Team:Uppsala/results
From 2013.igem.org
| Line 59: | Line 59: | ||
<li><a href="https://2013.igem.org/Team:Uppsala/metabolic-engineering">Metabolic engineering</a> | <li><a href="https://2013.igem.org/Team:Uppsala/metabolic-engineering">Metabolic engineering</a> | ||
<ul> | <ul> | ||
| - | <li><a href="https://2013.igem.org/Team:Uppsala/p-coumaric-acid"> | + | <li><a href="https://2013.igem.org/Team:Uppsala/p-coumaric-acid">p-Coumaric acid</a></li> |
<li><a href="https://2013.igem.org/Team:Uppsala/resveratrol">Resveratrol</a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/resveratrol">Resveratrol</a></li> | ||
<li><a href="https://2013.igem.org/Team:Uppsala/lycopene">Lycopene</a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/lycopene">Lycopene</a></li> | ||
| Line 80: | Line 80: | ||
<li><a href="https://2013.igem.org/Team:Uppsala/modeling" id="list_type1"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/6/63/Uppsala2013_Modeling.png"></a> | <li><a href="https://2013.igem.org/Team:Uppsala/modeling" id="list_type1"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/6/63/Uppsala2013_Modeling.png"></a> | ||
<ul> | <ul> | ||
| - | <li><a href="https://2013.igem.org/Team:Uppsala/P-Coumaric-acid-pathway"> | + | <li><a href="https://2013.igem.org/Team:Uppsala/P-Coumaric-acid-pathway">Kinetic model</a></li> |
<li><a href="https://2013.igem.org/Team:Uppsala/modeling-tutorial">Modeling tutorial </a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/modeling-tutorial">Modeling tutorial </a></li> | ||
| + | |||
| + | <li><a href="https://2013.igem.org/Team:Uppsala/toxicity-model">Toxicity model</a></li> | ||
</ul></li> | </ul></li> | ||
<li><a href="https://2013.igem.org/Team:Uppsala/parts" id="list_type2"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/e/eb/Uppsala2013_parts.png"></a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/parts" id="list_type2"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/e/eb/Uppsala2013_parts.png"></a></li> | ||
| Line 100: | Line 102: | ||
<li><a href="https://2013.igem.org/Team:Uppsala/public-opinion">Public opinion </a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/public-opinion">Public opinion </a></li> | ||
<li><a href="https://2013.igem.org/Team:Uppsala/Outreach">High school & media </a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/Outreach">High school & media </a></li> | ||
| - | + | <li><a href="https://2013.igem.org/Team:Uppsala/bioart">BioArt</a></li> | |
| + | <li><a href="https://2013.igem.org/Team:Uppsala/LactonutritiousWorld">A LactoWorld</a></li> | ||
| + | <li><a href="https://2013.igem.org/Team:Uppsala/killswitches">Killswitches</a></li> | ||
| + | <li><a href="https://2013.igem.org/Team:Uppsala/realization">Patent</a></li> | ||
</ul></li> | </ul></li> | ||
<li><a href="https://2013.igem.org/Team:Uppsala/attribution" id="list_type4"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/5/5d/Uppsala2013_Attributions.png"></a></li> | <li><a href="https://2013.igem.org/Team:Uppsala/attribution" id="list_type4"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/5/5d/Uppsala2013_Attributions.png"></a></li> | ||
Revision as of 19:53, 27 October 2013
Results and achievement
Achievements
Results
P-coumaric acid
We managed to clone out and biobrick tyrosine ammonia lyase and verify the biobrick by sequencing. Also we did succeful characterization on this part, showing that it works as expected. We managed to express our enzyme and detect it in a western blot, and also detect our metabolite in both spectrophotometry and hplc. The biobrick was characterized in E. coli d5 alpha and E. coli nissle. For detailed information about the characterization methods, see the protocol section. To read more about P-coumaric acid, click here

2. TAL with Cp8 promoter
3. TAL with J23110 promoter
4. Negative control
Figure 1:SDS-page and western blot. Expression of Tyrosine ammonia lyase with constitutive promoters. The negative control is empty, showing that there is no natural protein in E. coli with the same attributes.
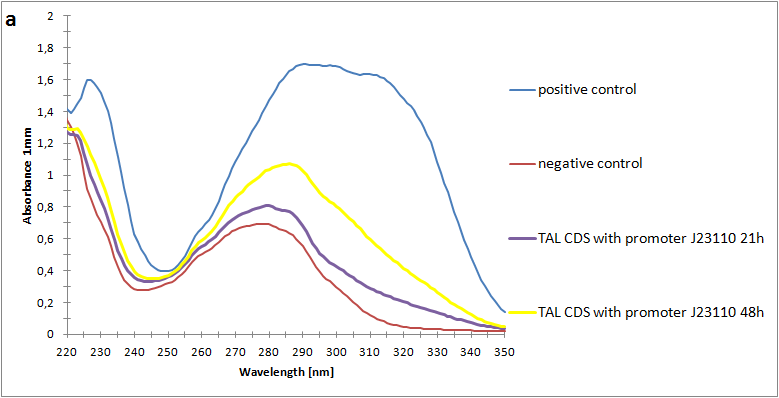
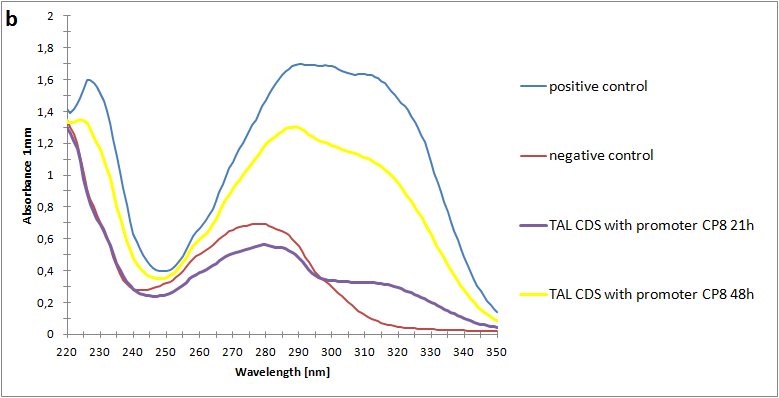

Figure 2:Absorbance spectra of extracts collected from bacterial cultures. Samples were collected 21 h and 48 h after 30 °C incubation. The negative control is an extract from a strain with no TAL gene on transformed plasmid. The positive control is an extract a culture of the same strain as the negative control but with added p-coumaric acid to a concentration of 500 µM before extraction. P-coumaric acid absorbance spectra has two peaks. The one around 305 nm is preferable to detect because of background from bacteria. (a) Spectra from the strain with TAL CDS with promoter J23110.(b) Spectra from the strain with TAL CDS with promoter CP8. (c) Spectra from the strain with TAL CDS with promoter J23101.
 | Figure 3: Graph showing the HPLC result of a sample prepared from E. coli expressing tyrosine ammonia lyase. Reverse phase HPLC with a C18 matrix was used. The peak for p-coumaric acid can be seen ~10 min, as shown by the standard sample below. |
 | Figure 4: Graph showing the HPLC result of a sample standard with p-coumaric acid. |
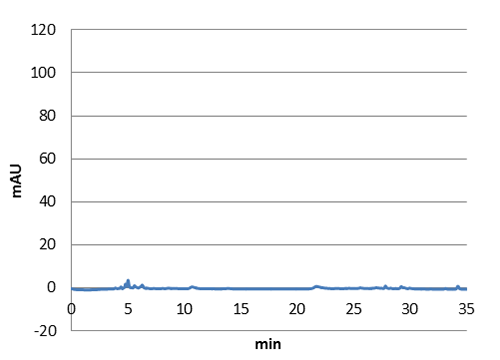 | Figure 5: E. coli culture injected to the hplc without our biobrick tyrosine ammonia lyase. Here we can see that there is originally no peak at 10 minutes. |
Resveratrol
Although we managed to clone out and sequence verify the genes for resveratrol production, we have had some problems in the characterization. The results are unclear, and we did not have time for further investigations.For detailed information about the characterization methods, see the protocol section and to read more about resveratrol, click here

Positive control -> 1, Stilbene synthase -> 2
4Cl-STS translational fusion expressed in E. coli

Resveratrol standard

Resveratrol standard scaled
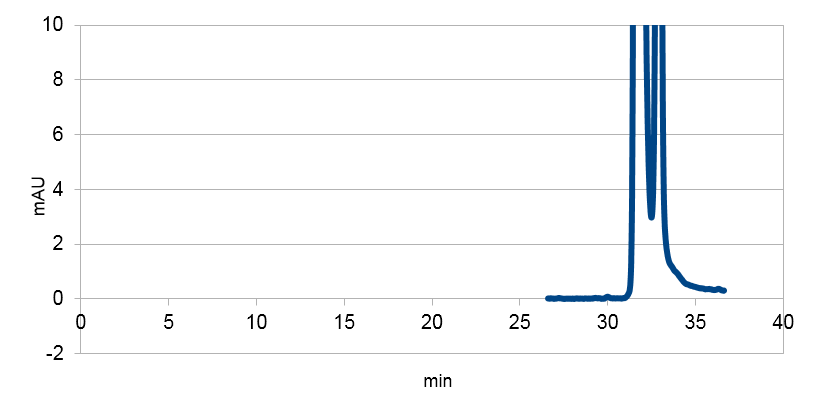
Lysed bacterial culture without plasmid of assembly

Shuttle vector
We constructed two shuttle vectors able to replicate and provide resistance in both Lactobacillus and E. coli. We have shown that they can both be used in E. coli for assembly and subcloning as well as transforming them to Lactobacillus. To read more about the shuttle vector, click here

Promoters
We have provided a constitutive promoter collection with 6 promoters with different strength measured by a fluorescence-activated cellsorting machine (FACS) in RPU, a unit based on the strength of the promoter J23101 in standard parts. These promoters works in both gram-negative bacterium E. coli and gram-positive bacterium Lactobacillus and are therefore also believed to be universal for all prokaryotic organisms. To read more about promoters, click here

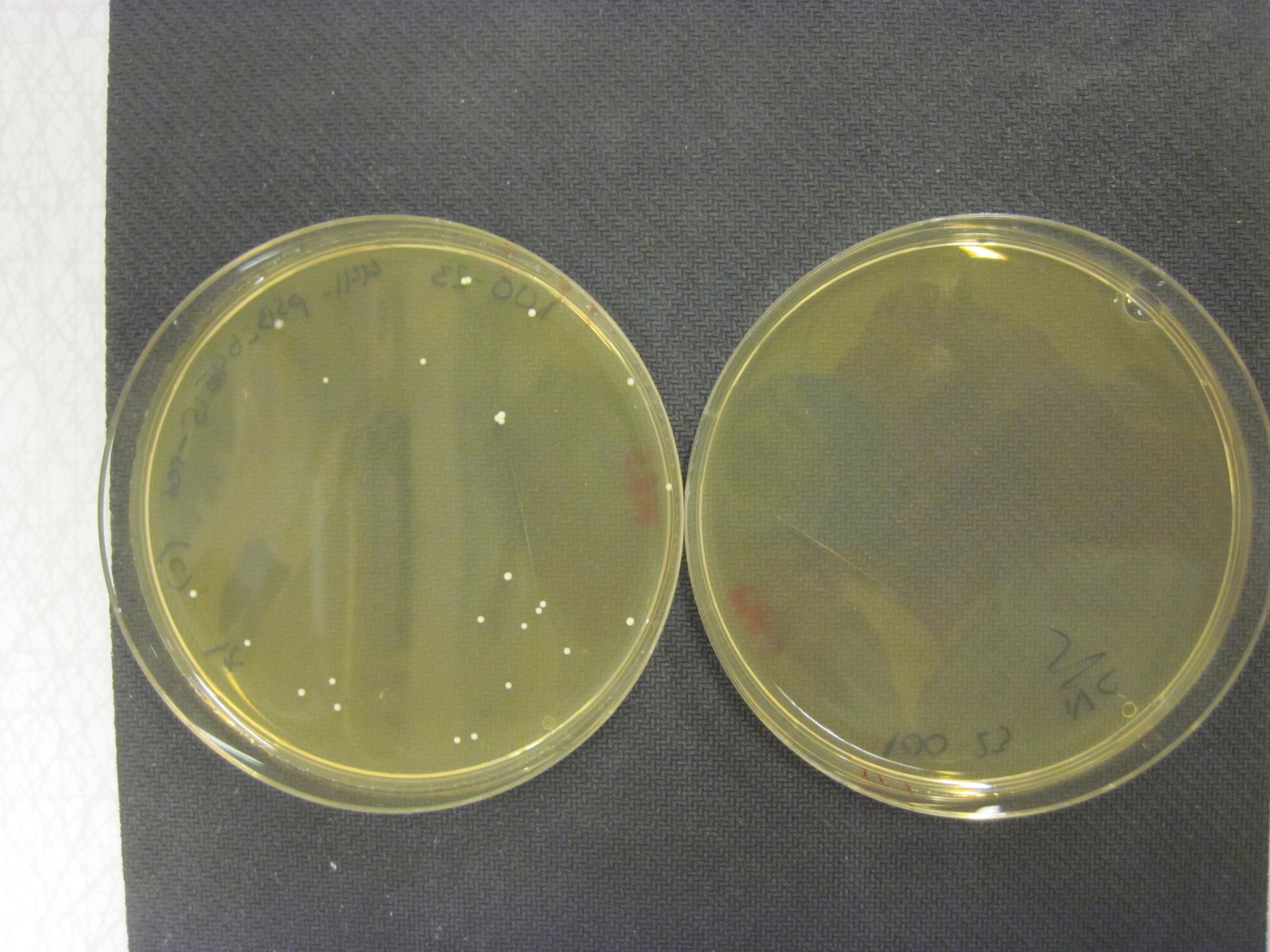
Competition test
We grew our E. coli Nissle containing the TAL biobrick together with wild type Nissle (mixed 1:1) in antiobiotic free LB medium. The culture was maintained by a thousandfold dilution in new medium every day, and letting it grow again overnight. To be able to track the fraction of each population over time, a dilution of the culture was plated on both selective, and non selective plates. The amount of colonies on the non selective plates gives us the total amount of cells, while the amount of colonies on the selective plates gives us the number of cells carrying the biobrick plasmid. The fraction of cells carrying the plasmid declined rapidly, and already after 5 days (approximately 50 generations of growth), we saw no trace of our E. coli Nissle with TAL on the selective plates, while the wild type still grew on the non selective plates. This clearly shows that our genetically engineered probiotic have a hard time competing against the wild type, and if released into the environment, they would most likely be outcompeted by other bacteria within days. To read more about the competition test, click here
Competition experiments between TAL and wild type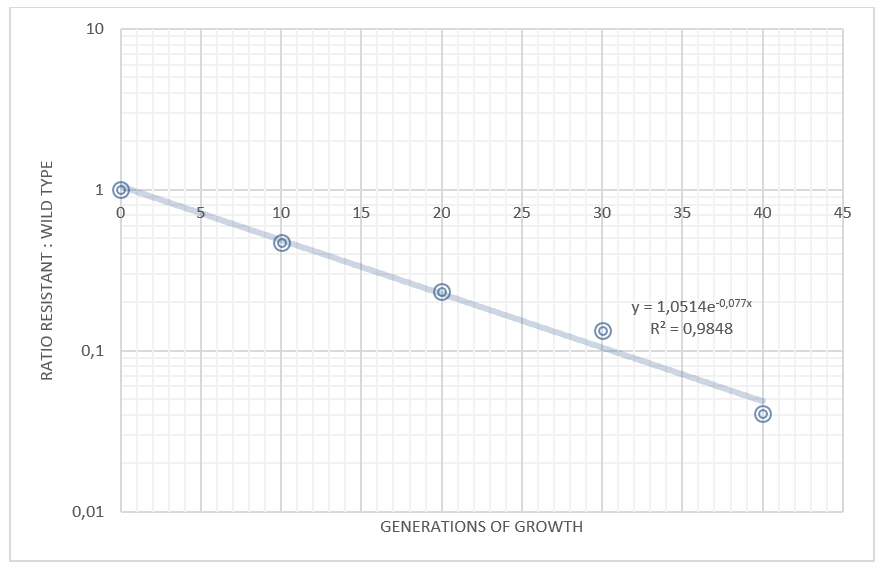
 "
"











