Team:NYMU-Taipei/Modeling/MainParts
From 2013.igem.org


Contents |
main part
Function of the parts:
circuit regulators:
• LacI regulated promoter(pLac) : when LacI exists, it will bind to LacI regulated promoter(pLac) and represses the promoter.
• pLux/cI hybrid promoter(plux/cI) : when luxR/AHL exists, it will open pLux/cI hybrid promoter(plux/cI), while cI will repress the promoter. What’s more, plux/cI is cI-dominant, which means when cI exists, the hybrid promoter will be repressed whether luxR/AHL exists or not.
Circuit regulation:
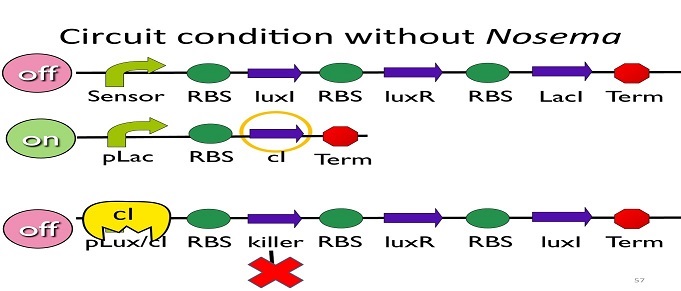
• Circuit condition without Nosema:
Without Nosema Ceranae, the first circuit will not open, and thus, no LacI will be produced. Since there is no LacI binding to pLac, pLac will not be repressed, which means cI will be produced. After that, cI will bind to pLux/cI hybrid promoter, leading the third circuit to be off.
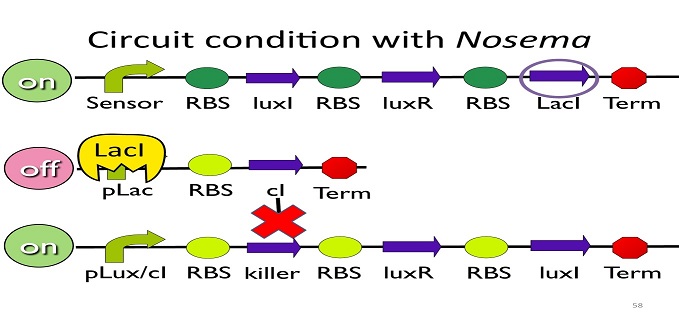
• Circuit condition with Nosema:
When Nosema Ceranae infects the bee, bee’s immune system will be activated, leading to the concentration of ROS (reactive oxygen species) to be high. In response, Bee. coli will enhance the production of oxyR, which forms a complex with ROS and binds to transcription binding site ahead of trxC (an oxyR-activated promoter; namely, the sensor), leading to the first circuit to be on.
Since the first circuit is on, the downstream LacI gene will be produced and bind to pLac, leading to the second circuit to be off.
After the second circuit is closed, no cI is produced, leading to the third circuit to be on (no repressor at all). And then, kill protein will be produced and kill Nosema Ceranae.
•Positive feedback:
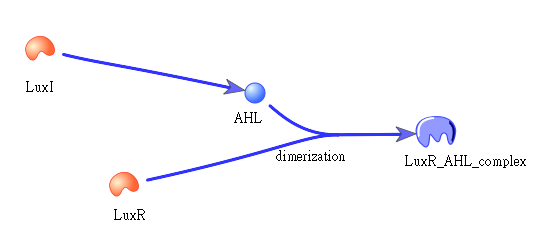
Besides the LuxI and LuxR(which then form a complex called luxR/AHL) produced by first circuit, the third circuit will also produce LuxI and LuxR, which acts as a positive feedback and strongly enhances the production of killer protein.
The picture shows how LuxI will transfer to AHL, which then bind to LuxR to form a dimer:
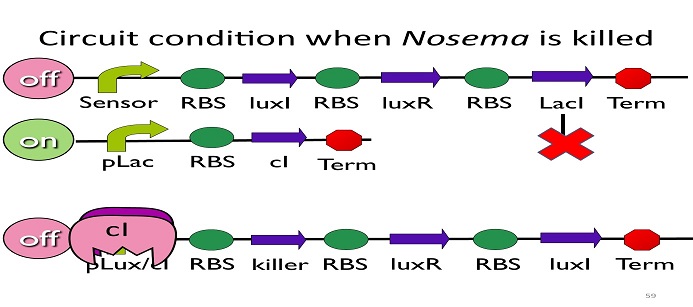
•Circuit condition when Nosema is killed:
After Nosema is killed, the sensor will be off and no lacI will be produced.
Without LacI, the pLac will not be repressed, and the second circuit will be on, leading to the production of cI. After cI binds to pLux/cI hybrid promoter, the third circuit to be off. After that, the overall circuit will switch to the beginning stage when there is no Nosema.
Aims
1. To know how much time it needs from sensing to killing the Nosema after infection
2. To know whether the pathway is effective or not
3. To know the range of kill protein concentration (from the minimal concentration which Is effective to kill Nosema to the maximal concentration which will not do harm to the bees)
It is assumed that AHL is abundant and thus the formation of AHL2LuxR complex is determined only by the concentration of LuxR; CI’s mechanism of binding to the promoter is assumed to act as hill effect form; kill protein will sustain a period of time before it is degraded naturally without any other types of decomposition.
Equations
Equation1:
KdLacI = dissociation constant of CI
nLacI = Hill coefficient of LacI
PoPSpLac = promoter strength of pLac
kdegmRNA = degrading constant of sensor promoter mRNA
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to knowmRNACI production rate and when it can reach the level to translate the desired concentration.
Equation2:
KdROSoxyR = dissociation constant ofROSoxyR
nROSoxyR = Hill coefficient of ROSoxyR
PoPStrxC = promoter strength of trxC
kdegmRNA = degrading constant of sensor promoter mRNA
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to know how trxC promoter strength (in PoPS) influences the production of lacI.
Equation3:
PoPSconstitutive= promoter strength of constitutive promoter (J23102)
N = number of plasmid in a single cell
V = volume of a cell
kdegmRNA = degrading constant of sensor promoter mRNA
The aim of the equation is to know the production rate ofmRNAoxyR, and choose the proper constitutive promoter for boosting OxyR's concentration.
Equation4:
KdROSoxyR = dissociation constant ofROSoxyR
nROSoxyR = Hill coefficient of ROSoxyR
PoPStrxC = promoter strength of trxC
KdLuxRAHL = dissociation constant ofLuxRAHL
nLuxRAHL = Hill coefficient of LuxRAHL
KdcI = dissociation constant of CI
ncI = Hill coefficient of CI
PoPSLuxcI = promoter strength of LuxcI hybrid promoter
kdegmRNA = degrading constant of sensor promoter Mrna
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to know how trxC promoter strength (in PoPS) influences the production of LuxI.
Equation5:
KdROSoxyR = dissociation constant ofROSoxyR
nROSoxyR = Hill coefficient of ROSoxyR
PoPStrxC = promoter strength of trxC
KdLuxRAHL = dissociation constant ofLuxRAHL
nLuxRAHL = Hill coefficient of LuxRAHL
KdcI = dissociation constant of CI
ncI = Hill coefficient of CI
PoPSLuxcI = promoter strength of LuxcI hybrid promoter
kdegmRNA = degrading constant of sensor promoter Mrna
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to know how trxC promoter strength (in PoPS) influences the production of LuxR.
Equation6:
KdLuxRAHL = dissociation constant ofLuxRAHL
nLuxRAHL = Hill coefficient of LuxRAHL
KdcI = dissociation constant of CI
ncI = Hill coefficient of CI
PoPSLuxcI = promoter strength of LuxcI hybrid promoter
kdegmRNA = degrading constant of sensor promoter Mrna
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to knowmRNA of kill protein production rate and when it can reach the level ofthe desired concentration.
Equation7:
KdROSoxyR = dissociation constant ofROSoxyR
nROSoxyR = Hill coefficient of ROSoxyR
PoPStrxC = promoter strength of trxC
KdT7 = dissociation constant of T7
NT7 = Hill coefficient of T7
PoPST7 = promoter strength of T7 promoter
kdegmRNA = degrading constant of sensor promoter Mrna
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to knowmRNA of T7 polymerase production rate and when it can reach the level to translate enough T7 polymerase.
Equation8:
KdT7 = dissociation constant of T7
NT7 = Hill coefficient of T7
PoPST7 = promoter strength of T7 promoter
KdegmRNA = degrading constant of sensor promoter mRNA
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to knowmRNA of enzyme PDC production rate and when it can reach the level to translate enough PDC.
Equation9:
KdT7 = dissociation constant of T7
NT7 = Hill coefficient of T7
PoPST7 = promoter strength of T7 promoter
kdegmRNA = degrading constant of sensor promoter mRNA
N = number of plasmid in a single cell
V = volume of a cell
The aim of the equation is to knowmRNA of enzyme ADH production rate and when it can reach the level to translate enough ADH.
Equation10:
RBS = binding site strength
KdegCI = degrading constant of CI
The aim of the equation is to know the production rate of CI and when it can reach the concentration to repress LuxCI hybrid promoter.
Equation11:
RBS = binding site strength
KdegLacI = degrading constant of LacI
The aim of the equation is to knowLacI production rate and when it can reach the concentration to repress promoter LacI.
'Equation12:
RBS = binding site strength
KAHL = LuxI to AHL rate constant
KdegLuxI = degrading constant of LuxI
The aim of the equation is to know the production rate of LuxI concerning the LuxIAHL reaction and LuxI degrading.
'Equation13:
RBS = binding site strength
KmAHL =AHL to LuxI rate constant
KdegLuxR= degrading constant of LuxR
KonAHL = 2AHL + LuxR to (AHL)2/LuxR complex rate constant
KdegAHL = degrading constant of AHL
The aim of the equation is to know the production rate of LuxR concerning the 2AHL + LuxR to (AHL)2/LuxR reaction , AHL to LuxI reaction, and LuxR, AHL degrading.
'Equation14:
KAHL = LuxI to AHL rate constant
KonAHL = 2AHL + LuxR to (AHL)^2/LuxR complex rate constant
KoffAHL = (AHL)2/LuxR complex to 2AHL + LuxR rate constant
KdegAHL = degrading constant of AHL
The aim of the equation is to know the production rate of AHL concerning the LuxIAHL reaction, 2AHL + LuxR to (AHL)2/LuxR reaction, (AHL)2/LuxR complex to 2AHL + LuxR reaction and AHL degrading.
'Equation15:
RBS = binding site strength
KdegT7 = degrading constant of T7
The aim of the equation is to know the threshold concentration of oxyR to conquer the terminal and when T7 polymerase can reach the required concentration to activate T7 promoter.
'Equation16:
RBS = binding site strength
PoPST7 = promoter strength of T7 promoter
N = number of plasmid in a single cell
V = volume of a cell
KdegPDC = degrading constant of PDC (pyruvate decarboxylase)
The aim of the equation is to know PDC production rate and when it can reach the concentration of ethanol pathway equilibrium.
'Equation17:
RBS = binding site strength
PoPST7 = promoter strength of T7 promoter
N = number of plasmid in a single cell
V = volume of a cell
KdegADH = degrading constant of ADH (Acetaldehyde)
The aim of the equation is to know ADH production rate and when it can reach the concentration of ethanol pathway equilibrium.
'Equation18:
KAHL = LuxI to AHL rate constant
KonAHL = 2AHL + LuxR to (AHL)2/LuxR complex rate constant
KoffAHL = (AHL)2/LuxR complex to 2AHL + LuxR rate constant
KdegAHL = degrading constant of AHL
The aim of the equation is to know the production rate of AHL concerning the LuxIAHL reaction, 2AHL + LuxR to (AHL)2/LuxR reaction, (AHL)2/LuxR complex to 2AHL + LuxR reaction and AHL degrading.
'Equation19:
RBS = binding site strength
Kdegkill = degrading constant of kill
The aim of the equation is to knowkiller protein production rate and when it can reach the effective concentration to killNosema.
'Equation20:
Kpyruvateacetaldehyde = pyruvate→acetaldehyde reaction rate constant
Kacetaldehydeethanol = acetaldehyde→ethanol reaction rate constant
Km = ethanol→acetaldehyde reaction rate constant
KdADH = dissociation constant of ADH
'The aim of the equation is to know ethanol production rate and when it can reach the concentration to kill the spores of Nosema Ceranae.
Parameters:
| Model | Parameter | Description | Value | Unit | Reference |
|---|---|---|---|---|---|
| Sensor | KdegOxyR | OxyR degrading rate | 107 | M-1 x min-1 | Regulation of the OxyR transcription factor by hydrogen peroxide and the cellular thiol—disulfide status |
| KdOxyR* | Dissociation constant of OxyR* | 10-7.33 | M | ||
| KOxyR | OxyR producing rate constant | 5.012 x 1014 | X | ||
| nOxyR* | Hill coefficient of OxyR* | Ranging from 0.75~3.5,depends on what kind of ROS it react with | X | OxyR: A Molecular Code for Redox-Related Signaling | |
| N | Copy number | 887(cells containing 25% plasmid bearing cells) | Single piece |
| |
| Ethanol
| pyruvate | Initial concentration of pyruvate in MG1655 | 1.18 x 2 | g/L | Expression of pyruvate carboxylase enhances succinate production in Escherichia coli without affecting glucose uptake |
| nPDC | Hill coefficient of PDC | 2.1 | - | Purification, characterization and cDNA sequencing of pyruvate decarboxylase Zygosaccharomyces biporus | |
| nADH | Hill coefficient of ADH | at high pH values, 30◦C, n=1; at low temperature, n=3 | - | Evidence for co-operativity in coenzyme binding to tetrameric Sulfolobus
solfataricus alcohol dehydrogenase and its structural basis: fluorescence, kinetic and structural studies of the wild-type enzyme and non-co-operative N249Y mutant | |
| SEIR(exponential)
| b | Infection rate constant of Nosema ceranae to the suspected | 24/75 | Period(days)-1 | |
| r1 | Infection rate constant of K12 to the suspected | 3/20 | Period(days)-1 | 1. Environment protection administration executive yuan of R.O.C Medical bacteriology of J.A.T | |
| r2 | Infection rate constant of K12 to the latent | 3/20 | Period(days)-1 | 2. Environment protection administration executive yuan of R.O.C. 3. Medical bacteriology of J.A.T | |
| e | rate of the latent turns infectious | 1/4 | Period(days)-1 | ||
| u | Death rate of the infected | 1/8 | Period(days)-1 | ||
| k | Rate of intaking capsule | 24/11 | Period(days)-1 | ||
| SEIR(exponential&linear) | S | x(1) | Amount of total population | ||
| E | x(2) | Amount of suspected individuals | |||
| I | x(3) | Amount of individuals in the latent period | |||
| R | x(4) | Amount of infected individuals |
 "
"