Team:Calgary/Project/PostRegionals
From 2013.igem.org
Iaingeorge (Talk | contribs) |
|||
| Line 7: | Line 7: | ||
<h1>Post-Regionals</h1> | <h1>Post-Regionals</h1> | ||
| - | <h2>For our <span class="blue">Sensor</span> we...</h2> | + | <h2>For our <span class="blue">Sensor</span> we are able to show individual components and our final system working!</h2> |
| + | |||
| + | <h2>So do These Coils Actually Bind?</h2> | ||
| + | |||
| + | <p> After putting in gratuitous effort to build parts containing these coils and successfully purifying these proteins we wanted to determine if the E and K coils interacted with each other. In order to characterize coil-coil interaction we performed an immunoprecipitation (IP) assay. We built a GFP with an E-coil (<a href=” http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189014”> BBa_K1189014</a>) and we also built TALE-B with a K-coil (<a href=” http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189014”>BBa_K1189030</a>). To characterize the binding of the coils we pulled down with either an immunoglobulin G antibody (IgG) that serves as a negative control or with GFP antibody. The idea behind this experiment is to pull down the E coil which is fused to GFP with a GFP antibody and then probe with a anti-his antibody which recognizes the his tag on the TALE fused to the K coil. Upon interaction between the E and K coils we will see an output at approximately 86 kDa when probed with a His-antibody as seen in Figure 7. Our test groups included the coils by themselves and both the E and K coils put together in solution. As seen in Figure 7 a band appears only when we pull E and K coil with a GFP and probe with an anti-his antibody indicating the presence of both GFP and TALE in the elution solution indicating that the coils interact with each other. </p> | ||
| + | <figure> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/a/a6/HimikaIP_Ucalgary.png"> | ||
| + | <figcaption><b>Figure 7.</b> Immunoprecipitation assay showing that the E and K coils interact with each other. The first three lanes show E-coil, K coil and E-coil with K-coil pulled down with IgG (control) antibody. The last three lanes were pulled down with a anti-GFP antibody and probed with a anti-his antibody. The E-coil is fused to GFP and the K-coil is fused to TALE with a his tag. Therefore, if we pull down with GFP and probe with his-antibody a band would appear the size of TALE. This indicates that both the GFP and the TALE are present and that is possible upon coil-coil interaction. | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | <h2>How does the use of Coils versus Direct Fusions of TALEs Affect our Prussian Blue reporter?</h2> | ||
| + | <p>After successfully confirming that we could convert our own ferritin proteins that were produced from the parts we constructed (<a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K1189018">BBa_K1189018,</a> <a | ||
| + | href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K1189021">BBa_K1189021</a>) into Prussian blue ferritin the next step was to evaluate how the design of our parts could potentially affect the reporter activity of our Prussian blue ferritin. Based on the spatial modelling performed by our team it was suggested that assembly of the ferritin nanoparticle with TALE proteins directly fused was highly unlikely. This is because the TALE proteins are significantly larger than the ferritin subunits. Their size would likely result in steric hindrance and prevent the assembly of the full ferritin protein. In order to test the predictions put forward by our modelling we ensured that our protein samples were balanced in order to have the same number ferritin cores in each sample. The catalytic activity of these proteins was then compared. From the data gathered we saw that the Prussian blue ferritin with fused coils (even if TALES are additionally bound to the ferritin via coils) was more effective as a reporter than having the TALE proteins directly fused to the ferritin nanoparticle (Figure 19). The results from this experiment suggest that the predictions made by our model were correct. Using coils however alleviates this issue as these coils are small and would not interfere in the ferritin self-assembly but can be used to attach our TALES to create the FerriTALE. </p> | ||
| + | |||
| + | <figure> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/0/0d/UCalgary2013TRPBFAssayNoColour.png" alt="Recombinant Prussian Blue FerritinMole Balanced" width="465" height="480"> | ||
| + | <figcaption> | ||
| + | <p><b>Figure 19.</b> Samples of our parts that were converted to Prussian Blue ferritin were mole balanced in order to ensure that the same number of effective ferritin cores are present in every sample. Additionally the ferritin-coil fusion was incubated with the TALE-coil fusion part in order to allow their binding for a separate trial. Negative controls include unconverted recombinant ferritin, bovine serum albumin and a substrate only control. Samples were incubated with a TMB substrate solution for 10 minutes at a pH of 5.6. Absorbance readings were taken at the 10 minute time-point at a wavelength of 650 nm. An ANOVA (analysis of variants) was performed upon the values to determine that there was statistical difference in the data gathered (based off of three replicates). A t-test was then performed which determined that the * columns are significantly different from the ** column (p=0.0012). Neither * column is significantly different from each other (p=0.67).</p> | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | <figure> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/c/c7/UCalgary2013TRBetalactamasecolourpsd.png" alt="Beta-lactamase Visual Assay" width="432" height="599"> | ||
| + | <figcaption> | ||
| + | <p><b>Figure 11.</b>Change in pH catalyzed by TALE A linked to β-lactamase (<a href=" http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189031">BBa_K1189031</a>) using benzylpenicillin. Bromothymol blue was used to keep track of this colour change. Absorbance readings were taken at 616 nm every 30 seconds. Different amounts of TALE A linked to β-lactamase (<a href=" http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189031">BBa_K1189031</a>) were tested. Commercial β-lactamase was used as a positive control. Negative controls included were bovine serum albumin, β-lactamase without the substrate and the substrate by itself.</p> | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | <h2>Can we successfully capture our DNA with our detectors with specificity and report it?</h2> | ||
| + | <figure> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/6/68/UCalgary2013TRBetalactamaseassay.png" alt="Beta-lactamase Catalytic Assay" width="800" height="442"> | ||
| + | <figcaption> | ||
| + | <p><b>Figure 12.</b>Change in pH catalyzed by TALE A linked to β-lactamase (<a href=" http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189031">BBa_K1189031</a>) using benzylpenicillin. Bromothymol blue was used to keep track of this colour change. Absorbance readings were taken at 616 nm every 30 seconds. Different amounts of TALE A linked to β-lactamase (<a href=" http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189031">BBa_K1189031</a>) were tested. Commercial β-lactamase was used as a positive control. Negative controls included were bovine serum albumin, β-lactamase without the substrate and the substrate by itself.</p> | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | |||
| + | <figure> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/5/54/TALE_B_and_TALE_A_B-lac_DNA_capture_assay.png" alt="TALE DNA Capture Assay" width="800" height="600"> | ||
| + | <figcaption> | ||
| + | <p><b>Figure 6: </b> TALE capture assay was done with TALE B (<a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189001"> | ||
| + | <span class="Green"><b> | ||
| + | BBa_K1189001 | ||
| + | </b></span> | ||
| + | </a >)and TALE A B-lac fusion (<a href=" http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189031"> | ||
| + | <span class="Green"><b> | ||
| + | BBa_K1189031 | ||
| + | </b></span> | ||
| + | </a >) and DNA (<a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K1189006"> | ||
| + | <span class="Green"><b> | ||
| + | BBa_K1189006 | ||
| + | </b></span> </a >) | ||
| + | with both target sequences. If capture is successful, the B-lac is present in the well giving a colour change from pink to yellow when subjected to benzylpenicillin substrate solution within 20 minutes. The only wells that change colour are the first four wells which contain TALE B, specific DNA, and TALE A β-lactamase fusion and our positive control wells which are the controls for our fusion TALE A β-lactamase protein and our positive recombinant β-lactamase. All our other controls including our test using a non-specific sequence of DNA remained pink . This preliminary characterization data demonstrates that the TALEs are able to bind to DNA with specificity. Additionally it also shows that our system of capturing DNA with two detector TALEs and then subsequent reporting of the DNA’s presence works. | ||
| + | </figcaption> | ||
| + | </figure> | ||
| + | |||
| + | <p>This assay shows that we can <b> capture our target DNA </b> with two detector TALEs with <b>specificity </b>. Additionally, <b> we can report whether that DNA has been captured</b> and is present in the sample, which is a very important concept for our sensor system. </p> | ||
| + | |||
<h2>For our <span class="orange">Human Practices</span> we...</h2> | <h2>For our <span class="orange">Human Practices</span> we...</h2> | ||
Revision as of 01:19, 29 October 2013
Post-Regionals
Post-Regionals
For our Sensor we are able to show individual components and our final system working!
So do These Coils Actually Bind?
After putting in gratuitous effort to build parts containing these coils and successfully purifying these proteins we wanted to determine if the E and K coils interacted with each other. In order to characterize coil-coil interaction we performed an immunoprecipitation (IP) assay. We built a GFP with an E-coil ( BBa_K1189014) and we also built TALE-B with a K-coil (BBa_K1189030). To characterize the binding of the coils we pulled down with either an immunoglobulin G antibody (IgG) that serves as a negative control or with GFP antibody. The idea behind this experiment is to pull down the E coil which is fused to GFP with a GFP antibody and then probe with a anti-his antibody which recognizes the his tag on the TALE fused to the K coil. Upon interaction between the E and K coils we will see an output at approximately 86 kDa when probed with a His-antibody as seen in Figure 7. Our test groups included the coils by themselves and both the E and K coils put together in solution. As seen in Figure 7 a band appears only when we pull E and K coil with a GFP and probe with an anti-his antibody indicating the presence of both GFP and TALE in the elution solution indicating that the coils interact with each other.

How does the use of Coils versus Direct Fusions of TALEs Affect our Prussian Blue reporter?
After successfully confirming that we could convert our own ferritin proteins that were produced from the parts we constructed (BBa_K1189018, BBa_K1189021) into Prussian blue ferritin the next step was to evaluate how the design of our parts could potentially affect the reporter activity of our Prussian blue ferritin. Based on the spatial modelling performed by our team it was suggested that assembly of the ferritin nanoparticle with TALE proteins directly fused was highly unlikely. This is because the TALE proteins are significantly larger than the ferritin subunits. Their size would likely result in steric hindrance and prevent the assembly of the full ferritin protein. In order to test the predictions put forward by our modelling we ensured that our protein samples were balanced in order to have the same number ferritin cores in each sample. The catalytic activity of these proteins was then compared. From the data gathered we saw that the Prussian blue ferritin with fused coils (even if TALES are additionally bound to the ferritin via coils) was more effective as a reporter than having the TALE proteins directly fused to the ferritin nanoparticle (Figure 19). The results from this experiment suggest that the predictions made by our model were correct. Using coils however alleviates this issue as these coils are small and would not interfere in the ferritin self-assembly but can be used to attach our TALES to create the FerriTALE.

Figure 19. Samples of our parts that were converted to Prussian Blue ferritin were mole balanced in order to ensure that the same number of effective ferritin cores are present in every sample. Additionally the ferritin-coil fusion was incubated with the TALE-coil fusion part in order to allow their binding for a separate trial. Negative controls include unconverted recombinant ferritin, bovine serum albumin and a substrate only control. Samples were incubated with a TMB substrate solution for 10 minutes at a pH of 5.6. Absorbance readings were taken at the 10 minute time-point at a wavelength of 650 nm. An ANOVA (analysis of variants) was performed upon the values to determine that there was statistical difference in the data gathered (based off of three replicates). A t-test was then performed which determined that the * columns are significantly different from the ** column (p=0.0012). Neither * column is significantly different from each other (p=0.67).

Figure 11.Change in pH catalyzed by TALE A linked to β-lactamase (BBa_K1189031) using benzylpenicillin. Bromothymol blue was used to keep track of this colour change. Absorbance readings were taken at 616 nm every 30 seconds. Different amounts of TALE A linked to β-lactamase (BBa_K1189031) were tested. Commercial β-lactamase was used as a positive control. Negative controls included were bovine serum albumin, β-lactamase without the substrate and the substrate by itself.
Can we successfully capture our DNA with our detectors with specificity and report it?
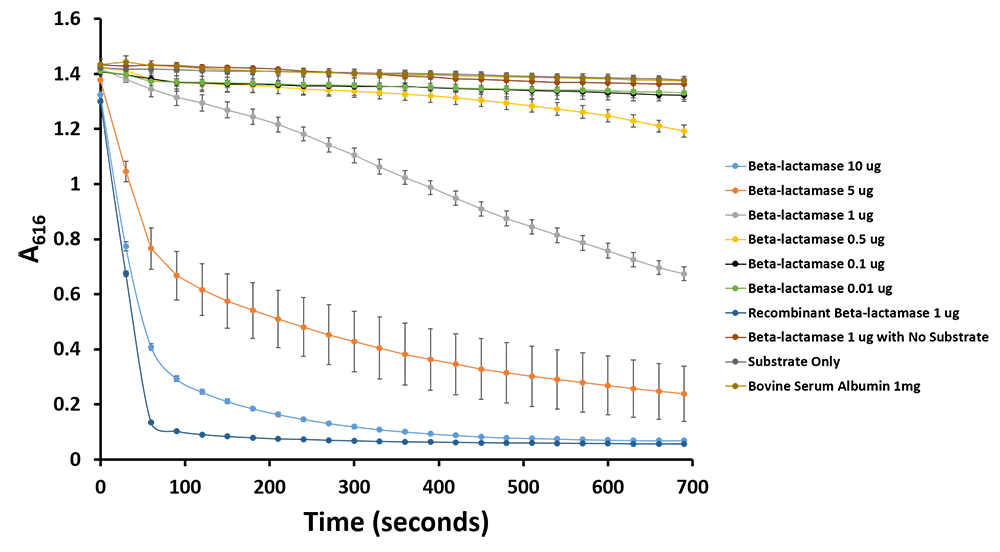
Figure 12.Change in pH catalyzed by TALE A linked to β-lactamase (BBa_K1189031) using benzylpenicillin. Bromothymol blue was used to keep track of this colour change. Absorbance readings were taken at 616 nm every 30 seconds. Different amounts of TALE A linked to β-lactamase (BBa_K1189031) were tested. Commercial β-lactamase was used as a positive control. Negative controls included were bovine serum albumin, β-lactamase without the substrate and the substrate by itself.

Figure 6: TALE capture assay was done with TALE B ( BBa_K1189001 )and TALE A B-lac fusion ( BBa_K1189031 ) and DNA ( BBa_K1189006 ) with both target sequences. If capture is successful, the B-lac is present in the well giving a colour change from pink to yellow when subjected to benzylpenicillin substrate solution within 20 minutes. The only wells that change colour are the first four wells which contain TALE B, specific DNA, and TALE A β-lactamase fusion and our positive control wells which are the controls for our fusion TALE A β-lactamase protein and our positive recombinant β-lactamase. All our other controls including our test using a non-specific sequence of DNA remained pink . This preliminary characterization data demonstrates that the TALEs are able to bind to DNA with specificity. Additionally it also shows that our system of capturing DNA with two detector TALEs and then subsequent reporting of the DNA’s presence works.
This assay shows that we can capture our target DNA with two detector TALEs with specificity . Additionally, we can report whether that DNA has been captured and is present in the sample, which is a very important concept for our sensor system.
 "
"
