Team:TU-Munich/Results/KillSwitch
From 2013.igem.org
(→Kill Switch) |
(→Level of promoter activity) |
||
| Line 23: | Line 23: | ||
==Level of promoter activity== | ==Level of promoter activity== | ||
| - | |||
| - | |||
[[File:PBI_376_f1.gif|thumb|right|400px| '''Figure 4''': Comparative expression performance of different constitutive mammalian and plant promoters in Physcomitrella patens, [[http://www.ncbi.nlm.nih.gov/pubmed/19021876 Gitzinger et al., 2009]]]] | [[File:PBI_376_f1.gif|thumb|right|400px| '''Figure 4''': Comparative expression performance of different constitutive mammalian and plant promoters in Physcomitrella patens, [[http://www.ncbi.nlm.nih.gov/pubmed/19021876 Gitzinger et al., 2009]]]] | ||
| + | |||
| + | We used the Actin 5 promotor from ''Physcomitrella patens'' [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1159306 BBa_K1159306], which is the strongest known Physco promotor (see Figure 4). The high intensity of nuclease expression might have lead to the early death of the moss. Equipping the Kill Switch with a promotor for a lower level of expression could fix this problem. | ||
==Photosensitivity== | ==Photosensitivity== | ||
Revision as of 23:36, 28 October 2013
Kill Switch
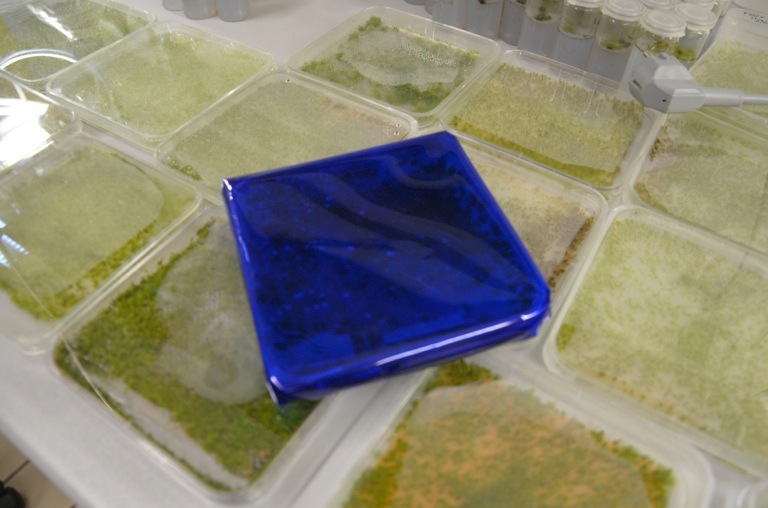
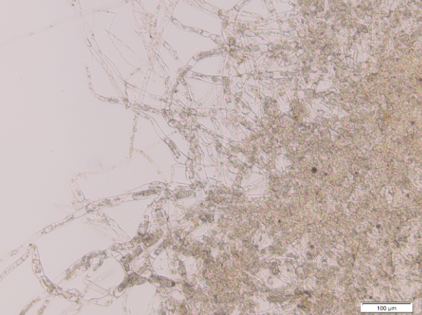
The Kill Switch mechanism is the most complex and ambitious aspect of our project on the protein level. For details on its design and function see here. We were very excited to start experimenting with the killswitch moss. However, when we opened the redlight filter foil, we found none of the allegedly transgenic moss lines had survived. There is a variety of possible reasons for this outcome that need to be discussed in order to improve our approach.
Transformation of the large DNA constructs
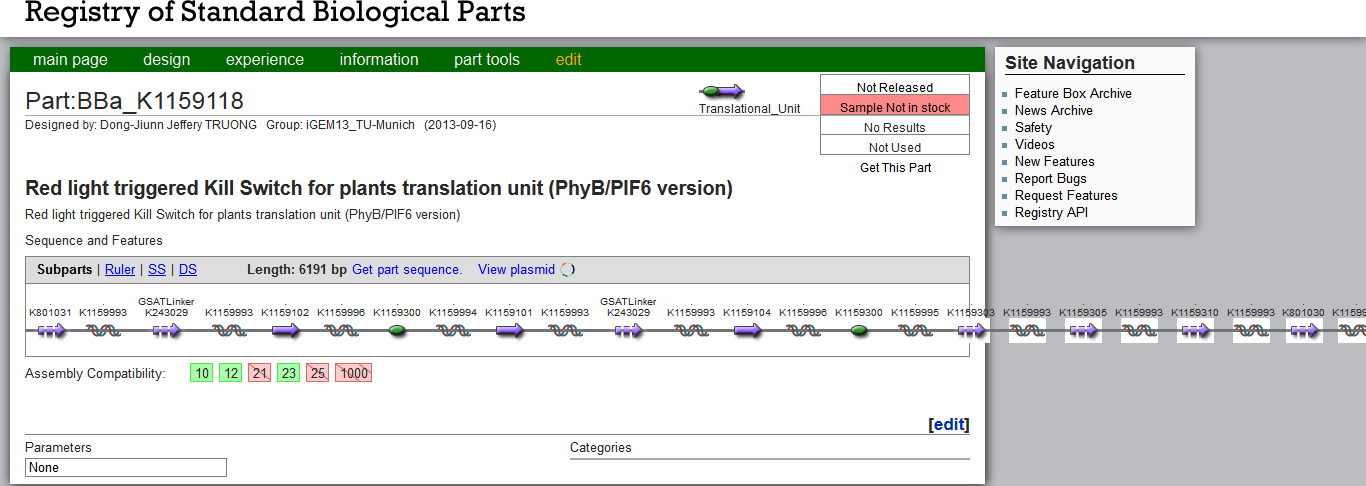
The Kill Switch DNA constructs are very large and even exceed the Part Registry´s frame (see Figure 3). For the PEG-mediated moss transformation, we used plasmids linearized with EcoRI. Contrary to the targeted gene transfer described by [http://www.plant-biotech.net/paper/CurrGenet_2003_hohe.pdf Hohe et al., 2003], our constructs contain no flanking homologuos regions. Our random integration transformation worked successfully for our other contructs, whereas the large Kill Switch construct might need further inquiry for a suitable integration site. Due to its size, the construct might be less stable and require sensitive handling.
We weren´t yet able to express the Kill Switch in E.coli, which is most likely due to codon usage discrepancy. While Physcomitrella patens is not very dicriminating regarding codon usage, the function and expression of proteins designed for Physco is not guaranteed to work in E.coli.
Level of promoter activity
We used the Actin 5 promotor from Physcomitrella patens [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1159306 BBa_K1159306], which is the strongest known Physco promotor (see Figure 4). The high intensity of nuclease expression might have lead to the early death of the moss. Equipping the Kill Switch with a promotor for a lower level of expression could fix this problem.
Photosensitivity
References
http://www.ncbi.nlm.nih.gov/pubmed/19021876 Gitzinger et al., 2009 Functional cross-kingdom conservation of mammalian and moss (Physcomitrella patens) transcription, translation and secretion machineries, Plant Biotechnol J. 2009 Feb;7(2):210
http://www.plant-biotech.net/paper/CurrGenet_2003_hohe.pdf Hohe et al., 2003 An improved and highly standardised transformation procedure allows efficient production of single and multiple targeted gene-knockouts in a moss, Physcomitrella patens, Curr Genet (2004) 44: 339–347
 "
"






AutoAnnotator:
Follow us:
Address:
iGEM Team TU-Munich
Emil-Erlenmeyer-Forum 5
85354 Freising, Germany
Email: igem@wzw.tum.de
Phone: +49 8161 71-4351