Team:NYMU-Taipei/Project/Inhibition/Id-Nosema
From 2013.igem.org
(Difference between revisions)
| Line 5: | Line 5: | ||
===''N. spp.'' Differentiation Methods=== | ===''N. spp.'' Differentiation Methods=== | ||
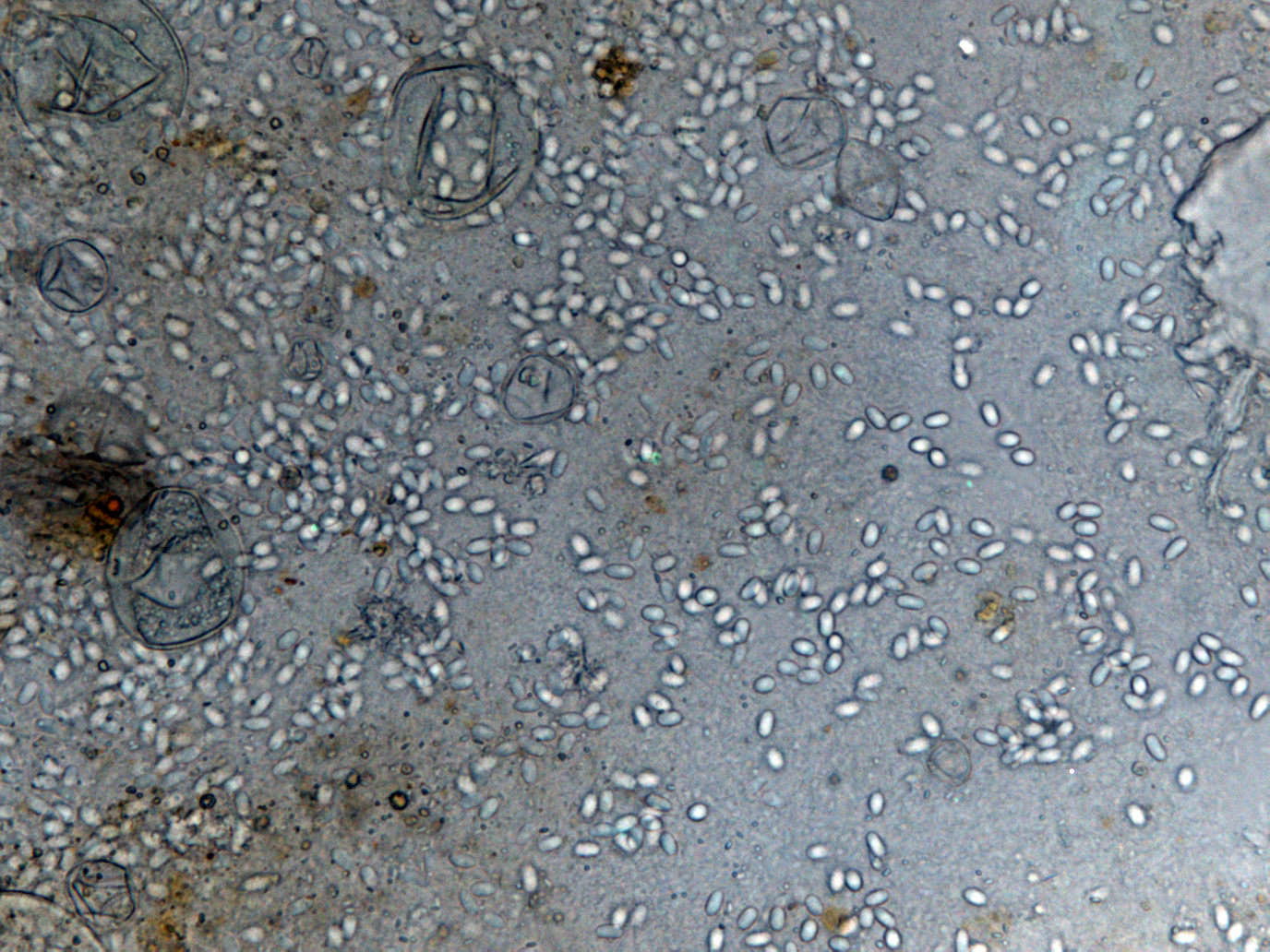
*Light microscopy method: It is based on morphology difference of ''N. apis'' and ''N. ceranae'', revealing that spores of N. ceranae are oval or rod shaped like a rice, and also slightly thinner than N. apis. Light microscopy method is convenient but not scientific enough since ''N. apis'' bears a striking resemblance to ''N. ceranae''. | *Light microscopy method: It is based on morphology difference of ''N. apis'' and ''N. ceranae'', revealing that spores of N. ceranae are oval or rod shaped like a rice, and also slightly thinner than N. apis. Light microscopy method is convenient but not scientific enough since ''N. apis'' bears a striking resemblance to ''N. ceranae''. | ||
| + | {| | ||
| + | |- | ||
| + | |[[File:Nymu-New40x-1.png|thumb|250px|'''Some ''N. ceranae'' spores shown in 400x manification rate.''']]||[[File:Nymu-New40x-3.png|thumb|250px|''' Many ''N. ceranae'' spores shown in 400x manification rate.''']] | ||
| + | |} | ||
| + | |||
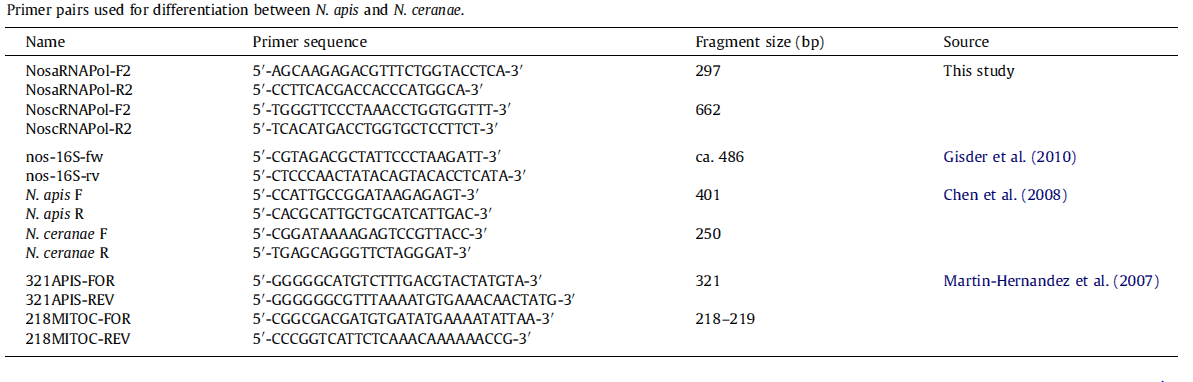
*Molecular methods: Species-specific primers are more accurate than light microscopy method. The problem is that so far nearly all protocols based on species–specific sequence differences are limited to the ribosomal RNA loci, which may lead to confusing results due to polymorphisms in and recombination between the multi-copy 16S rRNA genes, and hence overestimate the rate of Nosema infections. | *Molecular methods: Species-specific primers are more accurate than light microscopy method. The problem is that so far nearly all protocols based on species–specific sequence differences are limited to the ribosomal RNA loci, which may lead to confusing results due to polymorphisms in and recombination between the multi-copy 16S rRNA genes, and hence overestimate the rate of Nosema infections. | ||
| - | |||
| - | |||
[[File:Nymu-All kinds of primers.PNG|thumb|750px|center|'''Source:Molecular differentiation of Nosema apis and Nosema ceranae based on species–specific sequence differences in a protein coding gene '''<br>http://www.ncbi.nlm.nih.gov/pubmed/23352902]] | [[File:Nymu-All kinds of primers.PNG|thumb|750px|center|'''Source:Molecular differentiation of Nosema apis and Nosema ceranae based on species–specific sequence differences in a protein coding gene '''<br>http://www.ncbi.nlm.nih.gov/pubmed/23352902]] | ||
| + | *Our new primers: This year, we created better primers based on recently published, highly conserved and absolutely species–specific sequence differences in protein coding genes. To be more specific, the primers are based on species–specific genes instead of species–specific sequences. As more and more genes of ''Nosema spp.'' are revealed, we are able to choose the genes that are not homologs ''N. apis'' and ''N. ceranae'' for new primers. Our new primers are confirmed by BLAST through nucleotide blast and tblastn using Nucleotide collection (nr/nt) database and Whole-genome shotgun contigis (wgs) database. We propose that our new primers should become the preferred molecular tool for differentiation of ''Nosema spp.'' and thus assist in apiculture. | ||
| + | |||
==Experiment== | ==Experiment== | ||
Revision as of 23:01, 27 September 2013


National Yang Ming University
Contents |
Introduction
The identification of N. apis and N. ceranae is important and still need further research. So far the best differentiation method is the use of species–specific primers; however, the sequence differences are limited to the ribosomal RNA loci, leading to overestimation of Nosema infection rate. Thanks to recent genome study of N. apis and N. ceranae, we proposed novel primers for differentiation of N. spp. based on highly conserved, species–specific protein coding genes rather than species–specific sequences in rRNA genes, which is likely to improve the accuracy of species identification.
Background
N. spp. Differentiation Methods
- Light microscopy method: It is based on morphology difference of N. apis and N. ceranae, revealing that spores of N. ceranae are oval or rod shaped like a rice, and also slightly thinner than N. apis. Light microscopy method is convenient but not scientific enough since N. apis bears a striking resemblance to N. ceranae.
- Molecular methods: Species-specific primers are more accurate than light microscopy method. The problem is that so far nearly all protocols based on species–specific sequence differences are limited to the ribosomal RNA loci, which may lead to confusing results due to polymorphisms in and recombination between the multi-copy 16S rRNA genes, and hence overestimate the rate of Nosema infections.
- Our new primers: This year, we created better primers based on recently published, highly conserved and absolutely species–specific sequence differences in protein coding genes. To be more specific, the primers are based on species–specific genes instead of species–specific sequences. As more and more genes of Nosema spp. are revealed, we are able to choose the genes that are not homologs N. apis and N. ceranae for new primers. Our new primers are confirmed by BLAST through nucleotide blast and tblastn using Nucleotide collection (nr/nt) database and Whole-genome shotgun contigis (wgs) database. We propose that our new primers should become the preferred molecular tool for differentiation of Nosema spp. and thus assist in apiculture.
 "
"