Team:Evry/Modelmeta1
From 2013.igem.org
| Line 39: | Line 39: | ||
<h2>Materials and methods</h2> | <h2>Materials and methods</h2> | ||
<p> | <p> | ||
| - | + | The iron-FUR complex is simply formed that way:<br/> | |
| + | <img src="https://static.igem.org/mediawiki/2013/7/72/Reg1.png"/><br/> | ||
| + | We reduced this equation to:<br/> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/e/e8/Reg2.png"/><br/> | ||
| + | Which is not annoying, since we just have to divide our [FeFur] by to to get the real complex concentration.<br/> | ||
| + | We can easily write down both the formation (v) and the dissociation (v') speed:<br/> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/f/fe/Reg3.png"/><br/> | ||
| + | We chose to model the iron input in the bacteria using a linear function of the external iron concentration <i>Ferext</i>, the factor <i>p</i> being the cell-wall permeability for iron.<br/> | ||
| + | The FUR on the other hand is produced by the bacteria. Its evolution can also be considered as linear, using a mean production rate <i>Fur0</i>.<br/> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/5/51/Regfer.png"/><br/> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/6/68/Regfur.png"/><br/> | ||
| + | In this model, we only track the free Fe-FUR complex and not those attached to a FUR Binding Site. As <i>LacI</i> is the number of inhibited LacI, we can use this number to express how much Fe-FUR does bind to a FBS per unit of time.<br/> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/6/6d/Regfefur.png"/> | ||
</p> | </p> | ||
Revision as of 15:37, 22 October 2013
Sensor Model
Introduction
In order to determine our iron-coli's enterobactin production rate, we first have to know how much time our bacteria will take to sense the ambient iron. So, this first part of the Enterobactin production model focuses on the synthetic sensing system our team implemented in the bacteria.
Observations
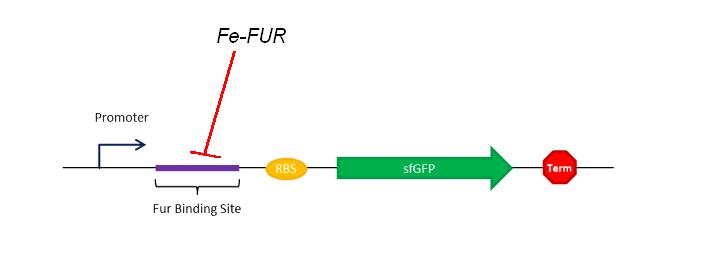
As shown on the Figure 1, our sensing system relies on a FUR Binding Site (FBS). In order to easily model the sensing delay and analyse its results, the first construction is composed of a GFP placed right after the FBS.
Goals
Our goal in this part of the model is to create a generic FBS-related sensing model so that:
- We can determine the iron-sensing delay of our bacteria
- The model can can be reused by other iron related projects
Materials and methods
The iron-FUR complex is simply formed that way:

We reduced this equation to:

Which is not annoying, since we just have to divide our [FeFur] by to to get the real complex concentration.
We can easily write down both the formation (v) and the dissociation (v') speed:

We chose to model the iron input in the bacteria using a linear function of the external iron concentration Ferext, the factor p being the cell-wall permeability for iron.
The FUR on the other hand is produced by the bacteria. Its evolution can also be considered as linear, using a mean production rate Fur0.


In this model, we only track the free Fe-FUR complex and not those attached to a FUR Binding Site. As LacI is the number of inhibited LacI, we can use this number to express how much Fe-FUR does bind to a FBS per unit of time.

Results
Conclusion
Models and scripts
References:
 "
"