreactor with our M.A.P.L.E system which degrades the waste and synthesises the bioplastic P(3HB). Click the tabs to find out more
Overview
Specifications
Design
Modelling
Assembly
Testing & results
Future work
Waste from recovery facilities is a mixture of plastics and cellulosic rich biomass such as fibres and wood. The Resource-full waste module utilises the large variety of materials organisms can naturally degrade to recycle the waste and to provide food for the production of the bioplastic poly-3-hydroxybutyrate.
Separating polyurethane and selling its breakdown products
Certain enzymes have the ability to degrade plastics as a result of their evolutionary history, dealing with tough and highly variable substrates such as lignin. Various enzymes have been biobricked to deal with most of the major petrochemical plastics, except for polyurethane. Polyurethane is one of the plastic constituents of this mixed waste. We have designed biobricks containing naturally occurring polyurethane esterases (also known as PUR esterases) to break polyurethane down into its constituent chemicals of ethylene glycol and isocyanate. These valuable chemicals will be separated using specialised filters and sold back to industry.
Using the residual material as a substrate for P3HB production
The remaining organic material will be first hydrolysed to release sugars for the bacteria to live upon. This is a current industrial pre-treatment technique to allow the fermentation of ligno-cellulosic biomass.
P3HB is produced by organisms containing the phaCAB operon, originally found in soil bacteria such as Ralstonia eutropha. The operon contains three genes which are members of the pathway required to make P3HB. P3HB is used as a storage molecule in the organisms which produce it. They do this when nutrients are limiting but when there is a plentiful carbon source.
P3HB is produced commercially by several companies. One of the crucial factors in whether production is economical or not is the efficiency of bioplastic production. We have hugely increased the efficiency of P3HB production; an important step in making our system feasible.
References:
- Harding KG, Dennis JS, von Blottnitz H, Harrison STL. Environmental analysis of plastic production processes: Comparing petroleum-based polypropylene and polyethylene with biologically-based poly-beta-hydroxybutyric acid using life cycle analysis. J Biotechnol 2007 MAY 31;130(1):57-66.
- Kim S, Dale BE. Energy and Greenhouse Gas Profiles of Polyhydroxybutyrates Derived from Corn Grain: A Life Cycle Perspective. Environ Sci Technol 2008 OCT 15;42(20):7690-7695.
- Kendall A. A life cycle assessment of biopolymer production from material recovery facility residuals. Resources, Conservation and Recycling 61 (2012) 69– 74 conomou A. Following the leader: bacterial protein export through the Sec pathway. Trends in microbiology 1999;7(8) 315-320.
- Chul-Hyung Kang, Ki-Hoon Oh, Mi-Hwa Lee, Tae-Kwang Oh, Bong Hee Kim, Jung- Hoon Yoon. A novel family VII esterase with industrial potential from compost metagenomic libraryMicrobial Cell Factories 2011, 10:41 doi:10.1186/1475-2859-10-41
- Da Almeida, Alejandra. Effect of the granule associated protein phasin (PhaP) on cell growth and poly(3-hydroxybutyrate) (PHB) accumulation from glycerol in bioreactor cultures of recombinant E. coli. Volume 131, Issue 2, Supplement, September 2007, Pages S167.
Specification
1. The bacteria should survive and grow in mixed waste
In order for the bacteria to produce bioplastic from the mixed waste they first need to be able to use it as a carbon source
2. The bacteria should secrete a functional polyurethane esterase
To recover the resources from polyurethane in the mixed waste, the enzymes should secrete in an active form.
3. Our Bacteria should be able to tolerate mixed waste degradation products, such as Ethylene Glycol
The products of Polyurethane degradation are toxic and so any bacteria growing with them must be able to survive to a concentration which will allow economical production of ethylene glycol. We have chosen a strain of E.coli-MG1655, which is resistant to ethylene glycol toxicity.
4. The bacteria should produce P3HB
5. The bacteria should produce P3HB from the mixed waste
PUR Degradation: Pathway ▼

Lignin degrading microbial enzymes are capable of degrading plastics. Some previous iGEM teams have exploited this ability. We have built on this work and extended the plastic degradation capabilities of the synthetic biology community by improving PUR degradation in particular as this has not been successfully achieved before. We have identified 5 PUR-esterase enzymes from the literature that are capable of catalyzing the below reaction.

We have synthesised all of the genes in the below table and are testing them for expression in E.coli, secretion, activity and PUR degradation capabilities. Our ultimate design is to be able to control the relative levels of different enzymes in a waste degrading bio-reactor in order to adjust it to the composition of waste. Therefore we designed the expression constructs accordingly. Our models predict the degradation rate of PUR at the bioreactor scale.
PUR Degradation: BioBrick Designs ▼
| enzyme | source organism | biobrick | reference |
|---|---|---|---|
| EstCS2 | uncultured unknown bacterium (GU256649.1) | BBa_K1149002 | Kang et.al 2011 |
| pueA | Pseudomonas chlororaphis | BBa_K1149003 | Stern et al., 2000 |
| pueB | Pseudomonas chlororaphis | BBa_K1149004 | Howard et al., 2001 |
| pudA | Comamonas acidovorans | BBa_K1149005 | Allen et al. 1999 |
| pulA | Pseudomonas fluorescens | BBa_K1149006 | Vega et al., 1999 |

We considered the safety aspects of using the PudA enzyme, since its sequences were originally from a Risk Group 2 (RG2) organism. We submitted our safety information to iGEM for review. Safety forms were approved on October 2nd, 2013 by the iGEM Safety Committee. We did not use the organism only the part itself and the final sequences were codon optimised for expression in E. coli, which included the elimination of the forbidden restriction sites.
Bioplastic Synthesis: Pathway ▼
Poly-3-hydroxybutyrate(P3HB) is a bioplastic which is naturally produced inside bacteria such as Ralstonia eutropha, where it accumulates as globules inside the cell. In its native bacteria it is produced as an energy store(1) but it is as a plastic that it interests researchers and industrialists. In our system, the bacteria will take up organic molecules from waste and use them for bioplastic production.

We have made P3HB in E.coli, transferring three genes, naturally found in Ralstonia eutropha into E.coli MG1655. These encode the three enzymes necessary for P3HB production; polyhydroxyalkanoate synthase(phaC), 3-ketothiolase(phaA) and acetoacetyl coenzyme A reductase(phaB). These are encoded by the phaCAB operon. We have altered the expression of these three genes to maximise the production of P3HB as high yields are required for it to be economically viable.
3-ketothiolase:

acetoacetyl coenzyme A reductase:

polyhydroxyalkanoate synthase:

Bioplastic Synthesis: Biobrick Designs ▼
We used the phaCAB biobrick [http://parts.igem.org/Part:BBa_K934001 BBa_K934001] to produce P3HB which contains the native operon of Ralstonia eutropha. Our system will be used at industrial scales. This requires high yields to be economically viable. We created P3HB synthesis models to inform our experimental attempts to increase P3HB yield. We were especially interested to find out if up or down-regulation of any of the enzymes involved in the biosynthetic pathway could increase yields. To analyse this, we needed to know the reactions in the metabolic pathway and also the relative parameters of the enzymes and their dynamic relationships in order to identify bottlenecks in the flux of metabolites. The results from the metabolic model suggest that the amount of phaB is especially critical and increasing it`s level should give us more PHB. (Please see our modelling section for details.) To do this, we designed constructs with stronger promoters to up-regulate the expression of the operon.
We designed and constructed two new constructs for increased expression of the operon (including phaB). The first construct uses the constitutive promoter [http://parts.igem.org/Part:BBa_J23104 J23104] and the [http://parts.igem.org/Part:BBa_B0034 RBS 0034] with a following scar site TACTAGAG in front of the ATG of the phaC gene. The second construct was designed as a hybrid promoter incorporating BBa_J23104 and the original promoter first, followed by the native promoter and RBS due to recent results of high expression from a hybrid promoter[http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0056321]. The region containing the native promoter and RBS is 352 nucleotides long and might contain important regulatory elements and therefore we were interested to test the hybrid construct in comparison to both the native and constitutive promoter configurations.

Chassis Design: Tolerance to Waste ▼
We chose the MG1655 E.coli strain as chassis because it constitutively expresses genes that make it resistant to toxic waste degradation products, such as Ethylene glycol. The [http://parts.igem.org/Part:BBa_K892010 AldA] and [http://parts.igem.org/Part:BBa_K892009 FucO] genes have an important role in decreasing the toxic effects of Ethylene-glycol by converting it into Glycolaldehyde which is a link to the cell`s central metabloism. We have received these genes from the registry and future work could be to express these in Bacillus or cellulose degrading organisms to make them tolerant as well.
Enzyme Secretion Strategy ▼
E.coli is commonly used as a chassis in innovative iGEM projects that aim to prove a concept and make the case for a novel function in a biologically engineered machine. In our case, we aim to degrade and synthesise plastic and the degradation part of our system needs to be extracellular. There are many strategies for secretion. On the following page, you can read about these and find out why we chose the pelB tag for all our degradation enzymes and why we are using osmY in our future work.
Alternative strategies for protein secretion in E.coli:
N terminal secretion-signal peptides are recognised by the sec pathway . The pathway transports proteins to the periplasm and additional mechanisms are usually necessary to further export the protein to the extracellular space. An approach for using this pathway is the addition of an N terminal signal peptide to the target protein which can obtain as high as 90% efficiency in secretion to the extracellular space. Such signals are pelB and phoA tags which are available as biobricks. PelB can be cleaved off by pelB peptidase in the periplasm whereas phoA will anchor your protein to this localisation.
Fusion partners are endogenous proteins that are naturally secreted in E.coli and can be fused to the target protein. Some such proteins were demonstrated to be a powerful carriers of medically relevant human proteins in E.coli. The yebF and ompF proteins exit the cell via the sec pathway and use additional mechanisms for leaving the periplasm where they interact with outer-membrane porins for extracellular secretion. The osmY is available as a Biobrick and we have submitted ompF this year.
The porin proteins such as ompF and ompA, can be used as fusion partners too and anchor proteins to the outer surface of E.coli.
ABC transporters use ATP for transport of specific proteins across the bacterial membrane. The ABC transporter of Erwinia chrysanthemi can be expressed in E.coli and export proteins that contain the LARD1 domain as a C terminal fusion. The system is biobricked in two separate plasmids (transporter, LARD1) with different antibiotic resistance and it is important to use both at the same time.
The [http://parts.igem.org/Protein_domains/Localization page] in the registry that lists localisation tags] is incomplete and contains eukaryotic and prokaryotic localisation signals mixed together. We have therefore put together the tables below for you to help choose an E.coli secretion strategy that is suitable for your project.
</p>
Extracellular Secretion in E.coli:
| Description | Biobrick | Team | |
|---|---|---|---|
| pelB | type II secretion signal peptide, cleaved off in periplasm | BBa_K208004 BBa_K1149022 BBa_K208004 BBa_J32015 | lots eg. Imperial 2013 |
| yebF | large fusion-protein, Sec-pathway and additional mechanism | BBa_K1149001 | Imperial 2013 |
| osmY | large fusion-protein, Sec-pathway and additional mechanism | BBa_K892008 | Washington 2012 |
| LARD 1 | Lipase ABC transporter recognition domain | BBa_K258001 | METU-GEne 2009 |
Outer Membrane Anchors and Periplasmic Expression in E.coli:
| Description | Biobrick | Team | |
|---|---|---|---|
| ompF | porin protein that can be fused to a protein, anchors to outer membrane | BBa_K864204 | Uppsala 2012 |
| ompA | porin protein that can be fused to a protein, anchors to outer membrane | BBa_K103006 | Wasraw 2008 |
| INP | short domain,anchors in periplasm | BBa_K523013 (INP-YFP) BBa_K632002 (INP-Silicatein) | Minnesota 2011 Edinburgh 2011 |
| torA | anchors in periplasm | Dundee 2013 | |
| phoA | tag for sec pathway, anchors to OM | BBa_K808028 | Darmstadt 2011 |
We chose the pelB secretion tag as it has been demonstrated to work in many cases with sometimes as high as 90% transport efficiency. The pelB has been used in iGEM project for many years and is part of 50+ constructs. The UC-Davis team last year used it to secrete LC-Cutinase, a PET plastic degrading enzyme ( BBa_K936013) successfully which is somewhat similar to our plastic degradation enzymes.
Assembly methods and toolkit:
An advantage of pelB is that it is relatively short (only 66 BP) and so we could get our gene synthesised with the tag. A problem with standard biobrick assembly is that the scar site contains a STOP codon and therefore [http://parts.igem.org/Protein_domains alternative strategies are needed]. One of the ways around this is to use Infusion/Gibson assembly which we also tried and you can find instructions under our protocols section. We have constructed a biobrick where LARD1 is after a constitutive promoter and RBS in order to facilitate quicker cloning with less assembly steps, [http://parts.igem.org/Part:BBa_K1149021 BBa_K1149021].
Unfortunately, the pelB tag did not prove to be the best strategy for secreting our degradation enzymes in the MG1655 strain. Therefore, we are changing the strategy to use osmY.
References
- Economou A. Following the leader: bacterial protein export through the Sec pathway. Trends in microbiology 1999;7(8) 315-320.
- Thanassi DG, Hultgren SJ. Multiple pathways allow protein secretion across the bacterial outer membrane. Current opinion in cell biology 2000;12(4) 420-430.
- Sletta H, Tondervik A, Hakvag S, Aune TEV, Nedal A, Aune R, et al. The presence of N-terminal secretion signal sequences leads to strong stimulation of the total expression levels of three tested medically important proteins during high-cell-density cultivations of E.coli. Applied and Environmental Microbiology 2007;73(3) 906-912.
- Qian Z, Xia X, Choi JH, Lee SY. Proteome-based identification of fusion partner for high-level extracellular production of recombinant proteins in E.coli. Biotechnology and bioengineering 2008;101(3) 587-601.
- Prehna G, Zhang G, Gong X, Duszyk M, Okon M, McIntosh LP, et al. A Protein Export Pathway Involving Escherichia coil Porins. Structure 2012;20(7) 1154-1166.
- Binet R, Letoffe S, Ghigo JM, Delepelaire P, Wandersman C. Protein secretion by Gram-negative bacterial ABC exporters - A review. Gene 1997;192(1) 7-11.
- Chung CW, You J, Kim K, Moon Y, Kim H, Ahn JH. Export of recombinant proteins in E.coli using ABC transporter with an attached lipase ABC transporter recognition domain (LARD). Microbial Cell Factories 2009;8 11.
- Francetic O, Pugsley AP. Towards the identification of type II secretion signals in a nonacylated variant of pullulanase from Klebsiella oxytoca. Journal of Bacteriology 2005;187(20) 7045-7055.
- Fu LL, Xu ZR, Li WF, Shuai JB, Lu P, hu CXH. Protein secretion pathways in Bacillus subtilis: Implication for optimization of heterologous protein secretion. Biotechnology Advances 2007;25(1) 1-12.
- Cho HY, Yukawa H, Inui M, Doi RH, Wong SL. Production of minicellulosomes from Clostridium cellulovorans in Bacillus subtilis WB800. Applied and Environmental Microbiology 2004;70(9) 5704-5707.
- Zhang XZ, Cui ZL, Hong Q, Li SP. High-level expression and secretion of methyl parathion hydrolase in Bacillus subtilis WB800. Applied and Environmental Microbiology 2005;71(7) 4101-4103.
Safety ▼
Safety issues regarding our chassis can be found here.
Modelling
What we learnt from the PUR degradation model:
- The model assisted the design of enzyme assays, in particular:
- The toxicity of the PUR degradation product, Ethylene glycol was taken into account in the modelling and it successfully determined the maximum tolerance of the system.
- Ethylene glycol toxicity assays were carried out in the wet lab and were designed according to the results from our model.
What we learnt from the P(3HB) synthesis model:
- Scanning concentration of PhaB and sensitivity analysis showed that increasing the concentration of PhaB would increase the production rate of P(3HB) in our engineered 'E.coli'.
- Simulations showed that the constitutive promoter J23104 could lead to a higher expression of PhaB than the original promoter used. As a result the Wet lab team designed and built [http://parts.igem.org/Part:BBa_K1149051 BBa_K1149051], which significantly increased bioplastic production.
- Results from the metabolic model suggested that the synthesis of P(3HB) would be accompanied by a drop of ATP over time and hence allowed us to identify potential limitations of our engineered system. This is important for the Industrial Implementation of our system.
1.0 Polyurethane (PUR) degradation model ▼
Introduction
The efficiencies for Polyurethane (PUR) degradation and ethylene glycol production are important for the performance of MAPLE system. We built a mathematical and deterministic model that is based on MATLAB extension Simbiology for Polyurethane degradation. The model contains the kinetic property of degradation enzymes that is helpful for the design of assays. As we scaled up the initial concentrations of all substrates to meet the conditions for a bio-reactor, the model can provide preliminary simulations and predictions for the MAPLE system.
Design
Objective
Here are some specific objectives for the model to achieve:
1. The model should contain the gene expression model of the degradation enzymes because the enzyme concentration determines the rate of plastic degradation. In our case for PUR degradation, we used [http://parts.igem.org/Part:BBa_K206000 pBAD strong promoter K206000] for most enzymes. We built the gene expression model based on inducible pBAD promoter, which gene expression rate can be regulated by inducer concentration.
2. The model should show the efficiency of the enzyme secretion to the culture from the cells. It's also important because the enzyme concentration in the culture depends on it. Here we used pelB secretion tag for most enzymes in order to achieve a high efficiency.
3. The model basically predict how long will take to degrade a known concentration of soluble polyurethane. It is assumed that the enzyme in our assays has the same kinetic properties as the enzyme used in the literature. The model can suggest a suitable concentration of the plastic to use in order to get good results from the assays.
4. It is known that ethylene glycol is toxic to E.coli. However, it has no clear effect on the growth of our MG1655 strain when the concentration of ethylene glycol is below 200mM. Therefore, the model should suggest a safe range of PUR concentration to avoid a high concentration (>200mM) of ethylene glycol produced.
The Model
Polyurethane (PUR) degradation involves 5 different degradation enzymes:
| enzyme | source organism | biobrick | reference |
|---|---|---|---|
| EstCS2 | uncultured unknown bacterium (GU256649.1) | [http://parts.igem.org/Part:BBa_K1149002 BBa_K1149002] | [http://www.microbialcellfactories.com/content/10/1/41 Kang et.al 2011] |
| pueA | Pseudomonas chlororaphis | [http://parts.igem.org/Part:BBa_K1149003 BBa_K1149003] | [http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2000.tb09056.x/pdf Stern et al., 2000] |
| pueB | Pseudomonas chlororaphis | [http://parts.igem.org/Part:BBa_K1149004 BBa_K1149004] | [http://www.sciencedirect.com/science/article/pii/S0964830501000427 Howard et al., 2001] |
| pudA | Comamonas acidovorans | [http://parts.igem.org/Part:BBa_K1149005 BBa_K1149005] | [http://www.sciencedirect.com/science/article/pii/S0964830598000663 Allen et al. 1999] |
| pulA | Pseudomonas fluorescens | [http://parts.igem.org/Part:BBa_K1149006 BBa_K1149006] | [http://www.sciencedirect.com/science/article/pii/S0964830598000687 Vega et al., 1999] |
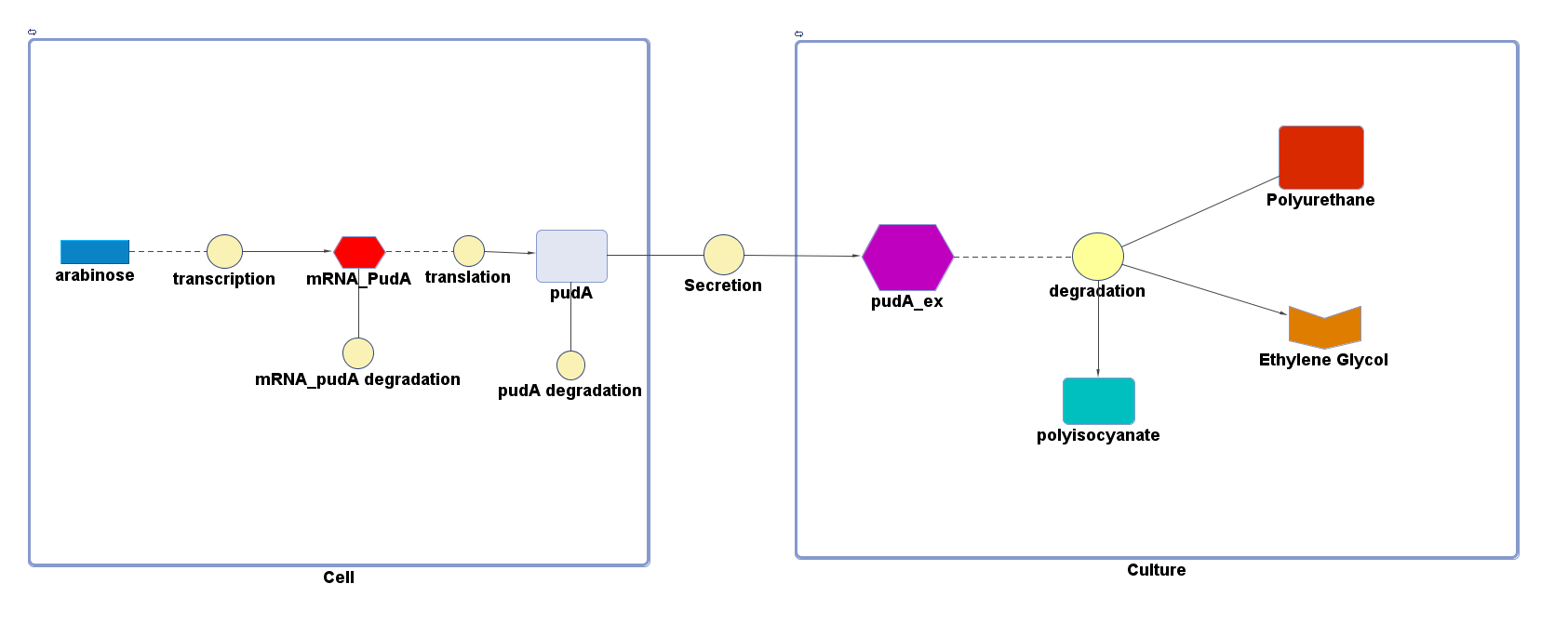
However, 4 of the 5 enzymes are not well characterised before, so we could't find enough kinetic data from the literature. The only well-characterised PUR degradation enzyme PudA is used in the model as an illustration of all PUR degradation enzymes. The model will be more complete when the kinetic data of the other enzymes are defined. The finished PUR degradation model is shown as below:
There are two compartments which represents cells and the culture from left to right. The "cell" compartment contains the gene expression module whereas the "culture" compartment contains the degradation module. The "secretion" block that connects two compartments is the secretion module.
Parameters and assumptions
| Parameter | Description | Value | Units | Source | Assumptions/Notes |
|---|---|---|---|---|---|
| β | maximum rate of transcription | 0.015 | mM/min | Please see derivation 1 below. | Please see derivation 1 below. |
| K | Activation coefficient | 0.0031 | mM | [http://parts.igem.org/Part:BBa_K206000:Characterization] | Taking the "switch point" as the activation coefficient |
| dmRNA | mRNA degradation rate | 0.10 | 1/min | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC420366/pdf/07X.pdf] | Taking the value of mRNA half-life in E.coli strain MG1655 as 6.8min. rate = ln2/half-life = ln2/6.8 = 0.10 |
| dprotein | Protein degradation rate | 0.050 | 1/min | [http://jb.asm.org/content/189/23/8746.full] | There is no active degradation pathway and that dilution is the dominant way by which the protein level decreases in a cell. Rate = 1/doubling time, where doubling time = 20min. Assuming steady-state growth in LB broth as presented in paper. |
| k2 | Protein production rate (PudA) | 2.2 | 1/min | Please see derivation 2 below. | Please see derivation 2 below. |
| [Arabinose] | Concentration of arabinose | Initial: 0.008 | mM | ||
| [mRNA] | Concentration of mRNA | - | mM | - | - |
| [PudA] | Concentration of PudA | - | mM | - | - |
1.Derivation of the maximal expression rate,β
- Average molecular weight (Mw) of a base pair = 660g/mol[http://www.geneinfinity.org/sp/sp_dnaprop.html][http://www.lifetechnologies.com/uk/en/home/references/ambion-tech-support/rna-tools-and-calculators/dna-and-rna-molecular-weights-and-conversions.html]
- Average mass of a base pair = 660g/mol x 1.66x10-24 = 1.1x10-21g
- Volume of an E.coli cell = 1µm3[http://kirschner.med.harvard.edu/files/bionumbers/fundamentalBioNumbersHandout.pdf] = 1x10-15L
- BioBrick assembly plasmid pSB1C3 is a high copy number plasmid (100-300 copies per cell)[http://parts.igem.org/Part:pSB1C3?title=Part:pSB1C3]
- assume 200 copies per cell
- ∴ concentration of the gene per cell = N x 200 x 1.66x10-6mM, where N = number of base pairs
- ∴ concentration of the gene PudA (N = 1644) in the volume of an E.coli cell is = 0.55mM
- Transcription rate in E.coli = 80bp/s[http://kirschner.med.harvard.edu/files/bionumbers/fundamentalBioNumbersHandout.pdf] = 80 x 1.66x10-6mM/s = 80 x 1.66x10-6 x 60mM/min = 7.97x10-3mM/min
- ∴ Rate of mRNA_pudA production under the control of pBAD = 7.97x10-3 ÷ 0.25 = 0.015/min
- Average molecular weight(Mw) of an amino acid(aa)= 110g/mol[http://www.genscript.com/conversion.html][http://www.promega.com/~/media/Files/Resources/Technical%20References/Amino%20Acid%20Abbreviations%20and%20Molecular%20Weights.pdf]
- Average mass of an amino acid = 110g/mol x 1.66x10-24=1.83x10-22g/L
- Translation rate = 20aa/s = (20 x 1.66x10-5 x 60)mM/min = 0.020mM/min
- PudA comprises of 548aa[http://www.sciencedirect.com/science/article/pii/S0964830598000663]
- ∴concentration of PudA's aa in the volume of an E.coli = 1.66x10-5mM x 548 = 9.10x10-3mM
- ∴ Rate of protein production = 0.020 ÷ 4.25x10-3 = 2.2/min
PUR degradation module
The reaction equation of the PUR degradation is:
[Polyurethane]+[PudA]= 5 [ethylene glycol] + 5 [polyisocyanate] + [PudA]
Assumptions:
We assumed 1 mole of polyurethane dispersion can produce 5 moles of ethylene glycol.
The molecular weight of a single polyurethane monomer is 470 g/mol whereas the molecular weight of the polyurethane dispersion is around 2000 g/mol.[http://www.polyurethanes.basf.de/pu/solutions/en/content/group/Arbeitsgebiete_und_Produkte/Grundprodukte/Lupraphen-Produktuebersicht_Gewicht]Therefore, the short-chain polyurethane consists approximately 5 monomers. 5 molecules of ethylene glycol will be produced by degrading one chain of polymer.
We also assumed a simple Michaelis-Menten mechanism for PudA
"[PudA]"is the concentration of the PudA enyme whereas "[Polyurethane]" is the concentration of the polyurethane. The kinetic parameters are defined as below:
| Parameter | Description | Value | Units | Sources | Species | |
|---|---|---|---|---|---|---|
| Km | Michaelis constant | 51.5 | mM | [http://www.sciencedirect.com/science/article/pii/S0964830598000663] | Comamonasacidovorans | |
| Kcat | Turnover number | 141.75 | 1/min | [http://www.sciencedirect.com/science/article/pii/S0964830598000663] | Comamonasacidovorans |
Assumptions:
1. The kinetic parameters of PudA are not from the strain E.coli K12 MG1665 we are using. So the assumption here is that the kinetic property of pudA in E.coli is close to that in Comamonasacidovorans
2. Considerations of temperature change and PH change are not included in the model. Therefore, the assumption is that they remain constant during the degradation
Secretion module
The efficiency of Secretion is assumed to be 90% secretion over 2 hours.[http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251600/] The rate of secretion in the model is therefore:
rate of secretion = 0.9[concentration of PudA]/120 (mM/mins)
Simulation Results
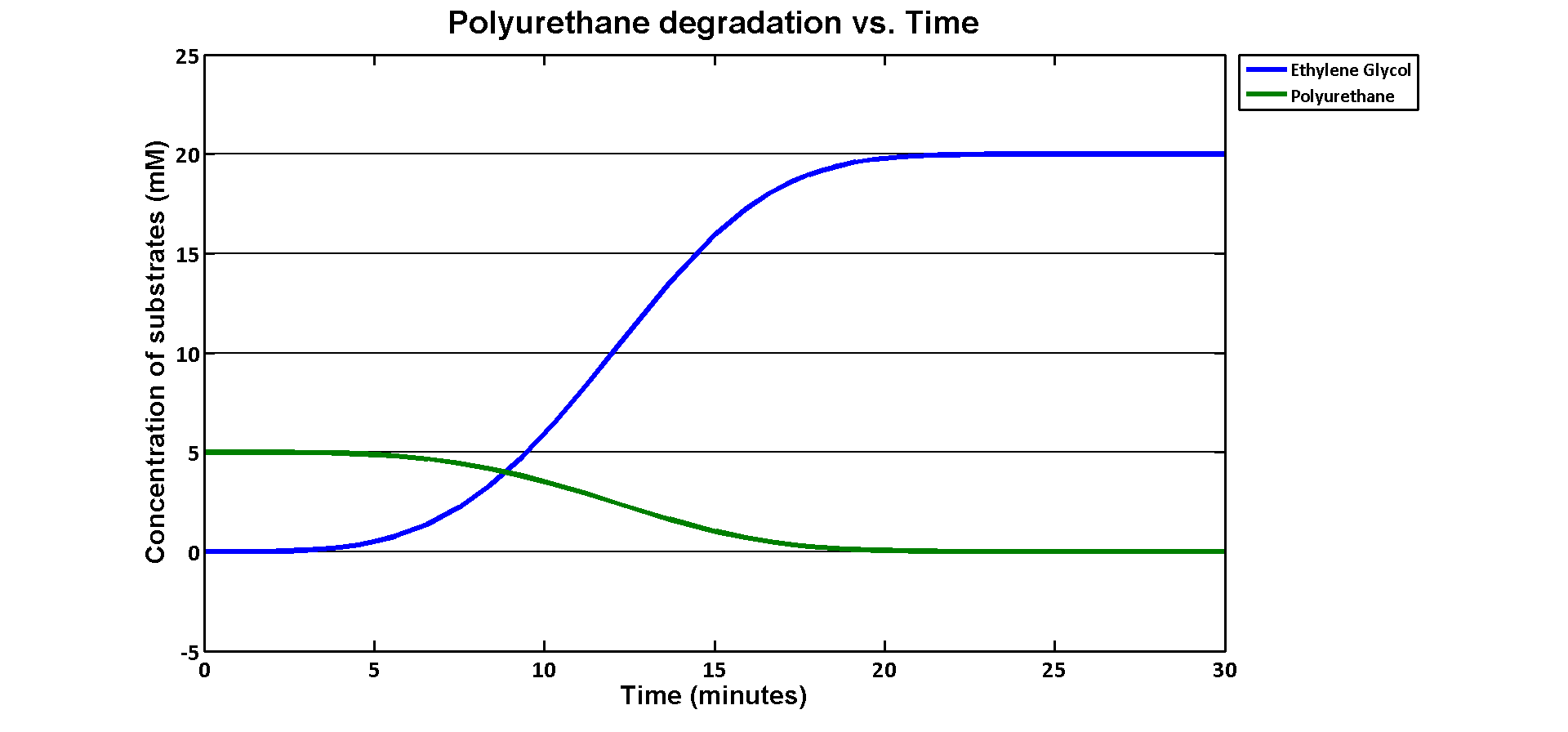
In terms of ethylene glycol toxicity, if we can't let the concentration of ethylene glycol over than 200mM, the maximum initial concentration of polyurethane dispersion is 40mM because one mole of polyurethane dispersion can produce 5 moles of ethylene glycol. (from the derivation above) If we put 40mM as the initial concentration in the model, the simulation result is:
40mM of polyurethane dispersion can be efficiently degraded in 20 mins. In case if we keep the concentration of the plastic below 40mM. The ethylene glycol won't affect the cell growth in our assays. 40mM is converted to 80g/L which suggests no more than 80 grams of polyurethane to be put in the system of 1L volume in order to avoid toxicity.
As for the implementation of MAPLE system, the polyurethane degradation enzymes in our system are very efficient. Therefore, we need an efficient filter which removing ethylene glycol from the system.
PUR degradation model:Download
2.0 P(3HB) synthesis model ▼
This section is about modelling the plastic-synthesising E.coli developed in our project. The plastic that this module synthesises is the bioplastic poly-3-hydroxybutyrate, also known as P(3HB). The content below shows the process of building, designing and optimisation of the model. Also, the considerations, assumptions and limitations will be discussed.
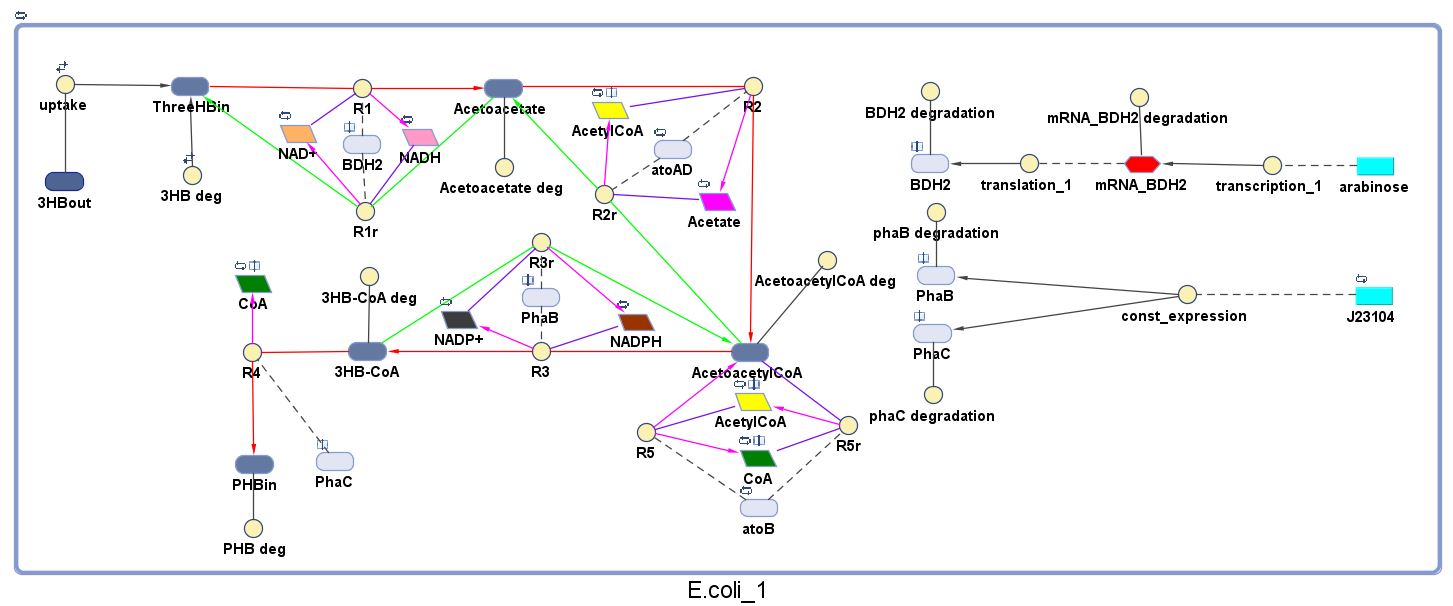
The model was built and simulated in Simbiology, a Matlab package designed for modelling biological systems. In our model we have a compartment (labelled "E.coli_1") that represents our engineered cell. The compartment contains the reactions and species that interact with the polymer synthesis pathway.
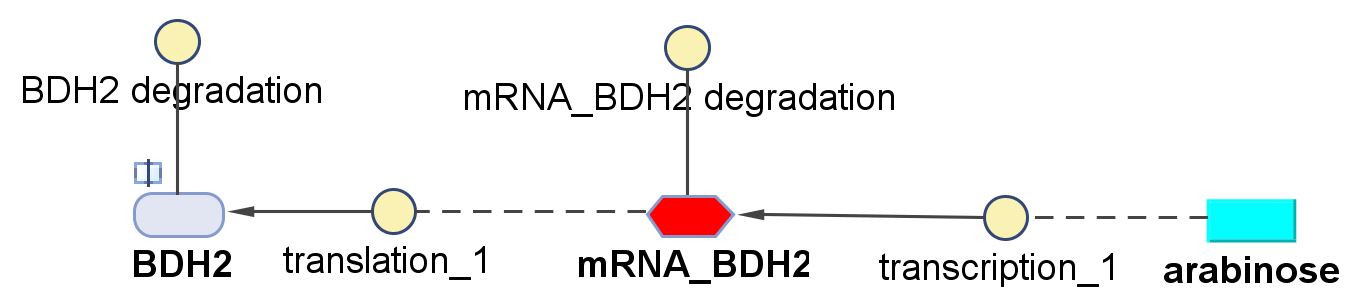
The pathways on the right half of the cell are genetic expressions of the 3 enzymes involved in the plastic synthesis pathway. The pathways on the left represent the actual P(3HB) production pathway.
As can be seen in the "Key", the yellow,solid circle represents a reaction object and for each object parameters the following needs to be specified for the simulation to work:
- Rate equation or kinetic law (e.g. mass action, Michaelis-Menten etc.)
- Parameters in the rate equation
- Species involved in the reaction
- Value and units for each parameter
Simbiology will then use the ODE solver (ode15) to solve these ODEs and give a plot of the specified output(s) (concentration level of a species over time, for instance).
2.1 Ordinary Differential Equations ▼
Values, sources and assumptions ▼
| Parameter | Description | Value | Units | Sources | Assumptions/Notes |
|---|---|---|---|---|---|
| β | maximum rate of transcription | 0.032 | mM/min | Please see derivation 1 below. | Please see derivation 1 below. |
| n | Hill coefficient | 2.0 | dimensionless | [http://parts.igem.org/Part:pSB1C3?title=Part:pSB1C3] | For pBAD strong. Taken from the parts registry page. Rounded to 2.0 from 2.26 as Simbiology wouldn't allow a non-integer value for such parameter. |
| K | Activation coefficient | 0.0031 | mM | [http://parts.igem.org/Part:BBa_K206000:Characterization] | For pBAD strong. Taking the "switch point" (from the corresponding parts registry page) as the activation coefficient. |
| dmRNA | mRNA degradation rate | 0.10 | 1/min | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC420366/pdf/07X.pdf] | Taking the value of mRNA half-life in E.coli strain MG1655 as 6.8min. rate = ln2/half-life = ln2/6.8 = 0.10 |
| dprotein | Protein degradation rate | 0.050 | 1/min | [http://jb.asm.org/content/189/23/8746.full] | There is no active degradation pathway and that dilution is the dominant way by which the protein level decreases in a cell. Rate = 1/doubling time, where doubling time = 20min. Assuming steady-state growth in LB broth as presented in paper. |
| k2 | Protein production rate (BDH2) | 4.7 | 1/min | Please see derivation 2 below. | Please see derivation 2 below. |
| k3 | Protein production rate (PhaCB) | 0.58 | mM/min | Please see derivation 3 below. | Please see derivation 3 below. |
| [Arabinose] | Concentration of arabinose | Initial: 0.008 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [mRNA] | Concentration of mRNA | - | mM | - | - |
| [BDH2] | Concentration of BDH2 | - | mM | - | - |
| [PhaB] | Concentration of PhaB | - | mM | - | - |
| [PhaC] | Concentration of PhaC | - | mM | - | - |
Derivations ▼
- Average molecular weight (Mw) of a base pair = 660g/mol[http://www.geneinfinity.org/sp/sp_dnaprop.html][http://www.lifetechnologies.com/uk/en/home/references/ambion-tech-support/rna-tools-and-calculators/dna-and-rna-molecular-weights-and-conversions.html]
- Average mass of a base pair = 660g/mol x 1.66x10-24 = 1.1x10-21g
- Volume of an E.coli cell = 1µm3[http://kirschner.med.harvard.edu/files/bionumbers/fundamentalBioNumbersHandout.pdf] = 1x10-15L
- BioBrick assembly plasmid pSB1C3 is a high copy number plasmid (100-300 copies per cell)[http://parts.igem.org/Part:pSB1C3?title=Part:pSB1C3]
- assume 200 copies per cell
- ∴ concentration of the gene per cell = N x 200 x 1.66x10-6mM, where N = number of base pairs
- ∴ concentration of the gene BDH2 (N = 768) in the volume of an E.coli cell is = 0.25mM
- Transcription rate in E.coli= 80bp/s[http://kirschner.med.harvard.edu/files/bionumbers/fundamentalBioNumbersHandout.pdf] = 80 x 1.66x10-6mM/s = 80 x 1.66x10-6 x 60mM/min = 7.97x10-3mM/min
- ∴ Rate of mRNA_BDH2 production under the control of pBAD = 7.97x10-3 ÷ 0.25 = 0.032/min
2.Protein production rate of BDH2, k2
- Average molecular weight(Mw) of an amino acid(aa)= 110g/mol[http://www.genscript.com/conversion.html][http://www.promega.com/~/media/Files/Resources/Technical%20References/Amino%20Acid%20Abbreviations%20and%20Molecular%20Weights.pdf]
- Average mass of an amino acid = 110g/mol x 1.66x10-24=1.83x10-22g/L
- Translation rate = 20aa/s = (20 x 1.66x10-5 x 60)mM/min = 0.020mM/min
- BDH2 comprises of 256aa[http://www.uniprot.org/uniprot/Q2PEN2&format=html]
- ∴concentration of BDH2's aa in the volume of an E.coli= 1.66x10-5mM x 256 = 4.25x10-3mM
- ∴ Rate of protein production = 0.020 ÷ 4.25x10-3 = 4.7/min
3.Protein production rate for PhaB and PhaC, k3
- Relative promoter strengths: J23104 = 1.3RPU, J23101 = 1.0RPU.[http://sb6.biobricks.org/poster/automated-bioparts-characterisation-for-synthetic-biology/]
- ∴ 104 is 1.3x stronger than 101.
- In absolute units: take GFP synthesis rate (molecules per min per cell) and approximate that as a generic protein synthesis rate for the promoter.
- GFP synthesis rate of 101 = 2232 molecules per min per cell.[http://sb6.biobricks.org/poster/automated-bioparts-characterisation-for-synthetic-biology/]
- ∴ GFP synthesis rate of 104 = 1.3 x 2232 = 2902 molecules per min per cell
- Assume 1 molecule in an E.coli cell gives a concentration of 1nM.
- ∴ GFP synthesis rate of 104 = 2902 x 1nM = 2.9x10-6nM/min per cell
- Plasmid copy number assumed as 200 (as in derivation 1)
- ∴ GFP synthesis rate of 104 in our E.coli = 200 x 2.9x10-6 = 0.00058nM/min = 0.58mM/min
2.2 Enzyme kinetics ▼
- Note:
- v = velocity of reaction
- Ac-CoA = acetyl-CoA
- AcAc-CoA = Acetoacetyl-CoA
- AcAc = acetoacetate
BDH2 (3-hydroxybutyrate dehydrogenase)
Values, sources and assumptions
| Parameter | Description | Value | Units | Source | Assumptions/Notes |
|---|---|---|---|---|---|
| kcat,BDH2_f | Turnover number of BDH2 in forward reaction | 22200 | 1/min | [http://jb.oxfordjournals.org/content/early/2009/01/03/jb.mvn186.full.pdf] | Sequences derived from Pseudomonas fragi but kinetic values from expression and purification of enzymes in E. coli XL1 Blue |
| kcat,BDH2_r | Turnover number of BDH2 in reverse reaction | 7200 | 1/min | [http://jb.oxfordjournals.org/content/early/2009/01/03/jb.mvn186.full.pdf] | Sequences derived from Pseudomonas fragi but kinetic values from expression and purification of enzymes in E. coli XL1 Blue |
| Ki,NAD+ | Inhibition constant of BDH2 with NAD+ | 2.5 | mM | [http://www.hal.inserm.fr/hal-00374321/] | Organism Tetrahymena pyriformis |
| Ki,NADH | Inhibition constant of BDH2 with NADH | 1.1 | mM | [http://www.hal.inserm.fr/hal-00374321/] | Organism Tetrahymena pyriformis |
| Km,AcAc | Michaelis constant for AcAc | 0.37 | mM | [http://jb.oxfordjournals.org/content/early/2009/01/03/jb.mvn186.full.pdf] | Values in paper reference |
| Km,NAD+ | Michaelis constant for NAD+ | 0.24 | mM | [http://jb.oxfordjournals.org/content/early/2009/01/03/jb.mvn186.full.pdf] | Values in paper reference |
| Km,NADH | Michaelis constant for NADH | 0.010 | mM | [http://jb.oxfordjournals.org/content/early/2009/01/03/jb.mvn186.full.pdf] | Values in paper reference |
| Km,3HB | Michaelis constant for 3HB | 0.80 | mM | [http://jb.oxfordjournals.org/content/early/2009/01/03/jb.mvn186.full.pdf] | source organism Pseudomonas fragi |
| [BDH2] | Concentration of BDH2 | - | mM | - | - |
| [NAD+] | Concentration of NAD+ | initial: 1.6x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [3HB] | Concentration of 3HB | initial: 0.0010 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [AcAc] | Concentration of AcAc | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [NADH] | Concentration of NADH | initial: 2.5x10-14 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
atoAD (Acetyl-CoA:acetoacetyl-CoA transferase (α and β subunits))
Values, sources and assumptions
| Parameter | Description | Value | Units | Sources | Assumptions/Notes |
|---|---|---|---|---|---|
| Vmax,atoAD_f | Maximum rate of atoAD in forward reaction | 0.00244 | mM/min | [http://www.sciencedirect.com/science/article/pii/000398617590003X] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Vmax,atoAD_r | Maximum rate of atoAD in reverse reaction | 0.0108 | mM/min | [http://www.sciencedirect.com/science/article/pii/000398617590003X] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,AcAc | Michaelis constant for AcAc | 1.86 | mM | [http://www.sciencedirect.com/science/article/pii/000398617590003X] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,Ac-CoA | Michaelis constant for Ac-CoA | 0.26 | mM | [http://www.brenda-enzymes.org/php/result_flat.php4?ecno=2.8.3.8] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,Acetate | Michaelis constant for Acetate | 53.1 | mM | [http://aem.asm.org/content/73/24/7814.full.pdf] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,AcAc-CoA | Michaelis constant for AcAc-CoA | 0.035 | mM | [http://www.sciencedirect.com/science/article/pii/000398617590003X] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| [AcAc] | Concentration of AcAc | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [Acetate] | Concentration of Acetate | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [AcAc-CoA] | Concentration of AcAc-CoA | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [Ac-CoA] | Concentration of Ac-CoA | initial: 1.0x10-14 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
phaB (Acetoacetyl-CoA reductase)
Values, sources and assumptions
| Parameter | Description | Value | Units | Sources | Assumptions/Notes |
|---|---|---|---|---|---|
| kcat,PhaB_f | Turnover number of PhaB in forward reaction | 6120 | 1/min | [http://www.ncbi.nlm.nih.gov/pubmed/23913421] | Gene in Ralstonia eutropha was engineered to put in E.coli and then purified |
| kcat,PhaB_r | Turnover number of PhaB in reverse reaction | 3600 | 1/min | [http://www.ncbi.nlm.nih.gov/pubmed/3286259] | originally in Zoogloea ramigera, expressed in E.coli and then purified |
| Km,NADPH | Michaelis constant for NADPH | 0.15 | mM | [http://www.ncbi.nlm.nih.gov/pubmed/23913421] | Gene in Ralstonia eutropha was engineered to put in E.coli and then purified |
| Km,AcAc-CoA | Michaelis constant for AcAc-CoA | 0.0057 | mM | [http://www.ncbi.nlm.nih.gov/pubmed/23913421] | Gene in Ralstonia eutropha was engineered to put in E.coli and then purified |
| Km,NADP+ | Michaelis constant for NADP+ | 0.0060 | mM | [http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.1997.tb12740.x/pdf] | purified from Methylobacterium extorquens |
| Km,3HB-CoA | Michaelis constant for 3HB-CoA | 0.026 | mM | [http://www.ncbi.nlm.nih.gov/pubmed/3286259] | originally in Zoogloea ramigera, expressed in E.coli and then purified |
| [PhaB] | Concentration of PhaB | - | mM | ||
| [AcAc-CoA] | Concentration of AcAc-CoA | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [NADPH] | Concentration of NADPH | initial:3.8x10-14 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [NADP+] | Concentration of NADP+ | initial:1.5x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [3HB-CoA] | Concentration of 3HB-CoA | - | mM | - | - |
phaC (P(3HB) synthase)
Values, sources and assumptions
| Parameter | Description | Value | Units | Sources | Assumptions/Notes |
|---|---|---|---|---|---|
| kcat,PhaC | Turnover number of PhaC | 1680 | 1/min | [http://ac.els-cdn.com/S0003986101925226/1-s2.0-S0003986101925226-] | |
| Km,PhaC | Michaelis constant of PhaC with 3HB-CoA | 0.14 | mM | [http://www.brenda-enzymes.org/php/result_flat.php4?ecno=2.3.1.B5] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| [PhaC] | Concentration of PhaC | - | mM | - | - |
| [3HB-CoA] | Concentration of 3HB-CoA | - | mM | - | - |
atoB (acetyl-CoA acetyltransferase)
Values, sources and assumptions
| Parameter | Description | Value | Units | Sources | Assumptions/Notes |
|---|---|---|---|---|---|
| Vmax,atoB_f | Maximum rate of atoB in forward reaction | 3.8x10-5 | mM/min | [http://www.ncbi.nlm.nih.gov/pubmed/9904] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Vmax,atoB_r | Maximum rate of atoB in reverse reaction | 8.5x10-4 | mM/min | [http://www.ncbi.nlm.nih.gov/pubmed/9904] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,atoB | Michaelis constant of atoB with Ac-CoA | 0.47 | mM | [http://www.sciencedirect.com/science/article/pii/0003986176901521] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,AcAc-CoA | Michaelis constant for AcAc-CoA | 0.1 | mM | [http://www.sciencedirect.com/science/article/pii/0003986176901521] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| Km,CoA | Michaelis constant for CoA | 0.25 | mM | [http://www.sciencedirect.com/science/article/pii/0003986176901521] | Enzyme is naturally expressed in E.coli: assume the enzyme is stable during the time course of the simulation. |
| [Ac-CoA] | Concentration of Ac-CoA | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [AcAc-CoA] | Concentration of AcAc-CoA | initial: 1.0x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
| [CoA] | Concentration of CoA | initial: 5.5x10-13 | mM | see section 2.5 "Initial concentrations of metabolites" | see section 2.5 "Initial concentrations of metabolites" |
Reaction kinetics and mechanisms ▼
List of enzymes and the corresponding mechanisms by which they work in the synthesis pathway
- BDH2: Ordered-sequential bi-bi mechanism[http://www.brenda-enzymes.org/php/result_flat.php4?ecno=1.1.1.30]
- atoAD: Ping-pong bi-bi mechanism[http://www.sciencedirect.com/science/article/pii/000398617590003X]
- atoB: Ping-pong bi-bi mechanism[http://www.sciencedirect.com/science/article/pii/000398617590003X]
- PhaB: Ordered-sequential bi-bi mechanism[http://digitalcommons.usu.edu/engineering_datasets/1/]
- PhaC: Michaelis-Menten[http://connection.ebscohost.com/c/articles/85133798/new-insights-activation-substrate-recognition-polyhydroxyalkanoate-synthase-from-ralstonia-eutropha]
2.3 Simulation results ▼
Using the values presented in the tables above, the deterministic model was simulated to see how much P(3HB) could be formed inside the cell. It was simulated up to the doubling time of E.coli MG1655 (20 mins), as the concentration of P(3HB) would become inaccurate afterwards due to cell division. However, with the single cell model, it would be difficult to reflect how much of the plastic could actually be formed on a macroscopic scale. Therefore, modifications to the simulation algorithm were needed to give scaled-up simulation results.
In the scaled-up regime, though, new limitations were encountered. For example, growth, cell division and the concentration gradient that each cell encounters and how much can be taken up by the cells. Given the data available, what could be done was to assume that the bacteria were in a bioreactor, with constant nutrient and oxygen supply, and that they were in the phase of stationary growth (please see assumptions and explanations under the graphs below).
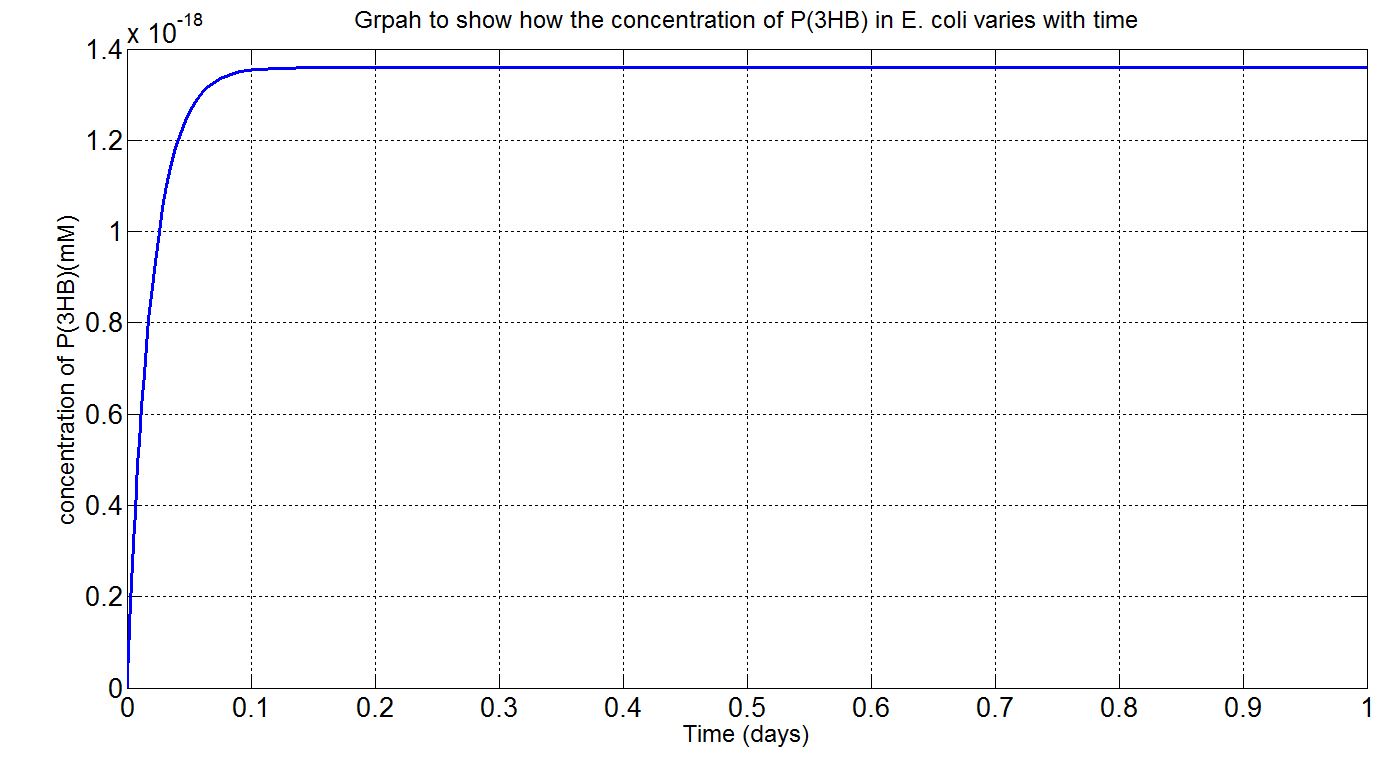
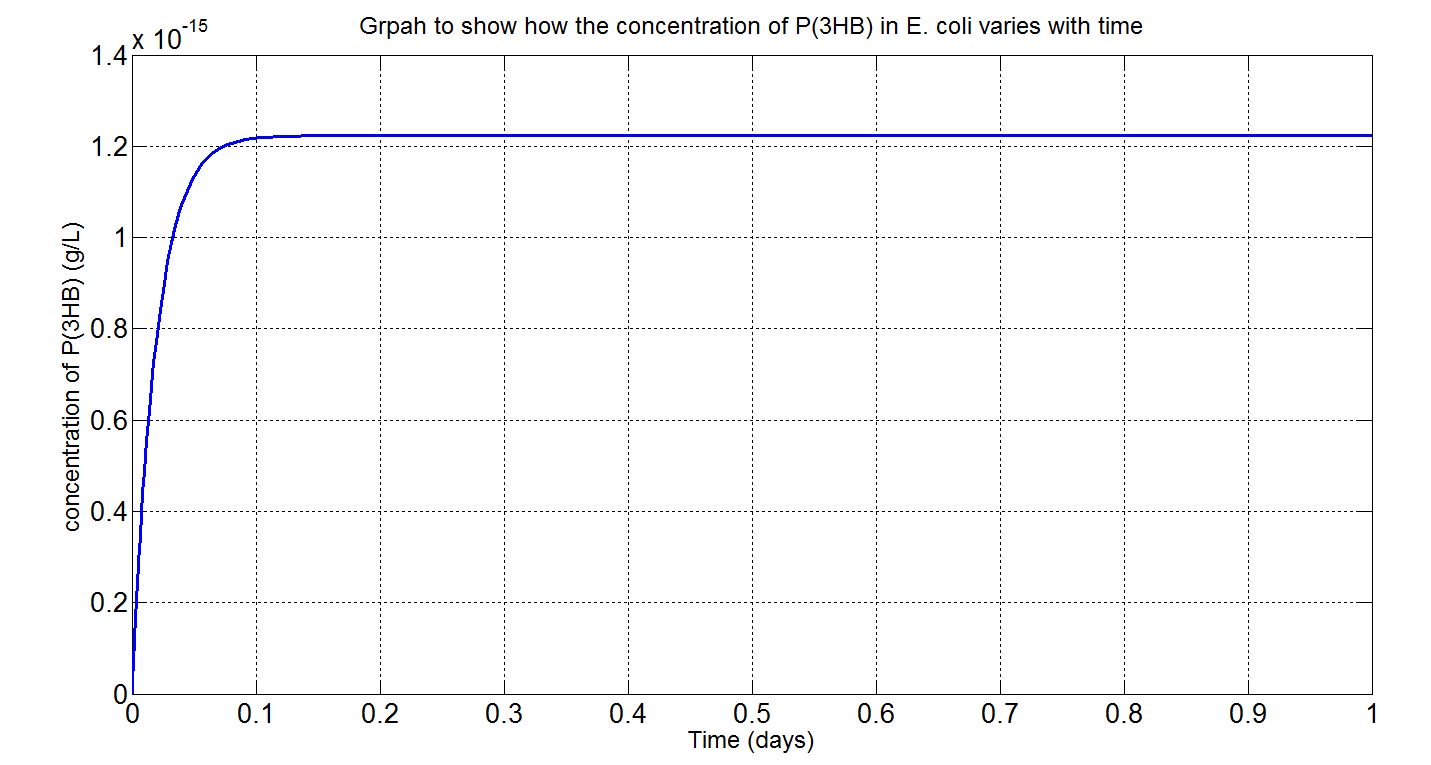
Here, three graphs are presented to show P(3HB)formation over time. The first two graphs are (i)single-cell model results, concentration in mM and (ii) single-cell model results, concentration in g/L. The first graph was the original graph produced from Simbiology as all concentrations were expressed in mM in the model. However, to gain a more intuitive sense of how much could be produced g/L was used. The third graph is the scaled-up simulation result. As it is a scaled-up version and the stationary growth (constant number of cells over time) assumption applies, the doubling time consideration is not included here and simulation time is now 1 day (for the purpose of a more realistic representation).
(i) please see assumptions and explanations below
(ii) please see assumptions and explanations below
(iii) please see assumptions and explanations below
Graphs (i) and (ii): assumptions and explanations
- To convert from mM to g/L:
- Molecular weight(Mw) of P(3HB) = 1.8x1012g/mol[http://webpages.charter.net/hvalentin/PDF-files/18.pdf]
(Please note that this value was taken from a paper about the "Production of poly(3-hydroxybutyrate-co-4-hydroxybutyrate) in recombinant Escherichia coli grown on glucose" instead of the production of poly(3-hydroxybutyrate). However, this is the closest we managed to get as far as species, substrate and type of plastic are concerned.
Also, it was noticed that the molecular weight would change depending on the cultivation time (as conditions such as growth and pH can also affect it) and it was hard to find a paper that contains the exact same conditions and the range of cultivation times for our simulations. Furthermore, in reality there wouldn't be a single value of molecular weight obtained, but a range of values (known as polydispersity).[http://aem.asm.org/content/78/9/3177.full] Therefore, it was assumed that the molecular weight would remain constant during the time course of the simulation and that the value should be considered as an average.)
- ∴P(3HB)concentration in mM x 10-3 x 1.8x1012g/mol = concentration in g/L
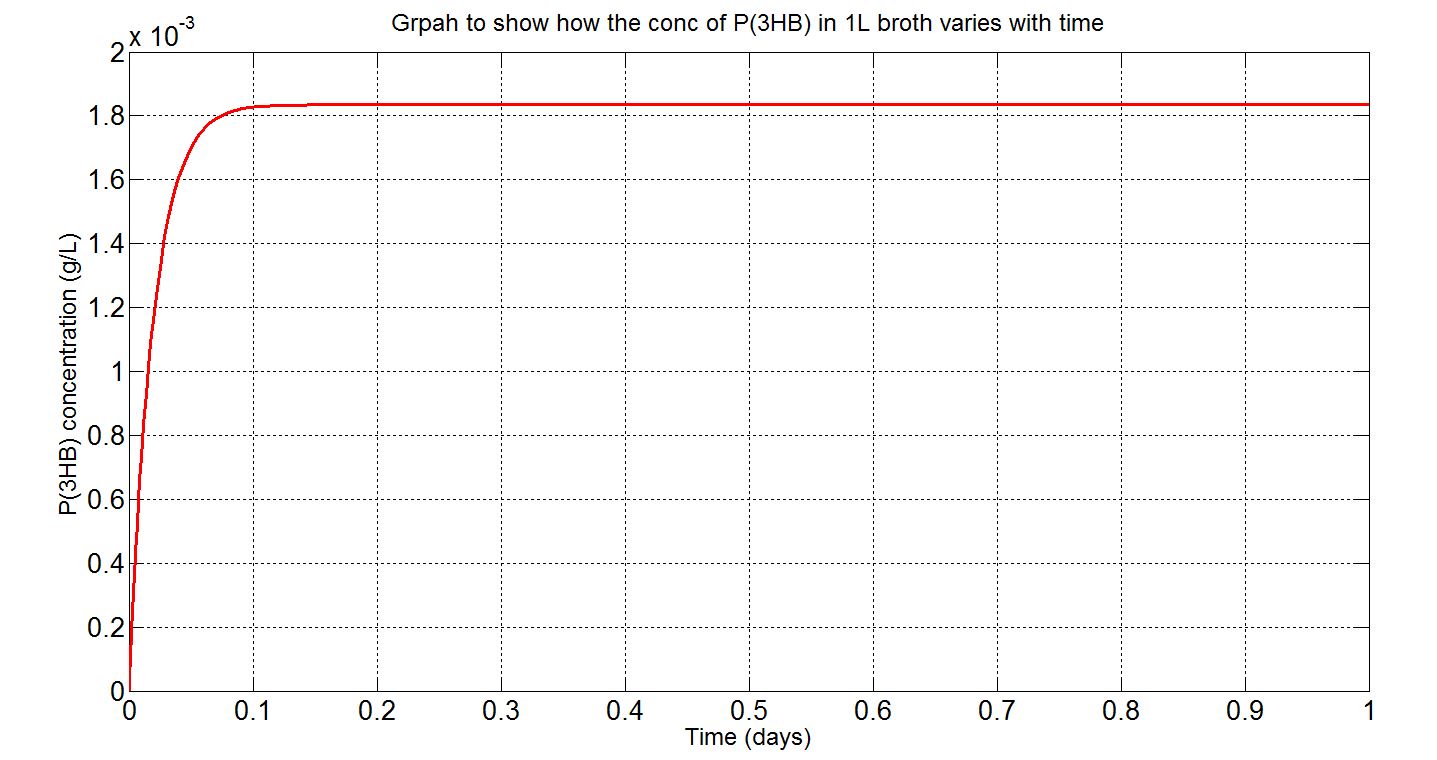
Graph (iii): assumptions and explanations
- Assume 1.5x1012cells in a 1L broth, calculated from value obtained from BioNumbers. A range was given: 1-2x109 cells/ml, so 1.5x109 was taken. [http://bionumbers.hms.harvard.edu/bionumber.aspx?s=n&id=104831&ver=5]
- Conditions and considerations accompanying this value are:
- growth medium: LB broth
- Cultivation temperature: 37°C
- wild-type E. coli K-12 strain MG1655 (NB: though not exactly our engineered one, it is the correct strain)
- Conditions and considerations accompanying this value are:
- Assume stationary growth phase.
- Assume that the amount of P(3HB) accumulated during the time course of simulation doesn't exceed the cell's capacity or ability to contain it.
- Same metabolic considerations as above (single-cell results)
- Simbiology single-cell simulation results multiplied by the number of cells stated above to give the red curve.
2.4 Model-guided design and optimisation ▼
Sensitivity analysis: species concentrations
What is sensitivity analysis and why do it?
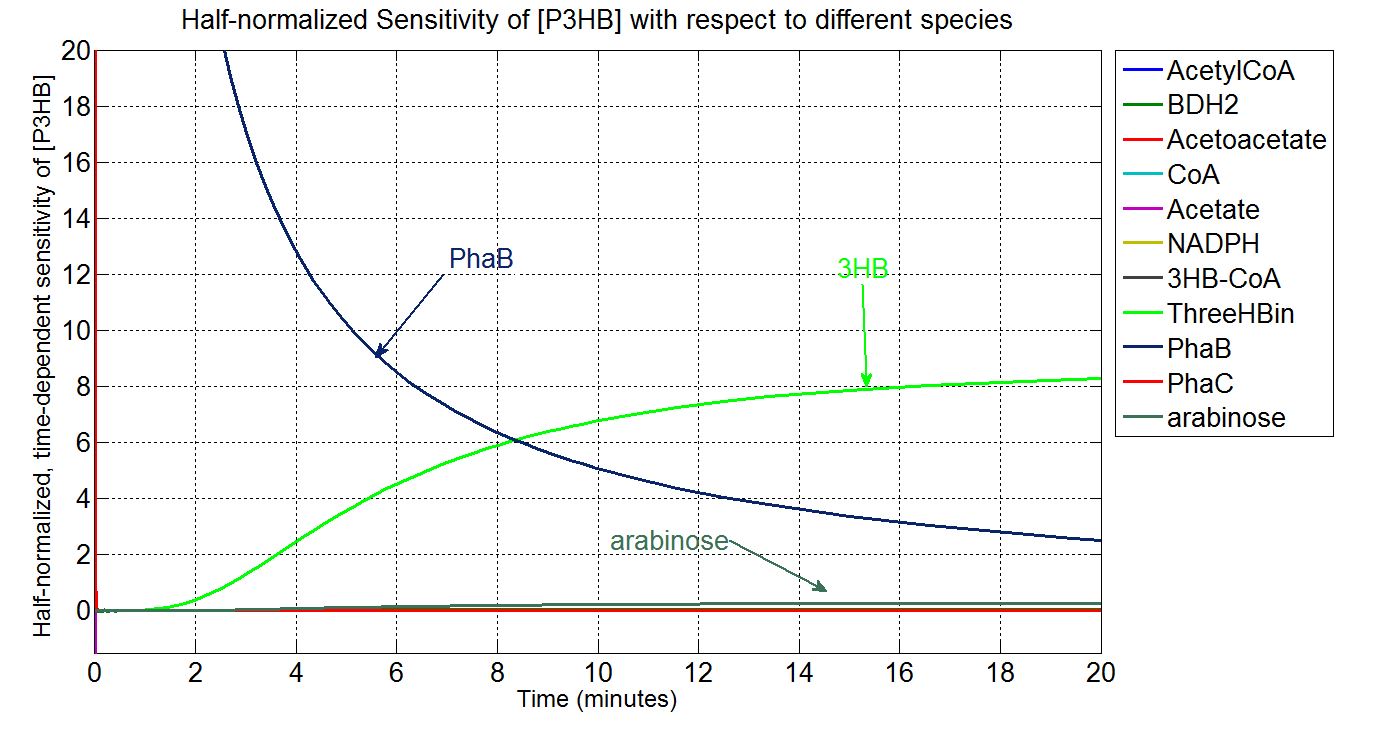
In order to look at how we could increase the production of P(3HB) we decided to run a sensitivity analysis (also in Simbiology) to identify what species in the model P(3HB) is sensitive to, given the specific conditions with which we set the model. In other words, we calculated the time-dependent sensitivity of P(3HB) with respect to the initial conditions of other species (species formed along the synthesis pathway and enzymes involved).[http://www.mathworks.co.uk/help/simbio/ug/calculating-sensitivities.html#brbumlr|ref]
For a species x, whose concentration depends on time, it can be expressed as x(t). To calculate the sensitivity of x(t) with respect to another species y(t), the sensitivity with normalisation relative to the numerator x(t) can be calculated as:
Therefore, the underlying algorithm for this sensitivity analysis of P(3HB) is:
Sensitivity analysis: enzyme concentrations
The sensitivity analysis of the enzymes involved in the synthetic pathway is carried out, which determine the most sensitive enzyme in the pathway
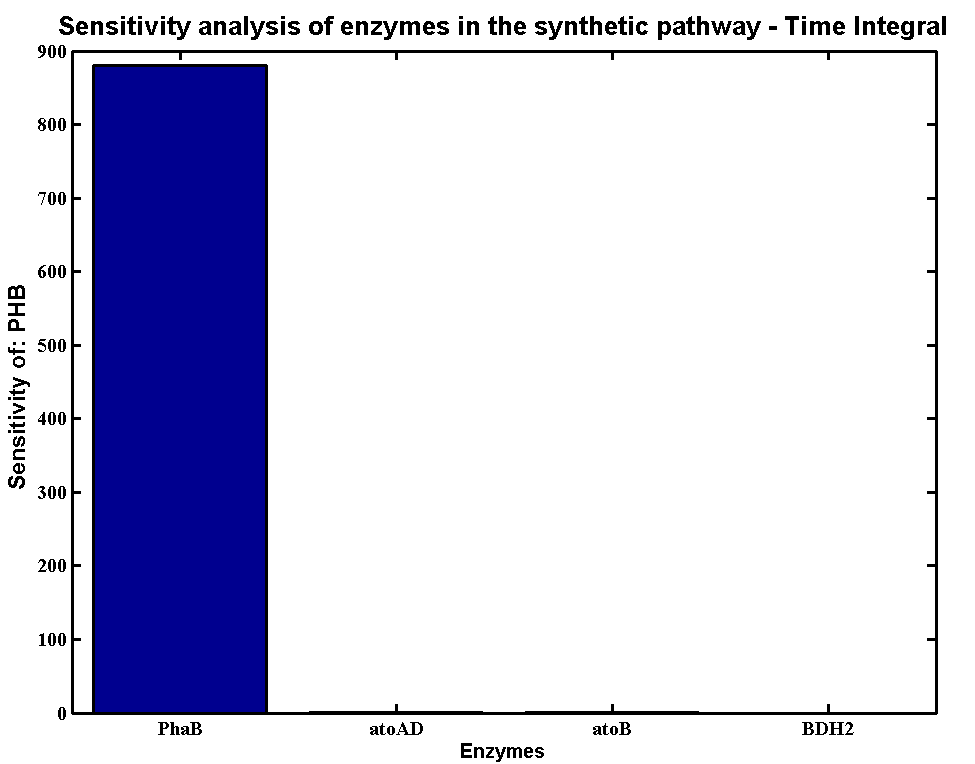
We plot a time integral of the sensitivity analysis which has the algorithm as:
The senstivity result is:
According to the simulation result, we found that the PHB production is highly sensitive to the concentration of PhaB and PhaC. In contrast, the system is not sensitive to the concentration of atoAD and atoB. In theory, increase the concentration of any of the enzyme will increase the flux of the system. However, for the strain E.coli K12 MG1655, both atoAD (acetoacetate:acetoacetyl-coA transferase) and atoB (acetoacetyl-coA thiolase) favour the reverse reactions, it becomes the rate limiting step in our pathway. Acetoacetyl-coA intends to be converted to acetoacetate instead of 3HB-coA. [http://www.sciencedirect.com/science/article/pii/0003986177904969]Therefore, the concentration of PhaB become really critical which push the system to flow in the forward direction.
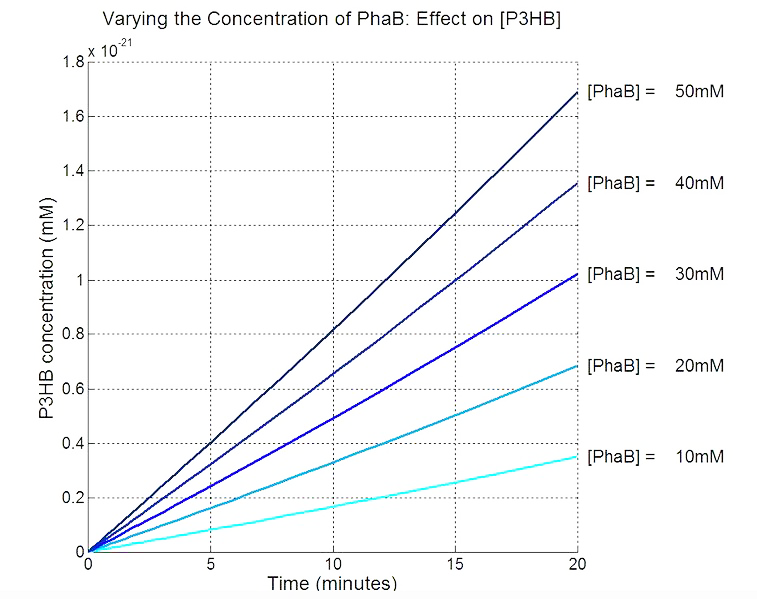
Scan with different levels of PhaB
The plot below is to show that increasing PhaB concentration within the engineered organism would have a positive effect on the concentration of the P(3HB) synthesised. As so many parameters within the model were taken from different sources, it would be difficult to give a simulation plot that is accurate quantitatively. It was decided to show the single cell simulation for this because the scans were carried out up to the doubling time of 20mins, hence avoided the issue of scaling-up which would have otherwise subjected the simulation to more errors. Therefore, this plot should be interpreted from a qualitative perspective and the trend should be observed.
Difference between promoter expressions after optimisation
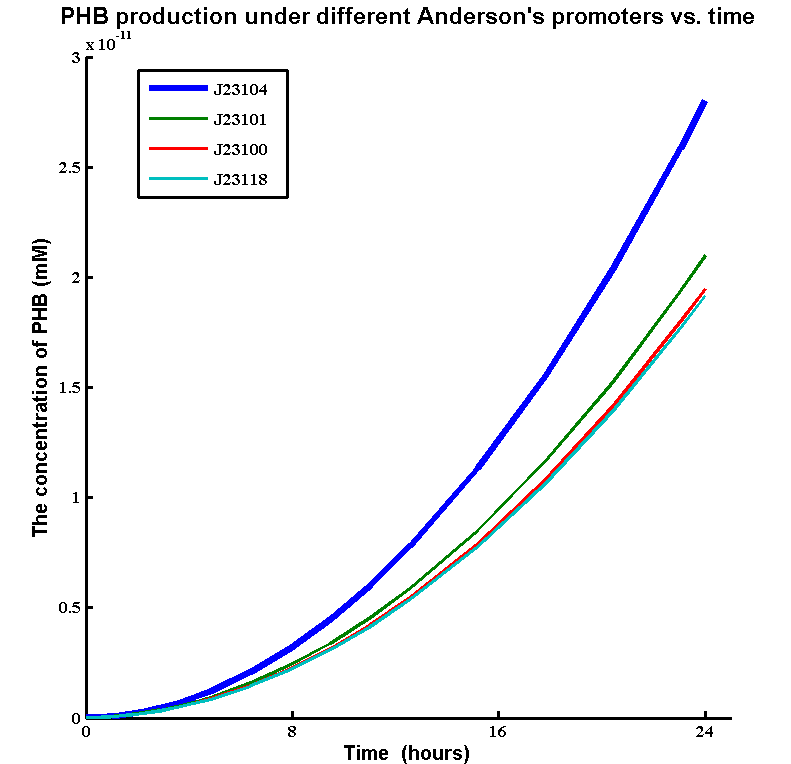
In order to choose the strongest promoter that is available to increase the gene expression. We ran a parameter scan of enzyme expression rates of a range of Anderson's constitutive promoters. The simulations shows the PHB production under different Anderson's promoters. The promoters we tested are:
| promoter | biobrick | GFP synthesis rate (mM/min) |
|---|---|---|
| J23100 | [http://parts.igem.org/Part:BBa_J23100] | 0.41 |
| J23101 | [http://parts.igem.org/Part:BBa_J23101] | 0.45 |
| J23104 | [http://parts.igem.org/Part:BBa_J23104] | 0.58 |
| J23118 | [http://parts.igem.org/Part:BBa_J23118] | 0.31 |
The derivations of the GFP synthesis rate are:
*Relative promoter strengths: J23104 = 1.3RPU, J23101 = 1.0RPU, J23100 = 0.92RPU, J23118 = 0.76RPU[http://sb6.biobricks.org/poster/automated-bioparts-characterisation-for-synthetic-biology/]
- In absolute units: take GFP synthesis rate (molecules per min per cell) and approximate that as a generic protein synthesis rate for the promoter.
- GFP synthesis rate of 101 = 2232 molecules per min per cell.[http://sb6.biobricks.org/poster/automated-bioparts-characterisation-for-synthetic-biology/]
- Assume 1 molecule in an E.coli cell gives a concentration of 1nM.
- ∴ GFP synthesis rate of 101 = 2232 x 1nM = 2.2x10-6nM/min per cell
- Plasmid copy number assumed as 200 (as in derivation 1)
- ∴ GFP synthesis rate of 101 in our E.coli = 200 x 2.2x10-6 = 0.00045nM/min = 0.45mM/min
- ∴ The GFP synthesis rates of other promoters can be calculated by multiplying the RPU value with the GFP synthesis rate of J23101 promoter.
The simulation result:
Although there is just a small increase the gene expression rate, J23104 promoter achieved a much higher PHB production than other constitutive promoters. Therefore, we decided to use J23104 promoter for our improved biobrick [http://parts.igem.org/Part:BBa_K1149051 BBa_K1149051].
Here is our experimental results that show we have 11 times more PHB after the change of promoter.
P(3HB) synthesis model:Download
Additional script:Download
2.5 Metabolic considerations ▼
In Module 1: Resourceful Plastic, glucose from the degradation is used as the input for the system that produces P(3HB). The P(3HB) synthetic pathway hence needs to be adapted for this purpose. To do so, 3HB monomer would no longer be the source input to the engineered system. So, BDH2 should be removed from the pathway. Acetyl-coA would then become the key resource to produce P(3HB). Also, a metabolic model was needed to show the production of Acetyl-coA from glucose. Furthermore, our synthetic pathway involves several key metabolites from the pathway such as NADPH, NADP+, NADH and NAD+. The initial concentration of those metabolites can be determined from the metabolic model by assuming they are in their steady states. More features and results of the metabolic model can be found in Module 2.
Initial concentrations of metabolites
The model presented in Section 2.0 contains a synthesis pathway that interact metabolic species involved in so many other metabolic reactions elsewhere in the cell. It was, therefore, difficult to determine how much of these metabolites would be left available for our plastic synthesis pathway to utilise. So, a simple metabolic model of the cell (more detail in next paragraph) was needed to simulate and show insight into this problem. We decided to use the steady state levels of these metabolites (without the engineered pathway) and plug the values into the P(3HB) synthesis model in Section 2.0 as the initial concentrations.
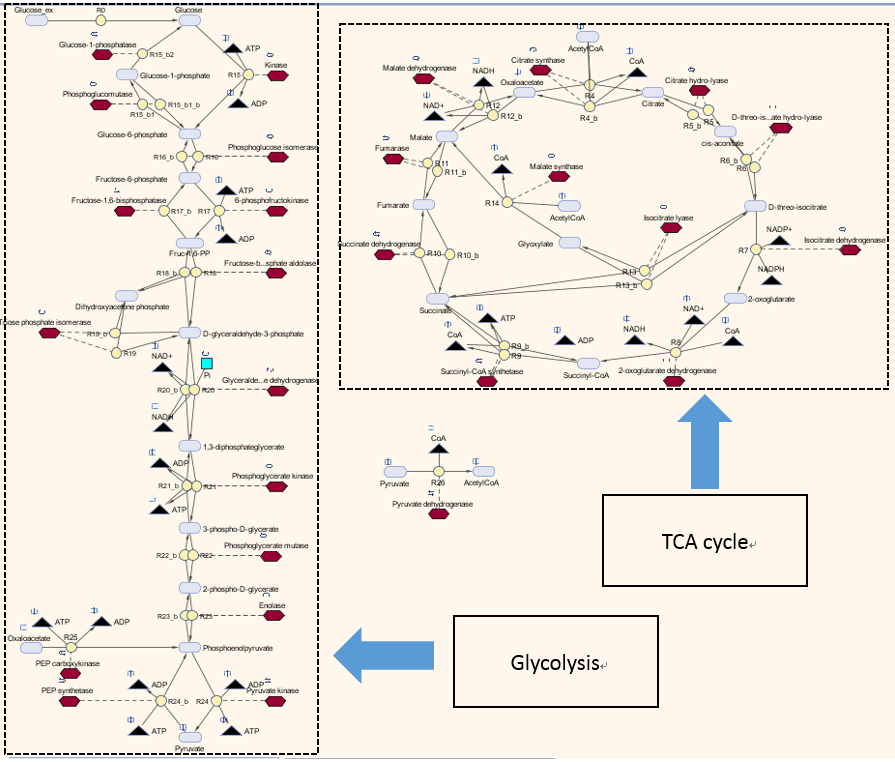
The metabolic model involves two pathways - glycolysis pathway and the TCA cycle, where glucose is used as the sole carbon source. The reason is that glucose is directly taken by the glycolysis pathway and there are interactions between Acetyl-coA and TCA cycle. This Simbiology model is based on the one built by Angela Dixon in Predictive Mathematical Model for Polyhydroxybutyrate Synthesis in Escherichia coli.[http://digitalcommons.usu.edu/engineering_datasets/1/]. All reactions and pathways are based on ODEs and kinetics data presented in the original model. It can dynamically predict the concentration of all metabolites involved over a defined simulation time course.
The metabolic model for determining steady state concentrations is shown as below:
Keys:
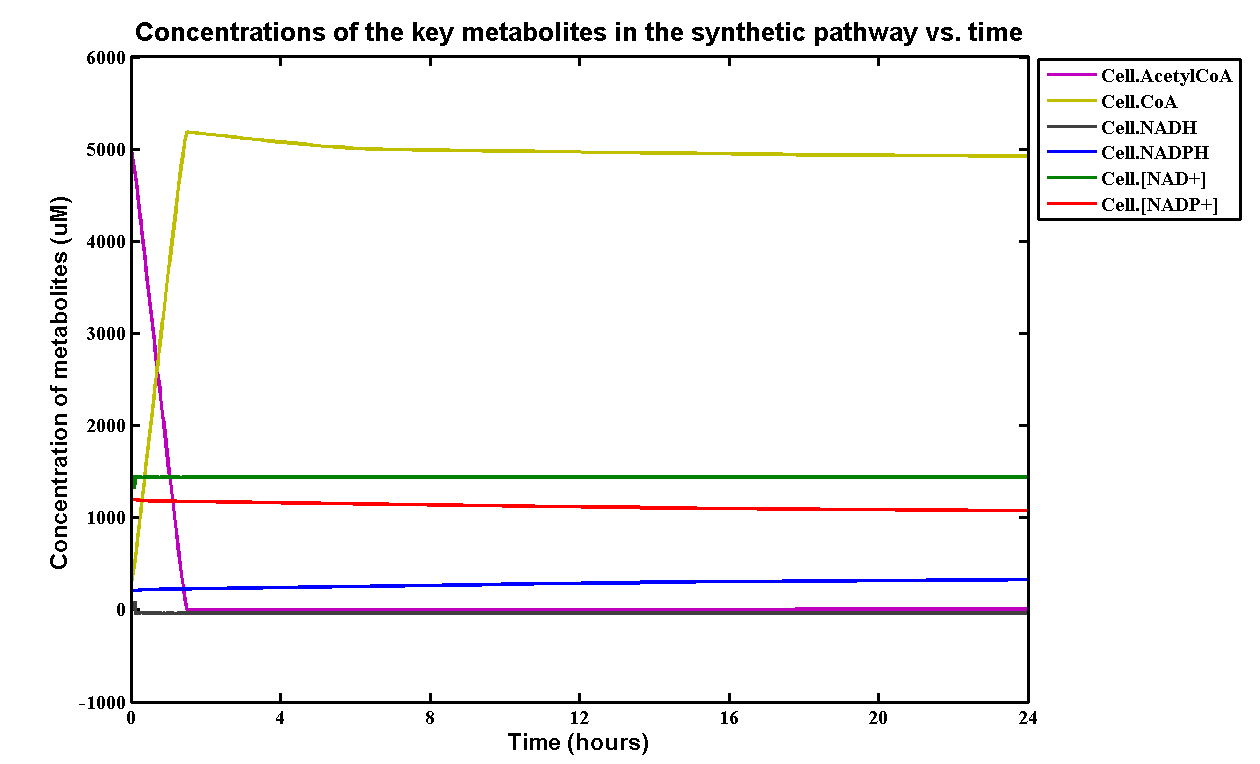
The initial concentrations of all metabolites and kinetic data are referenced in the paper.[http://digitalcommons.usu.edu/engineering_datasets/1/] The simulation of the metabolites is:
Table of initial and steady-state concentrations of the metabolites:
| Substrates | Initial concentration | Steady state concentration | Units | Sources |
|---|---|---|---|---|
| NADH | 200 | 250 | uM | [http://digitalcommons.usu.edu/engineering_datasets/1/] |
| NAD+ | 1200 | 1600 | uM | [http://digitalcommons.usu.edu/engineering_datasets/1/] |
| Acetyl-coA | 1000 | 100 | uM | [http://digitalcommons.usu.edu/engineering_datasets/1/] |
| Coenzyme A | 250 | 5500 | uM | [http://digitalcommons.usu.edu/engineering_datasets/1/] |
| NADPH | 200 | 380 | uM | [http://digitalcommons.usu.edu/engineering_datasets/1/] |
| NADP+ | 1200 | 1100 | uM | [http://digitalcommons.usu.edu/engineering_datasets/1/] |
- Note: The metabolic model was simulated on a macroscopic scale assuming 1.3x1013 cells. Therefore, to scale down the values for the single-cell model, all values would need to be divided by 1.3x1013 and units converted from uM to mM. This gave the following values which were used in the single-cell model in Section 2.0:
| Substrates | Steady state concentration | Units |
|---|---|---|
| NADH | 0.25 | mM |
| NAD+ | 1.6 | mM |
| Acetyl-coA | 0.10 | mM |
| Coenzyme A | 5.5 | mM |
| NADPH | 0.38 | mM |
| NADP+ | 1.1 | mM |
It should also be noted that acetate, acetoacetate and acetyl-CoA cannot be predicted by the metabolic model. Therefore, their initial concentrations are rough estimates (value similar to other metabolites) such that a reasonable P(3HB) output can be produced.
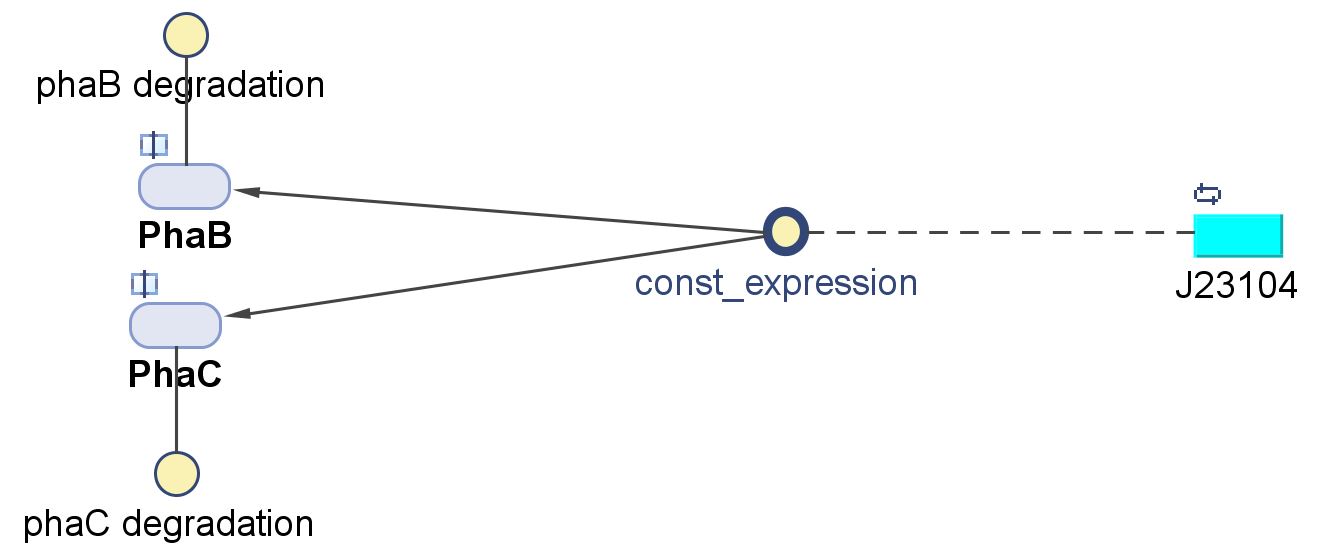
Assembling the optimised bioplastic producing operon
We made two new improved constructs by modifying the native operon:
<p> The modellers in the drylab predicted that an increased level of phaB enzyme will increase the amount of PHB produced and therefore we decided to change the native Ralstonia eutropha promoter and RBS to a strong constitutive E.coli promoter (J23104) and RBS (0034) in front of the operon. We have succeeded with the swap and built the new biobrick [http://parts.igem.org/Part:BBa_K1149052 BBa_K1149052] which we refer to as constitutive phaCAB. In parallel, we have also assembled a different configuration of transcription initiation region: the "hybrid". Surprisingly, our results show that our "hybrid" phaCAB construct [http://parts.igem.org/Part:BBa_K1149051 BBa_K1149051] significantly increased bioplastic production compared to native phaCAB. Please read on to discover more.We tested the promoter and RBS we were going to use by cloning them in front of amilCP blue chromoprotein ([http://parts.igem.org/Part:BBa_K1149020 BBa_K1149020]). It was bright blue and we had some fun playing around with it as part of our communication work "E.coli_Art".

In order to change the promoter, we designed a forward primer that binds to the beginning of the first gene in the operon and contains a Xba restriction site: Pha_Fw (cgcttctagagatggctactgggaaaggagccg). We used pha_Fw and the suffix primer G1005 primer pain to amplify the insert out of the backbone. The new promoter and RBS were J23104 and B0034 from the biobrick [http://parts.igem.org/Part:BBa_K608002 BBa_K608002] We digested the pSB1C3 backbone with this biobrick with SpeI and PstI.
Summary diagram of construction steps:

We performed DpnI digest on the gel purified PCR product (insert) in order to get rid of any contamination from the original plasmid. We did Alkaline Phosphatase treatment of the backbone to remove terminal phosphates in order to decrease self-ligation of plasmid. We transformed the ligation into NEB5 cells and we grew overnight cultures from single cell colonies. The next day, we purified the plasmids and digested 200 ng of them with Xba and PstI to screen for the right clones:

Colony #8 had the expected 4.2kB insert and therefore we sent this plasmid for sequencing. After having the sequenced it, we submitted the new part to registry as [http://parts.igem.org/Part:BBa_K1149051 BBa_K1149051]. This construct has double P+RBS since the native P+RBS stayed there in between the new P+RBS and the operon. We have identified a clone (from colony #6) that contains the new constitutive promoter only and submitted it as [http://parts.igem.org/Part:BBa_K1149052 BBa_K1149052] biobrick. We are testing both the hybrid and normal constitutive constructs for PHB bioplastic production.
Partial sequence of the constitutive phaCAB construct ([http://parts.igem.org/Part:BBa_K1149052 BBa_K1149052]):

Partial sequence of the hybrid phaCAB construct ([http://parts.igem.org/Part:BBa_K1149051 BBa_K1149051]):

Our chassis (MG1655) survives in mixed waste ▼
These assays were designed to test whether our chassis, E.coli (MG1655) could grow directly with waste over a long period of time.
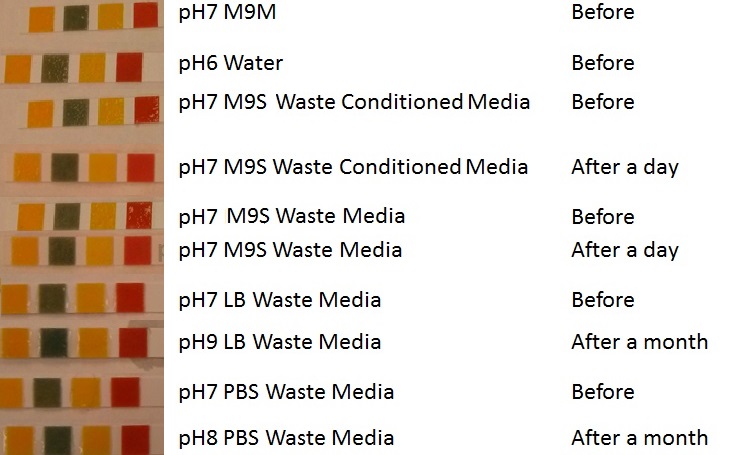
Waste media
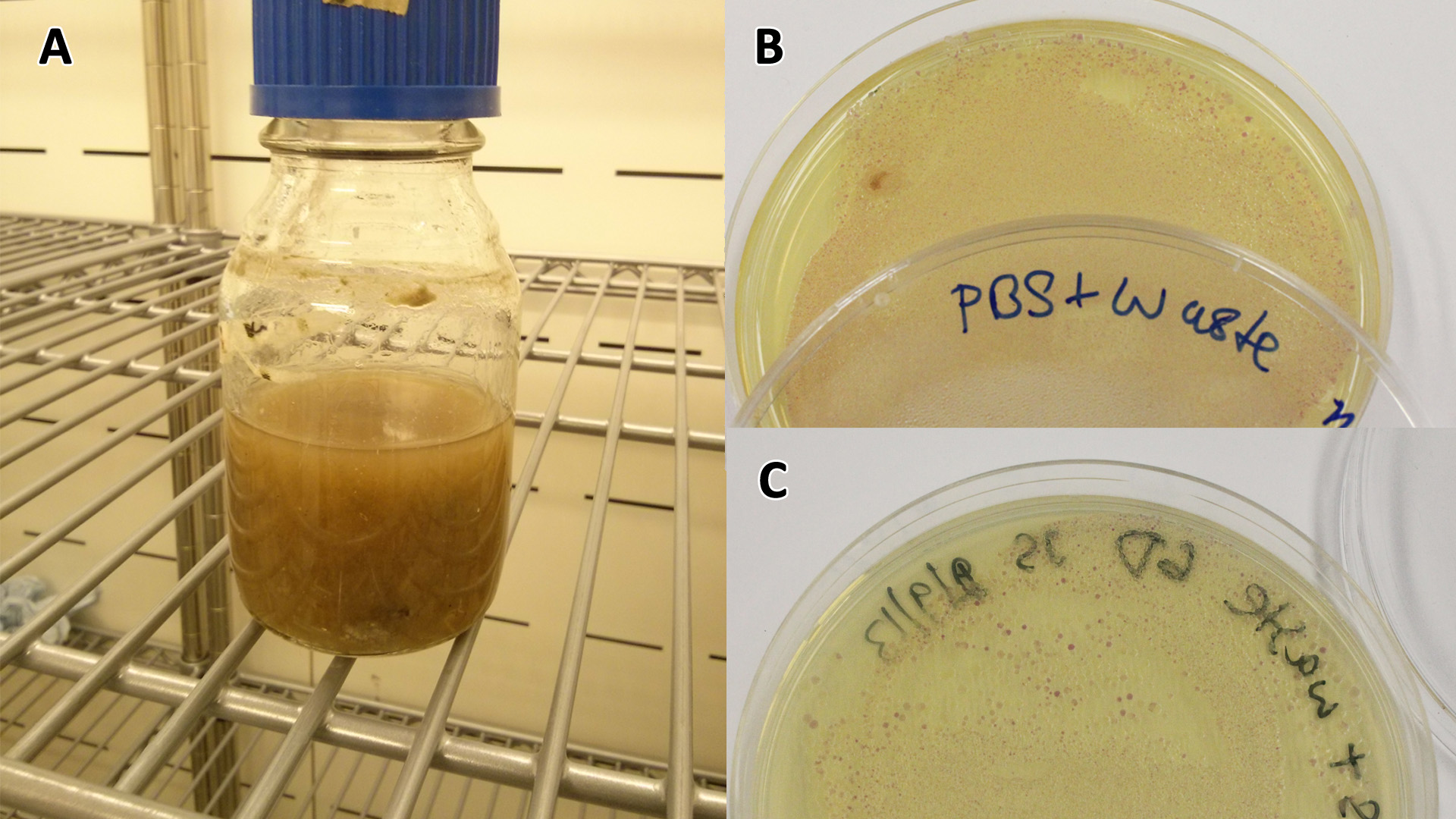
Conclusion: MG1655 E.coli are viable and grow on mixed waste alone. Therefore we have established that our chassis could survive in a mixed waste bio-reactor context, which is validation of our concept to industrially implement our system.
Waste conditioned media
These assays were designed to test whether our chassis, E.coli MG1655 strain could grow with waste conditioned media (WCM) over a period of 24-48 hours. Waste conditioned media is a filter sterilised version of the waste media and was designed for several reasons; Firstly we were unsure whether mixed waste would be toxic to E.coli and hence a less concentrated version may be more suitable and secondly large chunks of waste would prevent accurate OD600 measurements and therefore we decided to filter out the largest chunks.
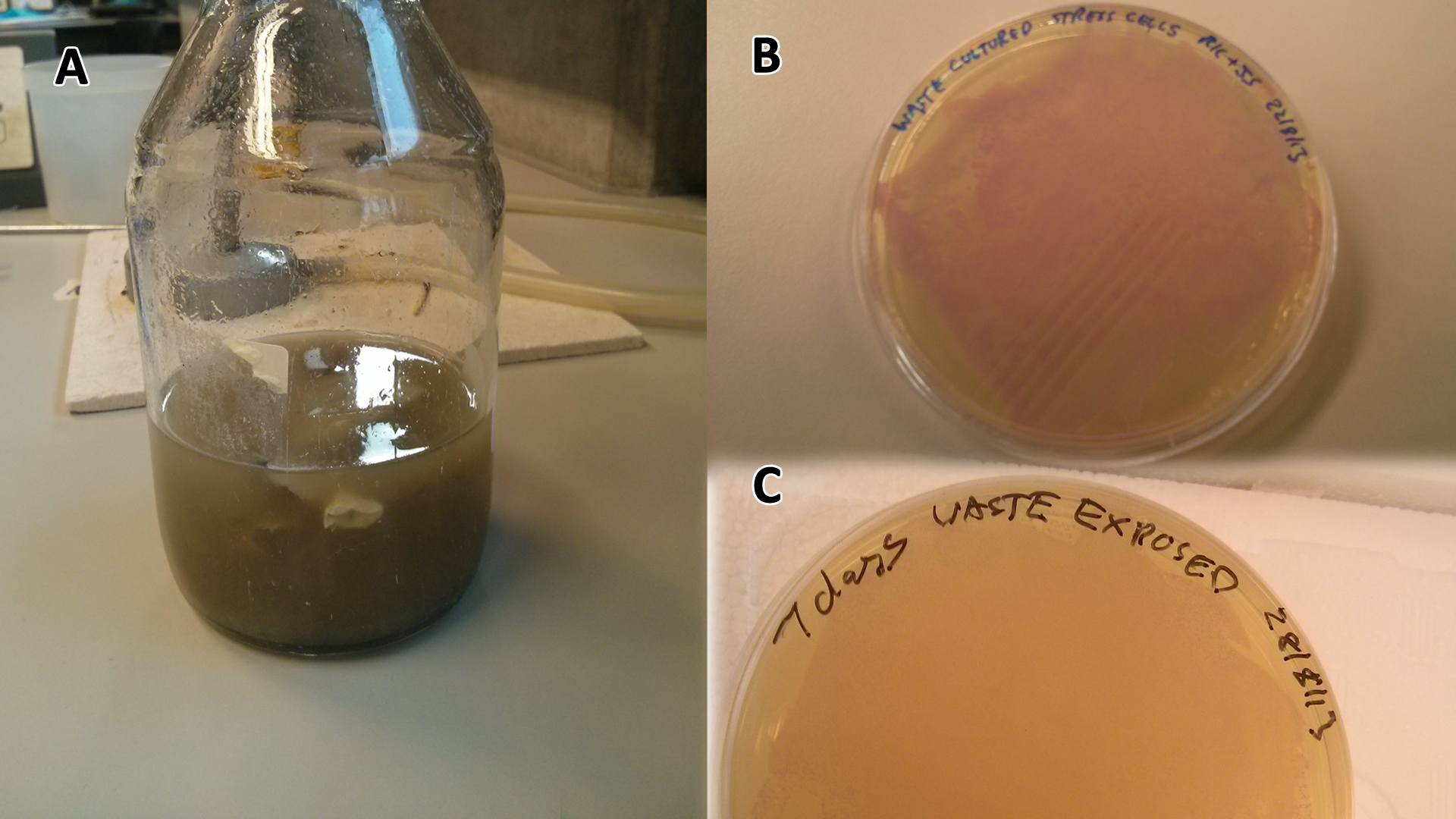
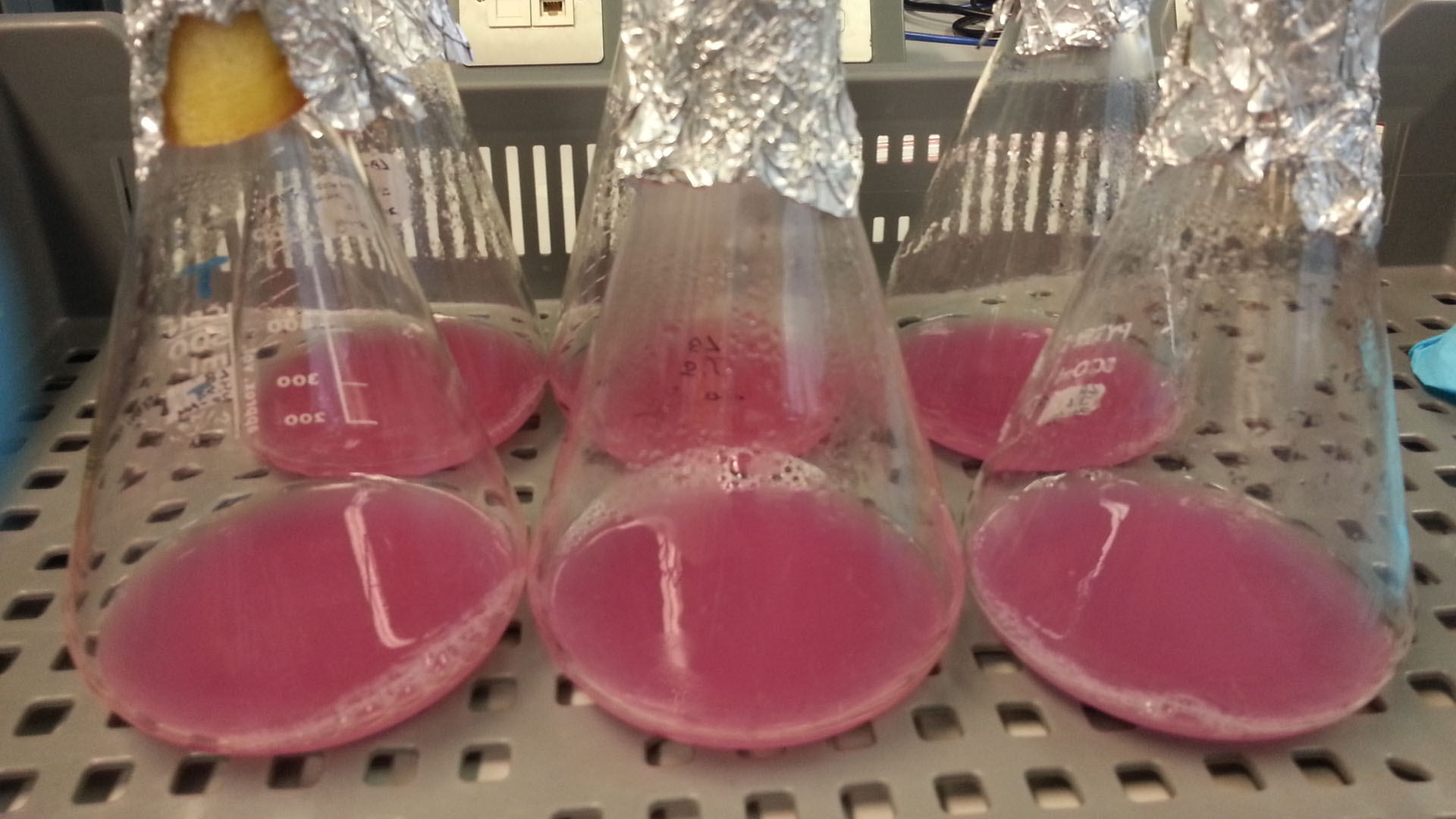 Photograph of waste conditioned media cultures mCherry stress biosensor (BBa_K639003) transformed MG1655 were grown with LB-WCM at 37ºC, with shaking for 48 hours. Figure made by Imperial College London 2013 iGEM. | |
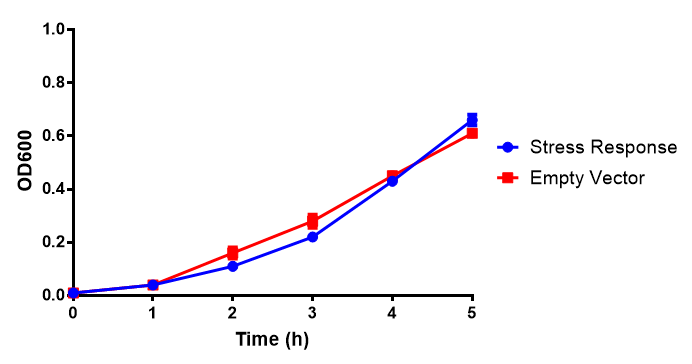
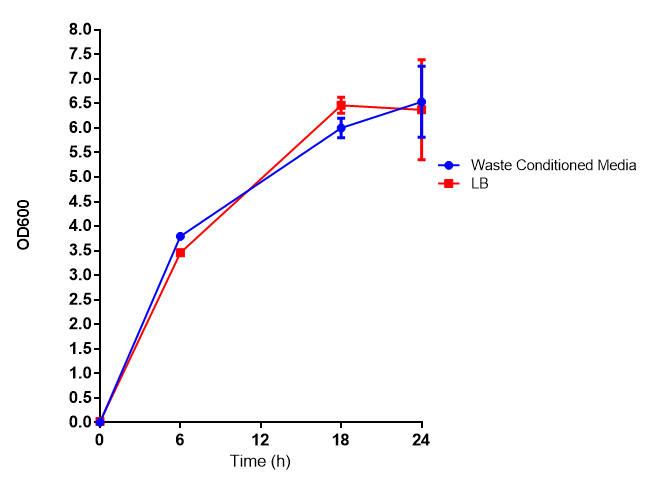 Growth assay in waste conditioned media (WCM).E.coli MG1655 strain transformed with [http://parts.igem.org/Part:BBa_K639003 mCherry stress biosensor.] were grown with LB-based waste conditioned media at 37ºC, with shaking. Error bars represent S.E.M., n=4. Figure made by Imperial College London 2013 iGEM. | |
Conclusion: MG1655 transformed with either empty vector (EV) control or mCherry stress biosensor (BBa_K639003) vector are viable and can grow in waste conditioned media. Therefore waste conditioned media is an appropriate and novel experimental media with which to characterise biobricks within a mixed waste/landfill context. These data are also characterisation of an existing biobrick (BBa_K639003)
Mixed waste degradation: Polyurethane (PUR) ▼
PUR Esterase enzyme activity
Cell lysate assay
Since the PUR Esterases were not secreted, initially the cells were lysed to obtain crude cell extracts in order to test whether the enzymes are active. The Western Blot results showed that the constructs EstCS2 [http://parts.igem.org/Part:BBa_K1149002 BBa_K1149002], PueB [http://parts.igem.org/Part:BBa_K1149004 BBa_K1149004] and PulA [http://parts.igem.org/Part:BBa_K1149006 BBa_K1149006] were being expressed. The cultures expressing these three constructs were grown, lysed by sonication and utilised in a colourimetric assay with the substrate analog para-Nitrophenyl butyrate. The data shows that PUR Esterase EstCS2 [http://parts.igem.org/Part:BBa_K1149002 BBa_K1149002] is definitely active.
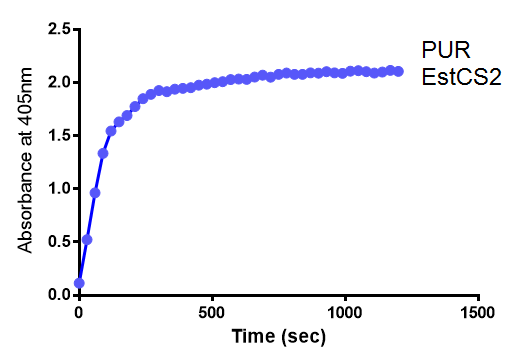
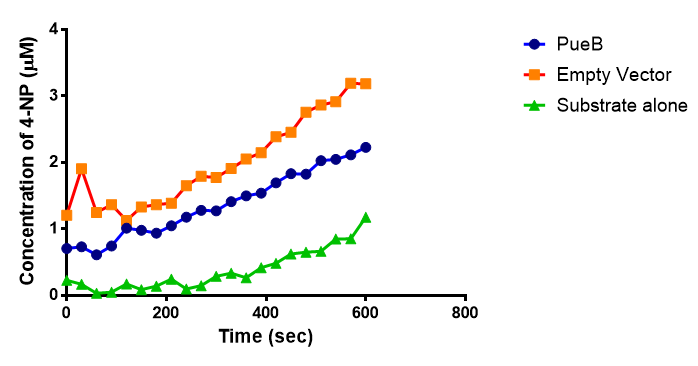
para-Nitrophenyl butyrate (p-NP) is commonly used to indicate an enzyme’s esterase activity. The enzyme cleaves the ester bond and releases the 4-nitrophenol (4-NP), thus causing a colour change from colourless to yellow and an increased absorbance at wavelength 405 nm.
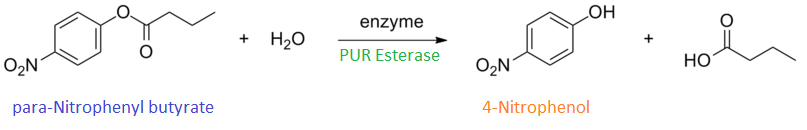
The assay was run in the [http://www.eppendorf.com/int/index.php?sitemap=2.1&action=products&contentid=1&catalognode=87236 Eppendorf BioSpectrometer] was used to automatically read the absorbance of the reaction mixture every 30 seconds. The concentration of 4-Nitrophenol produced from the reaction was calculated using the Beer-Lambert Law; the extinction coefficient of 4-NP is 18,000 M-1 cm-1 at 405 nm.
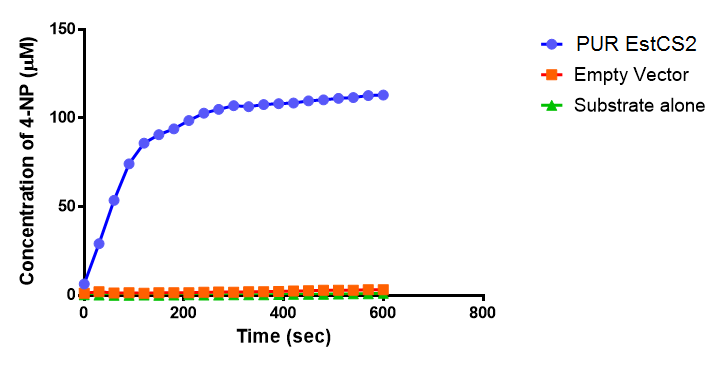
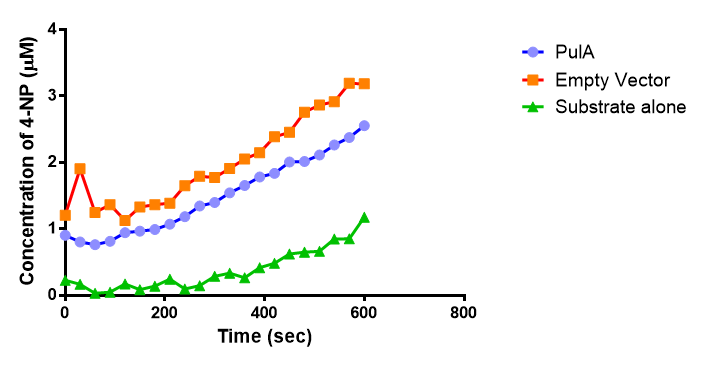
The results below show the concentration of 4-NP produced by the three different PUR esterases and compare them to the Empty Vector and Substrate alone as negative controls.
The above graphs clearly show that PUR Esterase EstCS2 is active and cleaving p-NP. The recorded concentrations of 4-NP, in the presence of this PUR Esterase, are much greater than with the Empty Vector or the Substrate alone. Is PUR Esterase EstCS2 active? Yes!
The enzymes expressed by both PueB and PulA do not appear to have esterase activity for this substrate. There is a tiny increase in 4-NP concentration for PueB, PulA, Empty Vector and Substrate alone. This probably indicates that the substrate is slowly degrading by itself. Other constituents of the cell lysates are likely to be causing a slight increase in p-NP degradation, as PueB, PulA and Empty Vector show higher concentrations of 4-NP than the Substrate alone.
Conclusion: PUR Esterase EstCS2 is active!
Mixed waste degradation: Toxicity ▼
Ethylene glycol
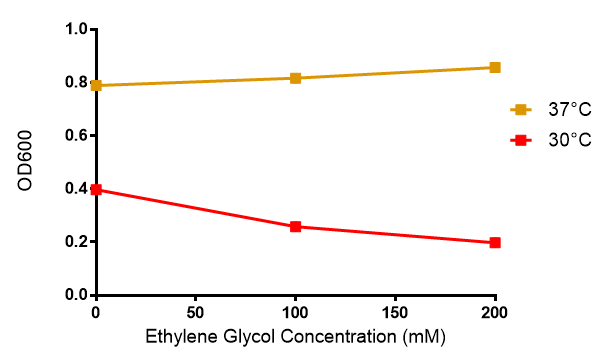
Reduced growth at 30°C likely due to decreased efficiency of MG1655 ethylene glycol break down enzymes. These enzymes (see UC Davis 2012) are endogenously expressed and detoxify Ethylene Glycol.
Bioplastic production ▼
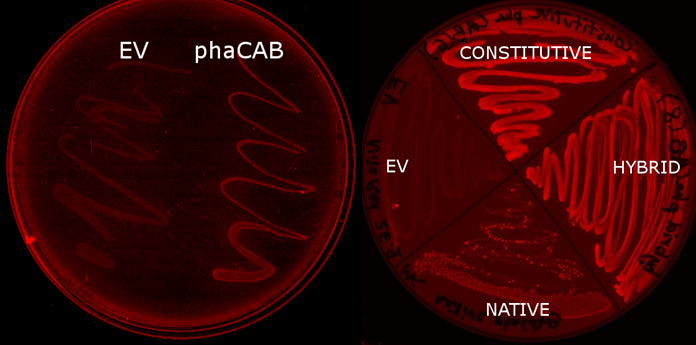
E.coli MG1655 containing the PhaCAB construct Part:BBa_K934001[http://parts.igem.org/wiki/index.php?title=Part:BBa_K934001] accumulates the bioplastic P3HB inside its cells. P3HB can be visualised through staining with Nile Red, which fluoresces strongly in a hydrophobic environment, such as when it binds to the P3HB. However staining will be much lower in cells where P3HB is not present. To see whether P3HB was being produced we grew E.coli containing the phaCAB operon under the native promoter from Ralstonia eutropha. This produced the results we hoped for as seen in the image below.
The P3HB production model indicated that upregulating atoB would lead to an increase in bioplastic production. We changed the promoter before the phaCAB operon to the strong promoter J23104 and a hybrid promoter we created.
The Nile Red plates also provide a preliminary way in which to investigate the effect of changing the promoter to a stronger version. The darker staining of the streaks of the constitutive and hybrid promoter containing constructs suggests the production of higher levels of P3HB in these strains.
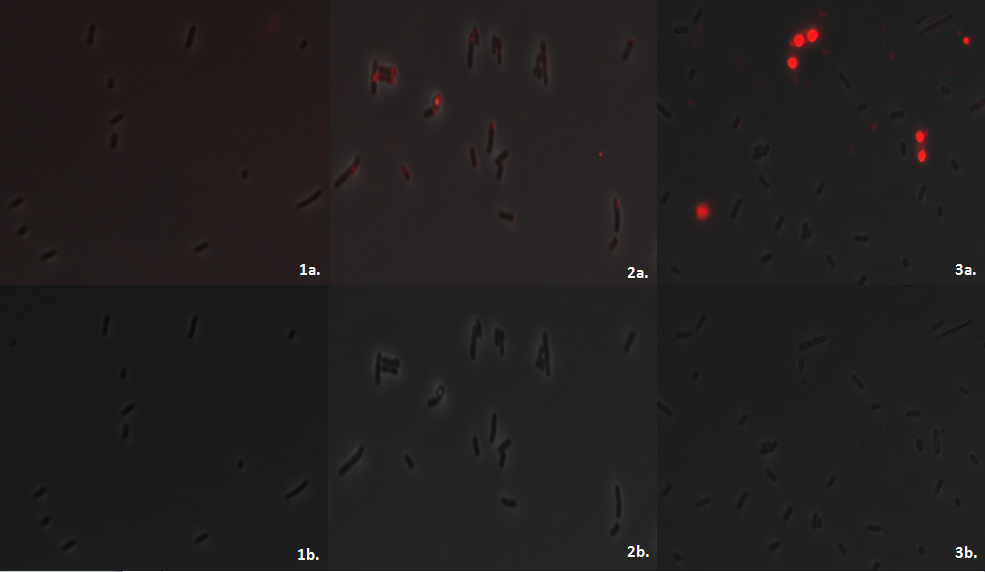
We imaged the cells using the fluorescent microscope to qualitatively understand how much P3HB is produced per cell. As indicated by other authors(1), we saw that P3HB is found in localised granules within the bacteria. Empty vector containing cells show little fluorescence. In those that do, the staining is qualitatively different than in the phaCAB containing strains. Nile Red is also used as a lipid stain and this is indicated by its staining of the membranes of EV cells. In contrast, in phaCAB containing cells, staining is restricted to points within the cells.
Each fluorescent image was created by exciting the stain at 530nm for 100ms. Despite this standard treatment of samples the intensity of fluorescence produced by the cells containing the hybrid promoter is greater than from the constitutive ones. This could indicate a difference in the strength of the promoters which lead to phaCAB expression and therefore the amount of P3HB produced. The images also suggest that not every single E.coli is stained. This may be due to the conditions in which the cells were grown or may be dependent upon the stage in the lifecycle of the E.coli.
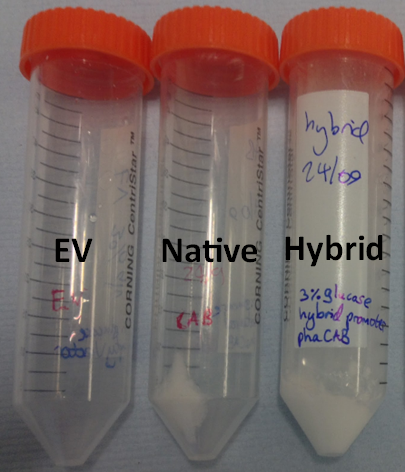
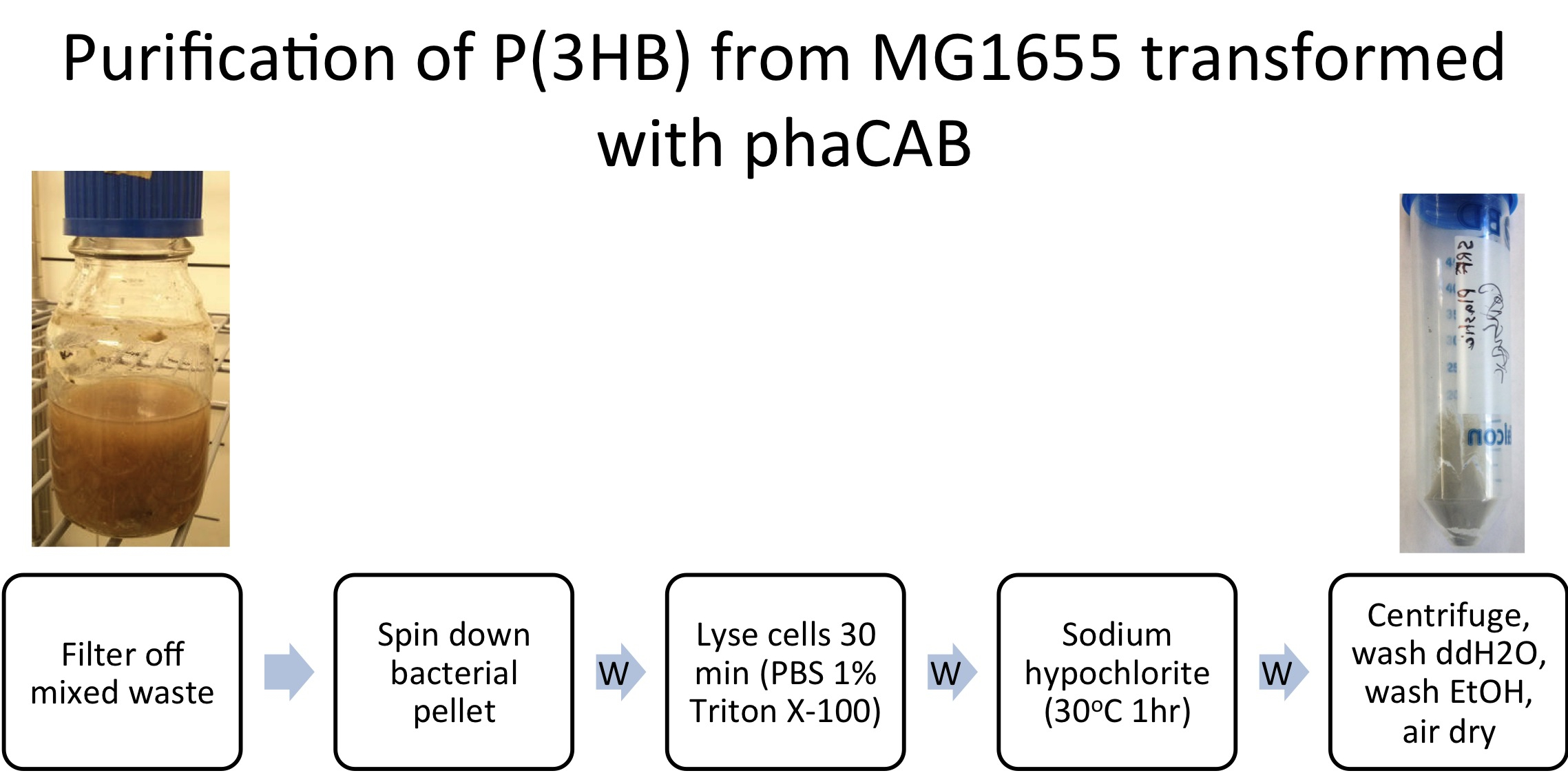
We extracted the P3HB using a simple technique which maintains the chain length of the P3HB while releasing it from the cells. The amount of plastic extracted from the hybrid promoter-phaCAB(BBa_K1149051[http://parts.igem.org/Part:BBa_K1149051]) was much larger compared to the native promoter-phaCAB. All other conditions were kept the same.
Increasing Bioplastic Production ▼
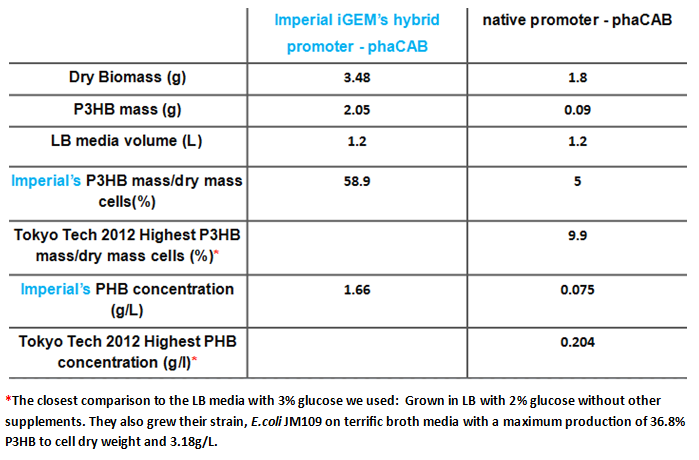
We converted the data into the format with which industry compares P3HB production. These are mass of plastic per dry biomass and mass of plastic per litre of culture media. The strain containing the hybrid promoter had a ten fold higher ratio of plastic to biomass than that produced by the original part BBa_K934001
Comparing the amount of plastic produced by the two constructs is visually striking.
Bioplastic from mixed waste ▼
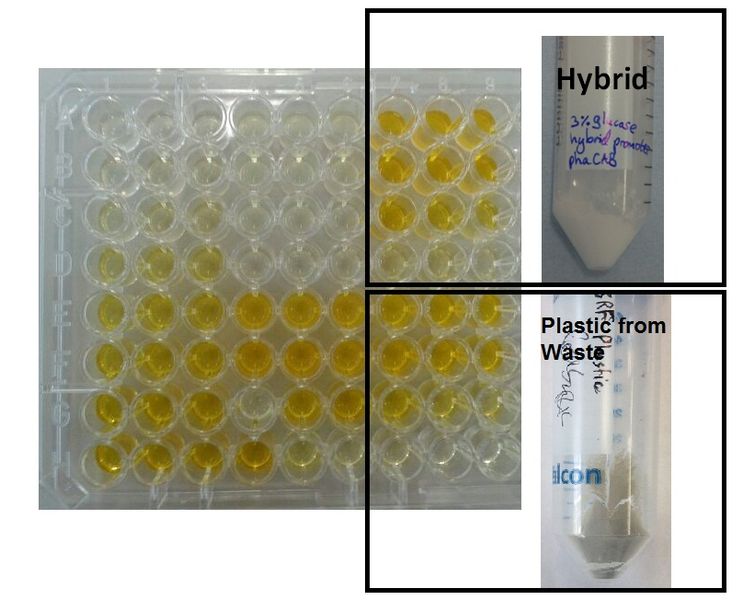
P(3HB) From Mixed Waste
E. coli (MG1655) transformed with native phaCAB (BBa_K934001) were grown with 600ml waste media (12g mixed waste in 600ml M9M- Minimal Media) overnight (16h) at 37oC with shaking. Prior to centrifugation large mixed waste chunks were filtered out. P(HB) was purified as described in Protocols.
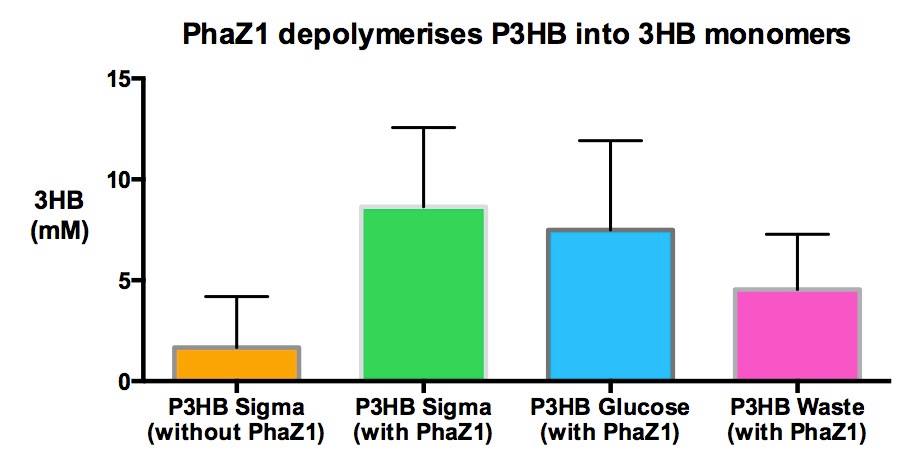
3HB Assay: Confirming production of 3HB
Please see data here.
Extras ▼
Growth assays with different experimental media
In additional to standard LB and minimal media, several novel experimental media were developed in order to characterise Biobricks within a mixed waste/landfill setting. These media were characterised through an examination of pH and through an array of growth assays with the project chassis, E.coli MG1655 strain.
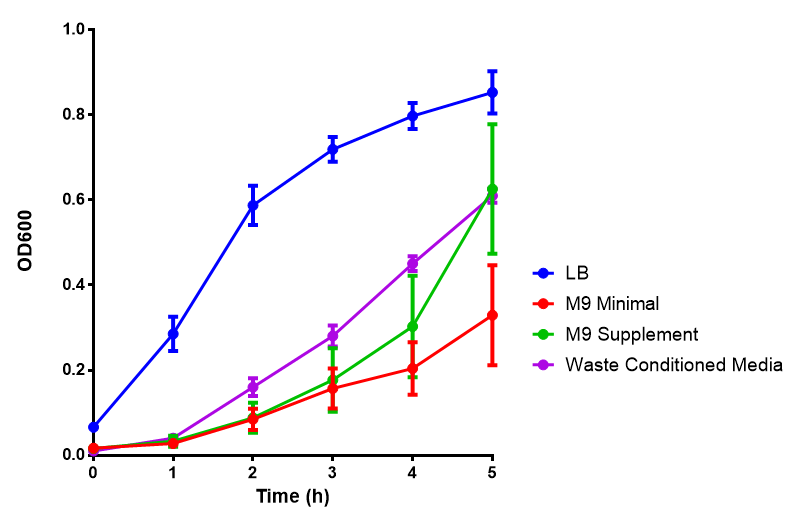 Media characterisation. E.coli strain MG1655 were transformed with a control plasmid and grown in different experimental media over a period of 5 hours. LB media, minimal media (M9M), supplemented minimal media (M9S), as described here or waste conditioned media (WCM), which is made from sterile filtrated mixed waste, see here. OD600 measured, error bars are S.E.M., n=4. Figure made by Imperial College London 2013 iGEM. |
Conclusion: E.coli strain are viable and grow in all of our experimental medias. We have established a novel media that is optimised for characterisation of biobricks within a mixed waste/landfill context.
Protocols ▼
Waste Conditioned Media (WCM)
We added 1g [http://en.wikipedia.org/wiki/Refuse-derived_fuel SRF] (Solid Recovered Fuel) /50 mL LB and then autoclaved the mixture. Once autoclaved, large waste chunks were removed through filter sterilisation (0.2 µm filters) before being used in growth assays as waste conditioned media (WCM). SRF refers to the refuse from recycling facilities that has no value currently and is incinerated to produce power at a cost to the recycling facility, it is composed of 30% plastics while the rest is cellulosic waste
Waste Growth Assay
Overnight (O/N) cultures of MG1655 transformed with (BBa_K639003) were diluted to OD 0.05 in either fresh LB or waste conditioned media and plated into 96 well plates (200 µl/well). OD600 was read at the indicated time points and media only (LB) was taken away as background signal. In addition to this a qualitative assay as performed. 9 mL mCherry bacteria were transferred into 300 mL of autoclaved SRF in LB, which were placed in a shaking incubator at 37°C. After 2 days 200 µL of this solution was plated out on chloramphenicol plates, this was then repeated a week later to gauge whether or not the bacteria were still alive. To show that the MG1655 could grow on waste solely, they were grown in PBS (phosphate buffered saline) and autoclaved waste. In 100 mL PBS + waste, 1 mL MG1655 mCherry E.coli were added. They were then grown in a shaking incubator at 37°C and samples were plated after 2 days and 6 days.
MSDS ▼
Some of the reagents we used were slightly toxic to human. In order to reduce the risks of using toxic reagents, we always wear gloves and labcoats. For some toxic reagents, we performed in fume cupboard. Click on the reagents to see the MSDS.
1. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=W222305&brand=ALDRICH&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Faldrich%2Fw222305%3Flang%3Den Tributyrin]
2. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=458392&brand=ALDRICH&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Faldrich%2F458392%3Flang%3Den Poly(diethylene glycol adipate)].
3. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=19123&brand=SIGMA&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Fsigma%2F19123%3Flang%3Den Nile red].
4. [http://www.sigmaaldrich.com/MSDS/MSDS/PleaseWaitMSDSPage.do?language=&country=GB&brand=SIGMA&productNumber=A3256&PageToGoToURL=http://www.sigmaaldrich.com/catalog/product/sigma/a3256?lang=en®ion=GB Arabinose].
5. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=N9876&brand=SIGMA&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Fsigma%2Fn9876%3Flang%3Den 4-Nitrophenyl butyrate].
6. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=270717&brand=ALDRICH&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Faldrich%2F270717%3Flang%3Den Acetonitrile].
7. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=S8511&brand=SIGMA&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Fsigma%2Fs8511%3Flang%3Den N-Succinyl-Ala-Ala-Pro-Leu p-nitroanilide].
8. [http://www.sigmaaldrich.com/MSDS/MSDS/DisplayMSDSPage.do?country=GB&language=en&productNumber=L3126&brand=SIGMA&PageToGoToURL=http%3A%2F%2Fwww.sigmaaldrich.com%2Fcatalog%2Fproduct%2Fsigma%2Fl3126%3Flang%3Den Lipase from porcine pancreas].
Future work
We are changing the pelB secretion tag to OsmY-fusion partner for better secretion
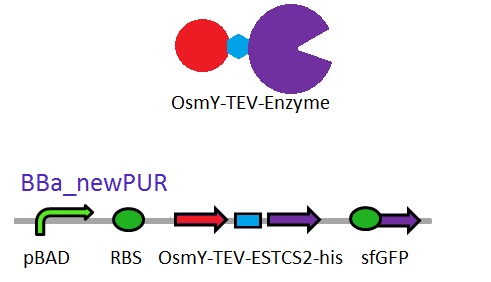
As seen in the Western Blot results, some of the enzymes designed for secretion were not secreted. These proteins may have been stuck in the periplasmic space. Thus we are optimising enzyme secretion by building a new construct with OsmY [http://parts.igem.org/Part:BBa_K892008 BBa_K892008] fusion for secretion. OsmY is naturally secreted from E.coli and will carry our PUR esterase enzyme with. Once in the culture medium, we think that the enzyme is going to be functional in the fusion form but we included a TEV site in between the domains that can be cleaved by TEV protease if needed.
The linker sequence we have designed between the domains is T G S E N L Y F Q G S (ACCGGCAGCGAGAACCTGTACTTCCAAGGCAGC) and includes the 7 amino acids of the TEV site as well as 4 other amino acids for flexibility.
This construct will be a substitute for [http://parts.igem.org/Part:BBa_K1149002 BBa_K1149002] which is the PUR esterase EstCS2 with the pelB seretion tag.
We will show that PUR esterase EstCS2 can break down plastic in emulsions
We have shown that PUR esterase EstCS2 [http://parts.igem.org/Part:BBa_K1149002 BBa_K1149002] is expressed and functional in a colourimetric assay with para-Nitrophenyl butyrate. As it stands, however, it has not degraded the polyurethane in an indicator plate, made with plastic emulsions and LB-agar [http://www.microbialcellfactories.com/content/10/1/41#B16 (4)]. The concentration of the plastics in the plate may be too high for the enzyme to degrade it. Next we will experiment with a variety of different concentrations to see if this has an effect on the enzyme activity. We have already experimented with and identified the mass spectrometry technique required to detect ethylene glycol. We will, therefore, also use mass spectrometry to show that polyurethane can be broken down by our PUR esterase EstCS2.
 "
"