Team:Leeds/Results
From 2013.igem.org
Results from the lab so far. Please note, many of the images have been uploaded at high resolution, and are best viewed from their file page. Access this by clicking the image. We may later introduce a gallery of the images to maximise the screen usage, but this is dependent upon Coffee-Induced coding by the Code Monkey.
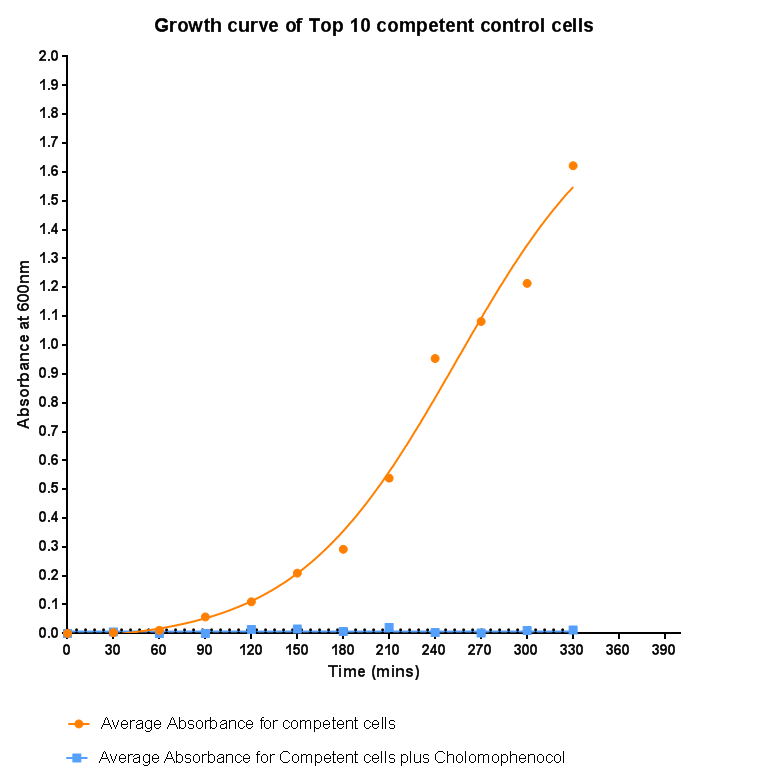
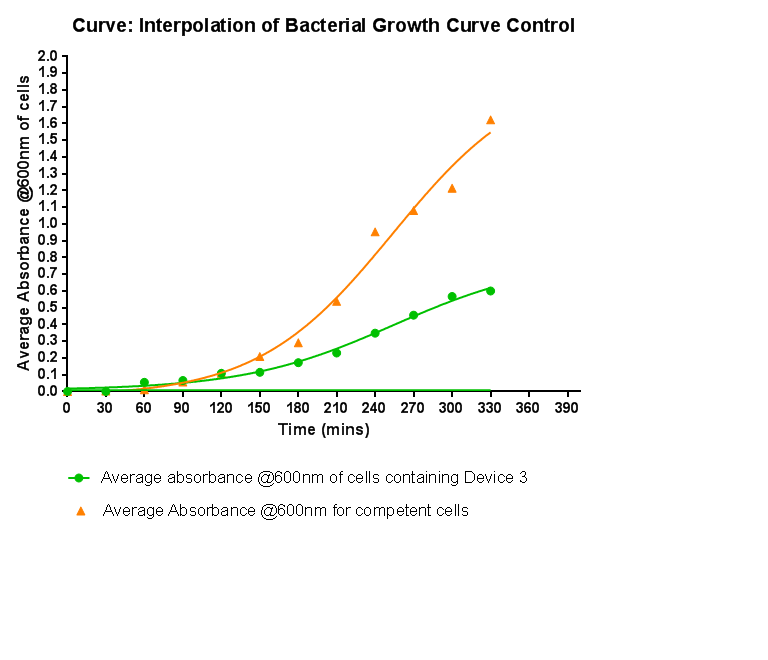
Bacterial growth curvesWe are monitoring the growth of our bacteria containing our devices to see if the genes we inserted have any effect on bacterial growth. Control Bacterial Growth CurveIn this experiment we used our chosen expression cells and monitored their growth on SB media both in the presence of chloramphenicol and without chloramphenicol. As the bacteria do not contain our plasmid that codes for chloramphenicol resistance, we didn't expect them to grow when it was present. The graph produced from the absorbance measurements taken over time, shows the growth of the bacteria. It has a clear lag and log phase when the bacteria are grown in SB media only. But as expected, the graph shows that the bacteria do not grow in the presence of chloramphenicol. This graph will be used as a control and other growth curves of bacteria containing our device will be compared to it in order to determine whether the genes we inserted cause an effect on cell growth.
Agarose Gels For DigestionsWhen a digestion of DNA is undertaken, it is important that an Agarose gel is run afterwards to ensure it was successful before moving onto the next step in the experimental procedure. Digestion of plasmid containing Green Fluorescent ProteinHere we digested the part [http://parts.igem.org/Part:BBa_K081012:Design K081012] with restriction enzyme EcoR1 and ran a gel to make sure the fragment(s) we obtained were of the correct length. This gel shows the bands obtained by EcoR1 digestion of BBa-K081012. There are 100bp ladders and 1kbp ladders in the first two wells. This gel was used to work out the size of the fragments obtained by plotting the results on a calibration graph.
DNA Ladder Calibration GraphFrom this graph we can work out that one fragment is around 2800bp long, while the other appears to be 4000bp long - far longer than the plasmid at 2858bp long. Through digestion, it has been linearised by a single EcoR1 digest. The fragment corresponds to what we expect so our digest was successful, despite not all of the plasmid being digested.
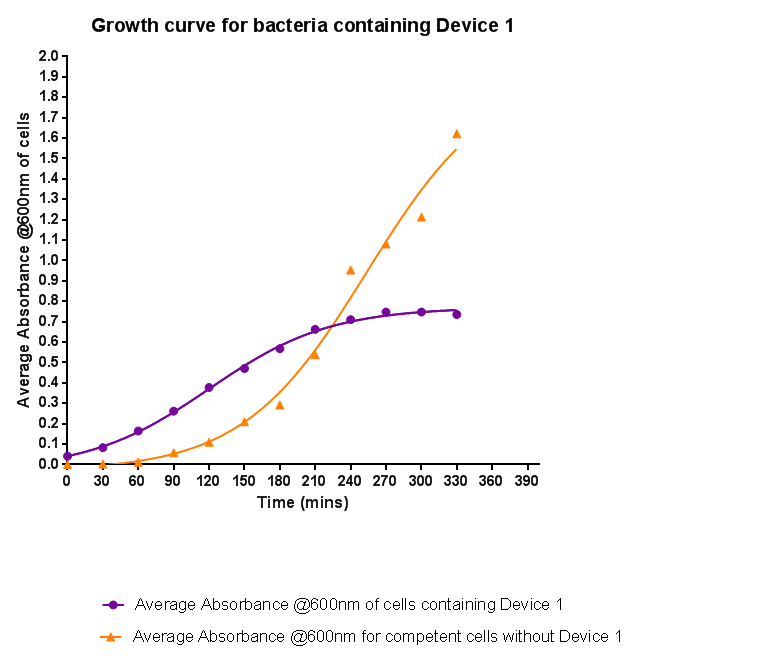
Characterisation of Device 1
Growth Curve
Restriction Digests
Plate Imaging
Membrane Stress - Hydrophobic Stress
Membrane Stress - Changes in pH
Membrane Stress- Atomic Force Microscopy
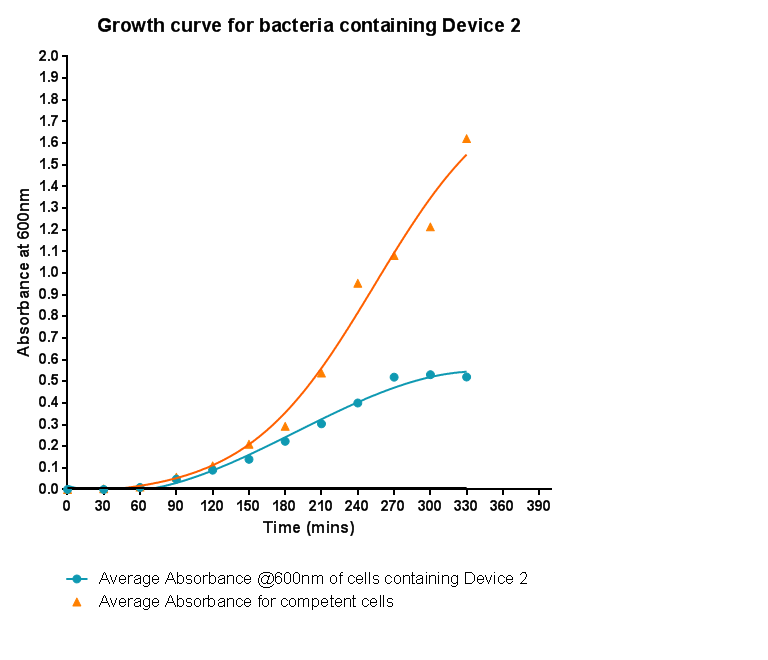
Characterisation of Device 2Growth Curve
Restriction Digests
Plate Imaging
Binding assay-washing beads
Binding assay-Confocal Microscopy
Characterisation of Device 3
Growth Curve
Restriction Digests
Plate Imaging
Binding Assay-Beads cause fluorescence?
| |||||||
 |
| ||||||

| |||||||

| |||||||
 "
"