Team:Paris Saclay/Notebook/August/28
From 2013.igem.org
(→lab work) |
(→2 - Electrophoresis to check the gel purification of pSB1C3 digested by DnpI) |
||
| (25 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{Team:Paris_Saclay/incl_debut_generique}} | {{Team:Paris_Saclay/incl_debut_generique}} | ||
| + | |||
='''Notebook : August 28'''= | ='''Notebook : August 28'''= | ||
| - | ==''' | + | =='''Lab work'''== |
| - | + | ==='''A - Aerobic/Anaerobic regulation system'''=== | |
| - | + | ||
| - | + | ||
| - | + | ===='''Objective : characterize BBa_K1155000 and BBa_K1155004'''==== | |
| - | + | ||
| - | ==''' | + | ===='''1 - Purification of colony transformed with ligation : NirB with RBS-LacZ-Term in pSB1C3, NirB with RBS-Amil CP-Term in pSB1C3, Pndh* with RBS-LacZ-Term in pSB1C3, Pndh* with RBS-Amil CP-Term in pSB1C3 by streaking in aerobic or anaerobic conditions'''==== |
| - | + | XiaoJing | |
| - | + | ||
| - | + | ||
| - | |||
| - | |||
{| | {| | ||
| - | | style=" | + | | style="border:1px solid black;padding:5px;background-color:#DE;" | |
| - | + | Transformation of 08/26/13 works. We will use it to characterize all ligations. PCR colony of 08/27/13 works for . We also will use them to characterize ligations : NirB with RBS-Amil CP-Term in pSB1C3, Pndh* with RBS-LacZ-Term in pSB1C3, Pndh* with RBS-Amil CP-Term in pSB1C3. | |
| - | * | + | |} |
| - | + | ||
| + | We streak colonies from construction : | ||
| + | * Pndh* with RBS-LacZ-Term in pSB1C3 with O2 and Xgal | ||
| + | * Pndh* with RBS-Amil CP-Term in pSB1C3 with O2 | ||
| + | * Pndh* with RBS-Amil CP-Term in pSB1C3 without O2 | ||
| + | * NirB with RBS-LacZ-Term in pSB1C3 without O2 and Xgal | ||
| + | * NirB with RBS-Amil CP-Term in pSB1C3 with O2 | ||
| + | * NirB with RBS-Amil CP-Term in pSB1C3 without O2 | ||
| - | + | ===='''2 - Culture of mutant strain MG1655Z1 Δfnr''' ==== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | XiaoJing | |
| - | | | + | |
| - | | | + | {| |
| + | | style="border:1px solid black;padding:5px;background-color:#DE;" | | ||
| + | Transformation of 08/27/13 works. We will do a purification on LB plates. | ||
|} | |} | ||
| - | + | We streaked our colonies on plates with LB and incubated them at 42°C. | |
| + | ==='''A - Aerobic/Anaerobic regulation system / B - PCB sensor system'''=== | ||
| + | ===='''Objective : obtaining FRN and BphR2 proteins'''==== | ||
| - | + | ===='''1 - Gel purification of pSB1C3 digested by DnpI '''==== | |
| + | XiaoJing | ||
| - | + | Protocol : [http://www.mn-net.com/tabid/1452/default.aspx Gel purification ] | |
| - | + | ||
| - | + | ||
| + | Nanodrop : | ||
| + | * pSB1C3 : 37.2ng/µL | ||
| + | ===='''2 - Electrophoresis to check the gel purification of pSB1C3 digested by DnpI'''==== | ||
| - | + | XiaoJing | |
| - | + | ||
| - | | style="width: | + | {| |
| - | * | + | | style="width:350px;border:1px solid black;" |[[File:Psgel12808.jpg]] |
| - | * | + | | style="width:350px;border:1px solid black;vertical-align:top;" | |
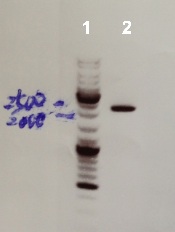
| - | * | + | * Well 1 : 6µL DNA Ladder |
| + | * Well 2 : 3µL of pSB1C3 digested by DpnI + 1µl of 6X loading dye | ||
| + | * Gel : 1% | ||
|} | |} | ||
| - | |||
| - | |||
| + | expected sizes : | ||
| + | * pSB1C3 : 2070 bp | ||
| - | + | {| | |
| - | + | | style="border:1px solid black;padding:5px;background-color:#DEDEDE;" | | |
| + | We obtained a fragment at the right size. The gel purification was good. We will use it for Gibson assembly. | ||
| + | |} | ||
| - | + | ===='''3 - Gibson assembly'''==== | |
| - | + | ||
| - | + | XiaoJing | |
| - | + | Used quantities : | |
| + | * RBS-BphR2 : | ||
| + | ** pSB1C3 : 3µL | ||
| + | ** BphR2 Part I : 1µL | ||
| + | ** BphR2 Part II : 1µL | ||
| + | ** Gibson mix : 15µL | ||
| - | * | + | * FNR : |
| + | ** pSB1C3 : 3µL | ||
| + | ** FNR Part I : 1µL | ||
| + | ** FNR Part II : 1µL | ||
| + | ** Gisbon mix : 15µL | ||
| + | * RBS-FNR : | ||
| + | ** pSB1C3 : 3µL | ||
| + | ** RBS-FNR Part I : 1µL | ||
| + | ** FNR Part II : 1µL | ||
| + | ** Gibson mix : 15µL | ||
| - | + | We incubate these mix at 50°C during 1h inside PCR machine. | |
| + | ===='''2 - Transformation of FNR, RBS-FNR and RBS-BphR2 in DH5α strain'''==== | ||
| - | + | XiaoJing | |
| + | Protocol : [[Team:Paris_Saclay/Protocols/Transformation|Bacterial transformation]] | ||
| - | |||
| - | |||
{| border="1" align="center" | {| border="1" align="center" | ||
| - | ||<big>Previous | + | |[[Team:Paris Saclay/Notebook/August/27|<big>Previous day</big>]] |
|[[Team:Paris_Saclay/Notebook|<big>Back to calendar</big>]] | |[[Team:Paris_Saclay/Notebook|<big>Back to calendar</big>]] | ||
| Line 95: | Line 110: | ||
|[[Team:Paris Saclay/Notebook/August/29|<big>Next day</big>]] | |[[Team:Paris Saclay/Notebook/August/29|<big>Next day</big>]] | ||
|} | |} | ||
| - | |||
{{Team:Paris_Saclay/incl_fin}} | {{Team:Paris_Saclay/incl_fin}} | ||
Latest revision as of 01:20, 5 October 2013
Notebook : August 28
Lab work
A - Aerobic/Anaerobic regulation system
Objective : characterize BBa_K1155000 and BBa_K1155004
1 - Purification of colony transformed with ligation : NirB with RBS-LacZ-Term in pSB1C3, NirB with RBS-Amil CP-Term in pSB1C3, Pndh* with RBS-LacZ-Term in pSB1C3, Pndh* with RBS-Amil CP-Term in pSB1C3 by streaking in aerobic or anaerobic conditions
XiaoJing
|
Transformation of 08/26/13 works. We will use it to characterize all ligations. PCR colony of 08/27/13 works for . We also will use them to characterize ligations : NirB with RBS-Amil CP-Term in pSB1C3, Pndh* with RBS-LacZ-Term in pSB1C3, Pndh* with RBS-Amil CP-Term in pSB1C3. |
We streak colonies from construction :
- Pndh* with RBS-LacZ-Term in pSB1C3 with O2 and Xgal
- Pndh* with RBS-Amil CP-Term in pSB1C3 with O2
- Pndh* with RBS-Amil CP-Term in pSB1C3 without O2
- NirB with RBS-LacZ-Term in pSB1C3 without O2 and Xgal
- NirB with RBS-Amil CP-Term in pSB1C3 with O2
- NirB with RBS-Amil CP-Term in pSB1C3 without O2
2 - Culture of mutant strain MG1655Z1 Δfnr
XiaoJing
|
Transformation of 08/27/13 works. We will do a purification on LB plates. |
We streaked our colonies on plates with LB and incubated them at 42°C.
A - Aerobic/Anaerobic regulation system / B - PCB sensor system
Objective : obtaining FRN and BphR2 proteins
1 - Gel purification of pSB1C3 digested by DnpI
XiaoJing
Protocol : [http://www.mn-net.com/tabid/1452/default.aspx Gel purification ]
Nanodrop :
- pSB1C3 : 37.2ng/µL
2 - Electrophoresis to check the gel purification of pSB1C3 digested by DnpI
XiaoJing
|
|
expected sizes :
- pSB1C3 : 2070 bp
|
We obtained a fragment at the right size. The gel purification was good. We will use it for Gibson assembly. |
3 - Gibson assembly
XiaoJing
Used quantities :
- RBS-BphR2 :
- pSB1C3 : 3µL
- BphR2 Part I : 1µL
- BphR2 Part II : 1µL
- Gibson mix : 15µL
- FNR :
- pSB1C3 : 3µL
- FNR Part I : 1µL
- FNR Part II : 1µL
- Gisbon mix : 15µL
- RBS-FNR :
- pSB1C3 : 3µL
- RBS-FNR Part I : 1µL
- FNR Part II : 1µL
- Gibson mix : 15µL
We incubate these mix at 50°C during 1h inside PCR machine.
2 - Transformation of FNR, RBS-FNR and RBS-BphR2 in DH5α strain
XiaoJing
Protocol : Bacterial transformation
| Previous day | Back to calendar | Next day |
 "
"