Team:TU-Eindhoven/StochasticModel
From 2013.igem.org



Contents |
Decoy Sites
To tune the level of protein expression, we used decoy sites. Decoy sites are tandem repeats of DNA where transcriptional factor can bind to but don't promote gene expression. Therefore it competes with promoter sites and lowers the gene expression rate. LeeDecoyTek-Hyung Lee and Narendra Maheshri, A regulatory role for repeated decoy transcription factor binding sites in target gene expression. Molecular Systems Biology 8, 576 (2012)
The Model
Decoy Sites
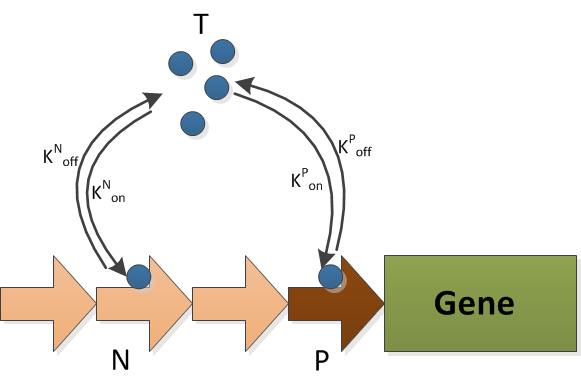

The T is the transcriptional factor, which is FNR. It can binds to either the decoy sites(N) or the promoter(P). The production and degradation of T is included in the FNR model.
Copy Number
Protein Expression Rate
Simulation Result
The input is the concentration of FNR protein. This result is taken from the part of FNR protein and converted from micro molar concentrations to number of molecules.

The simulated result of decoy site binding ratio
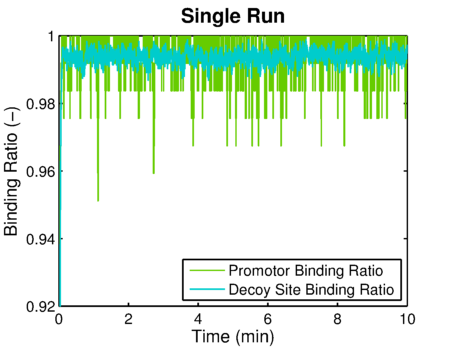 The simulated result of promoter binding ratio
The simulated result of promoter binding ratio
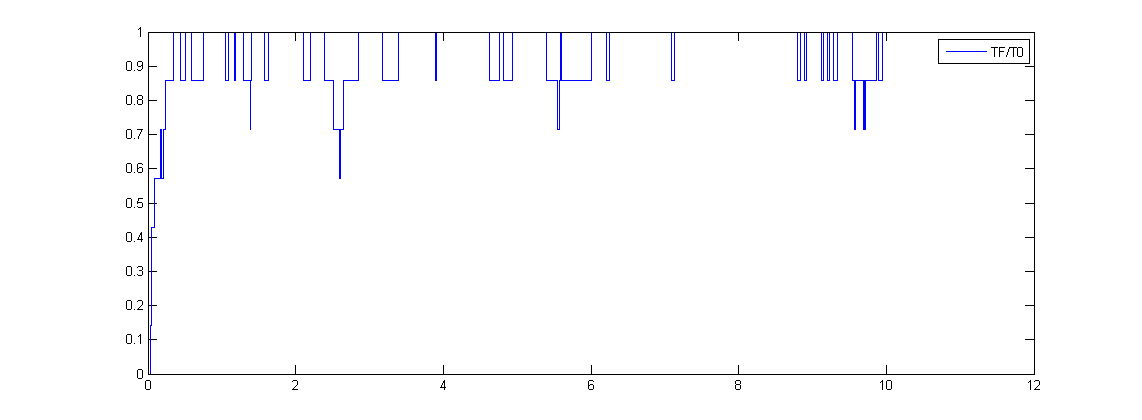 This would be the input for the modeling of protein generation. We assume the protein production rate is linear to the promoter binding ratio.LeeDecoy
This would be the input for the modeling of protein generation. We assume the protein production rate is linear to the promoter binding ratio.LeeDecoy
References
 "
"