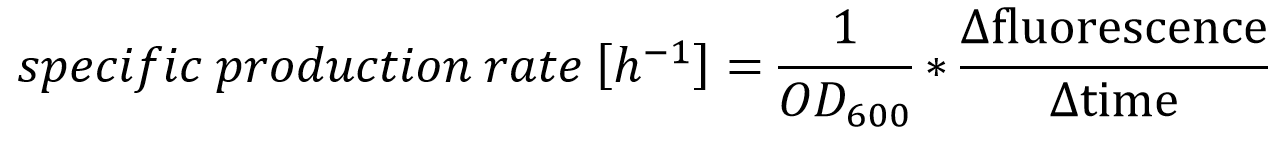Team:Bielefeld-Germany/Biosafety/Biosafety System S
From 2013.igem.org
| (36 intermediate revisions not shown) | |||
| Line 66: | Line 66: | ||
==Overview== | ==Overview== | ||
| - | [[File:IGEM Bielefeld 2013 Biosafety e.coli energypower.png|left|thumb|250px| | + | [[File:IGEM Bielefeld 2013 Biosafety e.coli energypower.png|left|thumb|250px|'''Biosafety-System AraCtive'''.]] |
<p align="justify"> | <p align="justify"> | ||
| - | The Biosafety-System araCtive <bbpart>BBa_K1172909</bbpart> is an improvement of the | + | The Biosafety-System araCtive <bbpart>BBa_K1172909</bbpart> is an improvement of the BioBrick <bbpart>BBa_K914014</bbpart> by replacing the first promoter by the rhamnose promoter P<sub>''Rha''</sub>, integration of the alanine racemase <bbpart>BBa_K1172901</bbpart> and utilization of the regulator AraC to control the transcription of the Barnase behind the P<sub>''BAD''</sub> promoter. Because of the tight repression of this promoter, this system has the lowest basal transcription of the Barnase and is therefore the most active and attractive one. |
</p> | </p> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<br><br> | <br><br> | ||
| Line 87: | Line 82: | ||
[[File:IGEM Bielefeld 2013 biosafety Rhamnose-promoter.png|left]] | [[File:IGEM Bielefeld 2013 biosafety Rhamnose-promoter.png|left]] | ||
<p align="justify"> | <p align="justify"> | ||
| - | The promoter P<sub>Rha</sub> (<bbpart>BBa_K914003</bbpart>) naturally regulates the catabolism of the hexose L-rhamnose. The advantage | + | The promoter P<sub>''Rha''</sub> (<bbpart>BBa_K914003</bbpart>) naturally regulates the catabolism of the hexose L-rhamnose. The advantage to use this promoter is its solely positive regulation. The regulon consists of the gene ''rhaT'' for the L-rhamnose transporter and the two operons ''rhaSR'' and ''rhaBAD''.<br> |
| - | The operon | + | The operon ''rhaSR'' encodes the two transcriptional activators RhaS and RhaR, who are responsible for the positive activation of the L-rhamnose catabolism, while the operon ''rhaBAD'' encodes the genes for the direct catabolism of L-rhamnose.<br> |
| - | When L-rhamnose is present, it acts as an inducer by binding to the regulatory protein RhaR. RhaR regulates | + | When L-rhamnose is present, it acts as an inducer by binding to the regulatory protein RhaR. RhaR regulates its own expression and the expression of the regulatory gene ''rhaS'' by repressing or, in the presence of L-rhamnose, activating the operon ''rhaSR''. Normally, the expression level is modest, but it can be enhanced by a higher level of intracellular cAMP, which increases in the absence of glucose. So, in the presence of L-rhamnose and a high concentration of intracellular cAMP, the activator protein RhaR is expressed on higher level, resulting in an activation of the promoter P<sub>''rhaT''</sub> for an efficient L-rhamnose uptake and an activation of the operon ''rhaBAD''. The L-rhamnose is than broken down into dihydroxyacetone phosphate and lactate aldehyde by the enzymes encoded by ''rhaBAD'' ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Wickstrum ''et al.'', 2005]).<br> |
| - | A brief schematic summary of the regulation is shown in Figure | + | A brief schematic summary of the regulation is shown in Figure 1. |
</p><br> | </p><br> | ||
<br> | <br> | ||
| - | [[File:IGEM Bielefeld 2013 biosafety Rhamnosepromoter 3.png|600px|thumb|center|'''Figure | + | [[File:IGEM Bielefeld 2013 biosafety Rhamnosepromoter 3.png|600px|thumb|center|'''Figure 1:''' The catabolism of L-rhamnose in ''E. coli'' is turned off in general but inducible by L-rhamnose. The induction activates the transcription of the genes ''rhaS'' and ''rhaR'', which regulate the L-rhamnose catabolism by positive activation of the rhamnose uptake (''rhaT'') and its metabolization (''rhaBAD'').]] |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | Dihydroxyacetone phosphate can be metabolized in the glycolysis pathway, while lactate aldehyde is | + | Dihydroxyacetone phosphate can be metabolized in the glycolysis pathway, while lactate aldehyde is oxidized to lactate under aerobic conditions and reduced to L-1,2,-propandiol under anaerobic conditions.<br> |
| - | + | The degradation of L-rhamnose can be separated in three steps. In the first step the L-rhamnose is turned into L-rhamnulose by an isomerase (gene ''rhaA''). L-rhamnulose is in turn phosphorylated to L-rhamnulose-1-phosphate by a kinase (gene ''rhaB'') and the sugar phosphate is finally hydrolyzed by an aldolase (gene ''rhaD'') to dihydroxyacetone phosphate and lactate aldehyde ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Baldoma ''et al.'', 1988]).<br> | |
| - | + | For our Safety-System araCtive, the rhamnose promoter P<sub>Rha</sub> is used to control the expression of the repressor AraC and the essential alanine racemase, because this promoter has an even lower basal transcription then the arabinose promoter P<sub>''BAD''</sub>. This is needed to tightly repress the expression of the alanine racemase (''alr'') and thereby take advantage of the double-kill switch. Although the rhamnose promoter P<sub>''Rha''</sub> is characterized by a very low basal transcription it can not be used for the control of the toxic RNase Ba ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#RNase_Ba_.28Barnase.29 Barnase]), which ist needed for the alanine racemase (''alr''), the other part of the double-kill switch. In conclusion this promoter is the best choice for the first part of the Biosafety-System, as it is tightly repressed in the absence of L-rhamnose, but activated in its presence.</p> | |
<br> | <br> | ||
| - | ===''' | + | ==='''Regulator AraC'''=== |
<br> | <br> | ||
[[File:IGEM Bielefeld 2013 biosafety araC.png|left]] | [[File:IGEM Bielefeld 2013 biosafety araC.png|left]] | ||
<p align="justify"> | <p align="justify"> | ||
| - | The araC gene is naturally found in ''E. coli'' and is coding for the 32 kDa AraC protein, which regulates the expression of the genes required for the uptake and catabolism of the pentose L-arabinose. The genes for the catabolism of arabinose are under the control of the arabinose promoter P<sub>BAD</sub>, which is both positively and negatively regulated by | + | The ''araC'' gene is naturally found in ''E. coli'' and is coding for the 32 kDa AraC protein, which regulates the expression of the genes required for the uptake and catabolism of the pentose L-arabinose. The genes for the catabolism of arabinose are under the control of the arabinose promoter P<sub>''BAD''</sub>, which is both positively and negatively regulated by AraC. Naturally in the presence of L-arabinose the expression of those genes is activated, while it is repressed in its absence. In addition the AraC protein regulates its own transcription under the control of the so called P<sub>C</sub> promoter. Compared to the P<sub>''BAD''</sub> promoter the P<sub>''C''</sub> promoter and the associated ''araC'' gene are therefore transcribed in opposite direction ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Schleif, 2010]).</p> |
<br> | <br> | ||
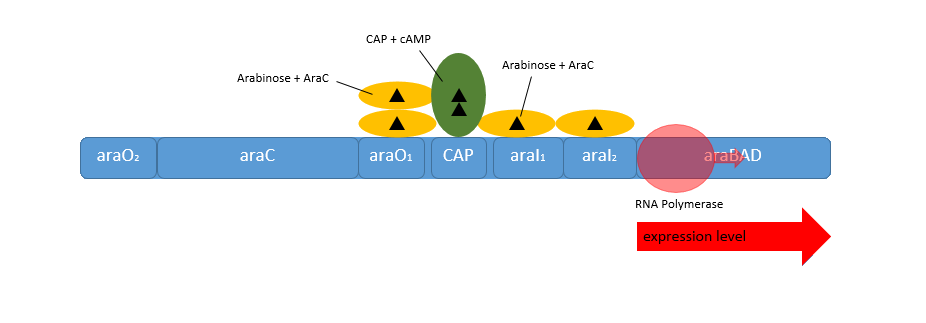
| - | [[File:IGEM Bielefeld 2013 Biosafety arabinosepromoter expression 2.png|600px|thumb|center|'''Figure | + | [[File:IGEM Bielefeld 2013 Biosafety arabinosepromoter expression 2.png|600px|thumb|center|'''Figure 2:''' Positive regulation of the arabinose promoter P<sub>''BAD''</sub>. When L-arabinose is present the dimer-structure of AraC is relaxed and the AraC protein functions as activator. The activation can additionally be enhanced by the CRP-cAMP-complex.]] |
<br> | <br> | ||
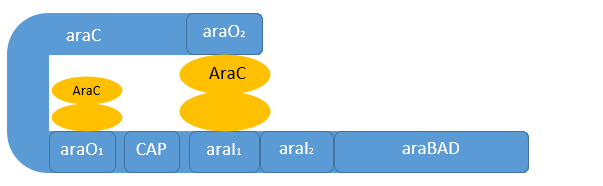
| - | [[File:IGEM Bielefeld 2013 Biosafety arabinosepromoter gehemmt.png|600px|thumb|center|'''Figure | + | [[File:IGEM Bielefeld 2013 Biosafety arabinosepromoter gehemmt.png|600px|thumb|center|'''Figure 3:''' Negative regulation of the arabinose promoter P<sub>''BAD''</sub>. In the absence of L-arabinose, AraC functions as repressor by forming a dimer, which inhibits the binding of the RNA-polymerase.]] |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | Naturally in the absence of L-arabinose the AraC protein is found in its repressor conformation, binding to the O<sub>2</sub>-region and I<sub>1</sub>-region. Binding two AraC proteins to | + | Naturally in the absence of L-arabinose the AraC protein is found in its repressor conformation, binding to the O<sub>2</sub>-region and I<sub>1</sub>-region. Binding of two AraC proteins to these domains results in a protein-protein-interaction of AraC forming a dimer. This AraC dimer is responsible for the tight repression of the P<sub>''C''</sub> and the P<sub>BAD</sub> promoter by forming a DNA-loop to inhibit the initiation of the RNA-polymerase (see Figure 3 above).<br> |
| - | When L-arabinose is present in the media the operon changes to the active form. Following this conditions the | + | When L-arabinose is present in the media the operon changes to the active form. Following this conditions the AraC protein functions as activator by binding to the I<sub>1</sub>- and I<sub>2</sub>-region of the operator region. This causes transcription of the genes behind the P<sub>''BAD''</sub> and the P<sub>''C''</sub> promoter. Depending on the intracellular level of cAMP the transcription of both the genes behind the promoters P<sub>''BAD''</sub> and P<sub>''C''</sub> can even be enhanced. In the absence of glucose the cell reaches a high level of cAMP as a signal molecule. cAMP binds as the typically CRP-cAMP complex binds in front of the AraC regulator and increases the binding affinity of the RNA-polymerase, so that the genes behind the P<sub>''BAD''</sub> promoter are expressed on higher levels. Besides the transcription affinity of the P<sub>''BAD''</sub> promoter, the AraC protein also regulates the genes ''araE'' and the operon consisting the genes ''araFGH'', which are necessary for an efficient uptake of L-arabinose ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Cass, 1988]).<br> |
| - | The auto-regulation of the ''araC'' gene itself works both in the presence and absence of L-arabinose. In presence of L-arabinose the DNA-loop which prevents also the transcription of the P<sub>BAD</sub> promoter also inhibits the transcription of ''araC''. When L-arabinose is present and the AraC concentration is high enough, | + | The auto-regulation of the ''araC'' gene itself works both in the presence and absence of L-arabinose. In presence of L-arabinose, the DNA-loop which prevents also the transcription of the P<sub>''BAD''</sub> promoter also inhibits the transcription of ''araC''. When L-arabinose is present and the AraC concentration is high enough, it auto-regulates its own transcription by dimerization between the ''araO''<sub>1</sub>- and the ''araO''<sub>2</sub>-region. This leads to an other DNA-loop, inhibiting solely the transcription of the genes behind the P<sub>''C''</sub> promoter ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Hamilton, 1988]).<br> |
| - | For our Biosafety-System we decided to use the arabinose P<sub>BAD</sub> promoter, because this promoter is | + | For our Biosafety-System we decided to use the arabinose P<sub>''BAD''</sub> promoter, because this promoter is tightly regulated by AraC and shows no basal transcription in the complete absence of AraC. Additionally this promoter is repressed by glucose and basal transcription is activated in the presence of cAMP.</p> |
<br> | <br> | ||
| Line 125: | Line 120: | ||
[[Image:IGEM Bielefeld 2013 biosafety alr test.png|left]] | [[Image:IGEM Bielefeld 2013 biosafety alr test.png|left]] | ||
<p align="justify"> | <p align="justify"> | ||
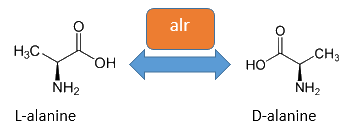
| - | The alanine racemase Alr (EC 5.1.1.1) from the | + | The alanine racemase Alr (EC 5.1.1.1) from the Gram-negative enteric bacteria ''Escherichia coli'' is a racemase, which catalyses the reversible conversion of L-alanine into the enantiomer D-alanine (see Figure 4). For this reaction, the cofactor pyridoxal-5'-phosphate (PLP) is necessary. The constitutively expressed alanine racemase (''alr'') is naturally responsible for the accumulation of D-alanine. This compound is an essential component of the bacterial cell wall, because it is used for the cross-linkage of peptidoglycan ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Walsh, 1989]).<br> |
| - | The usage of D-alanine instead of a typically L-amino | + | The usage of D-alanine instead of a typically L-amino acid prevents cleavage by peptidases. However, a lack of D-alanine causes to a bacteriolytic characteristics. In the absence of D‑alanine dividing cells will lyse rapidly. This fact is used for our Biosafety-Strain, a D-alanine auxotrophic mutant (K-12 ∆''alr'' ∆''dadX''). The Biosafety-Strain grows only with a plasmid containing the alanine racemase (<bbpart>BBa_K1172901</bbpart>) to complement the D-alanine auxotrophy. Consequently the alanine racemase is essential for bacterial cell division. This approach guarantees a high plasmid stability, which is extremely important when the plasmid contains a toxic gene like the [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#RNase_Ba_.28Barnase.29 Barnase]. In addition this construction provides the possibility for the implementation of a double kill-switch system. Because if the expression of the alanine racemase is repressed and there is no D-alanine supplementation in the medium, cells will not grow.</p> |
<br> | <br> | ||
| - | [[Image:IGEM Bielefeld 2013 alr isomerase bearbeitet.png|600px|thumb|center|'''Figure | + | [[Image:IGEM Bielefeld 2013 alr isomerase bearbeitet.png|600px|thumb|center|'''Figure 4:''' The alanine racemase (<bbpart>BBa_K1172901</bbpart>) from ''E. coli'' catalyses the reversible conversion from L-alanine to D-alanine. For this isomerization the cofactor pyridoxal-5'-phosphate is necessary.]] |
<br> | <br> | ||
| Line 136: | Line 131: | ||
[[File:IGEM Bielefeld 2013 biosafety Terminator.png|left]] | [[File:IGEM Bielefeld 2013 biosafety Terminator.png|left]] | ||
<p align="justify"> | <p align="justify"> | ||
| - | Terminators are essential | + | Terminators are essential to terminate transcription of an operon. In procaryotes, two types of terminators exist. The rho-dependent and the rho-independent terminator. Rho-independent terminators are characterized by their stem-loop forming sequence. In general, the terminator-region can be divided into four regions. The first region is GC-rich and constitutes one half of the stem. This region is followed by the loop-region and another GC-rich region that makes up the opposite part of the stem. The terminator closes with a poly uracil region, which destabilizes the binding of the RNA-polymerase. The stem-loop of the terminator causes a distinction of the DNA and the translated RNA. Consequently the binding of the RNA-polymerase is cancelled and the transcription ends after the stem-loop ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Carafa ''et al.'', 1990]).<br> |
| - | For our Bioafety-System araCtive the terminator is necessary to avoid that the expression of the genes under control of the | + | For our Bioafety-System araCtive the terminator is necessary to avoid that the expression of the genes under control of the rhamnose promoter P<sub>''Rha''</sub>, like the Repressor AraC and the alanine racemase (''alr'') results in the transcription of the genes behind the arabinose promoter P<sub>''BAD''</sub>, which contains the toxic Barnase <bbpart>BBa_K1172904</bbpart> and would lead to cell death.</p> |
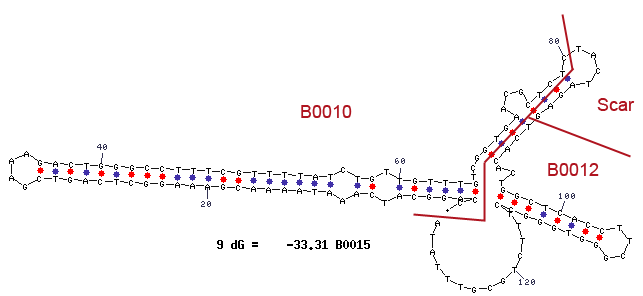
<br> | <br> | ||
| - | [[File:Team Bielefeld Biosafety Terminator.png|400x600px|thumb|center| '''Figure | + | [[File:Team Bielefeld Biosafety Terminator.png|400x600px|thumb|center| '''Figure 5:''' Stem-loop structure of the terminator <bbpart>BBa_B0015</bbpart>, which is used for the Biosafety-System araCtive. The terminator is used to make sure that solely both the repressor AraC and the alanine racemase Alr are expressed but the transcription of the toxic RNase Ba (Barnase) is avoided.]] |
<br> | <br> | ||
| Line 147: | Line 142: | ||
[[File:IGEM Bielefeld 2013 biosafety pBAD.png|left]] | [[File:IGEM Bielefeld 2013 biosafety pBAD.png|left]] | ||
<p align="justify"> | <p align="justify"> | ||
| - | The arabinose promoter P<sub>BAD</sub> controls the expression of the genes, which are necessary for the catabolism of the pentose arabinose. The expression is regulated by the | + | The arabinose promoter P<sub>''BAD''</sub> controls the expression of the genes, which are necessary for the catabolism of the pentose arabinose. The expression is regulated by the bifunctional regulator AraC. The genes of this operon convert the pentose to xylose, which can be converted into fructose-1,6-diphosphate and subsequently be fed into glycolysis. The conversion of L-arabinose is realized in three steps. First the isomerase (AraA) catalyzes the reaction from L-arabinose to the isomer L-ribulose. In the second step the ribulose kinase (AraB) phosphorylates the L-ribulose to L-ribulose-phosphate. For this reaction ATP is needed. Last, the L-ribulose-phosphate is converted to D-xylose-5-phosphat by L-ribulose-5-phosphate-epimerase (AraD, [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Schleif, 2010]).</p> |
<br> | <br> | ||
| - | [[File:IGEM Bielefeld 2013 Biosafety arabinosepromoter basale transkription 3.png|600px|thumb|center|'''Figure | + | [[File:IGEM Bielefeld 2013 Biosafety arabinosepromoter basale transkription 3.png|600px|thumb|center|'''Figure 6:''' Overview of the arabinose regulon from ''E. coli''. The transcription of the genes ''araBAD'', which are coding for the enzymes of the L-arabinose catabolism are positively induced and negatively repressed by the AraC protein (depending on the presence of arabinose), which additionally autoregulates his own transcription.]] |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| Line 160: | Line 155: | ||
[[Image:IGEM Bielefeld 2013 biosafety RNase Ba test.png|left]] | [[Image:IGEM Bielefeld 2013 biosafety RNase Ba test.png|left]] | ||
<p align="justify"> | <p align="justify"> | ||
| - | The Barnase (EC 3.1.27) is a 12 kDa extracellular microbial ribonuclease, which is naturally found in the | + | The Barnase (EC 3.1.27) is a 12 kDa extracellular microbial ribonuclease, which is naturally found in the Gram-positive soil bacteria ''Bacillus amyloliquefaciens'' and consists of a single chain of 110 amino acids. The Barnase (RNase Ba) catalyses the cleavage of single stranded RNA, preferentially behind Gs. In the first step of the RNA-degradation a cyclic intermediate is formed by transesterification and afterwards this intermediate is hydrolyzed yielding in a 3'-nucleotide ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Mossakowska ''et al.'', 1989]).</p> |
<br> | <br> | ||
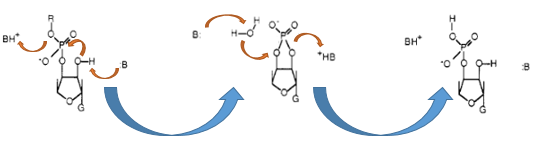
| - | [[Image:IGEM Bielefeld 2013 Biosafety Barnasemove.png|600px|thumb|center|'''Figure | + | [[Image:IGEM Bielefeld 2013 Biosafety Barnasemove.png|600px|thumb|center|'''Figure 7:''' Enzymatic reaction of the RNA-cleavage by the RNase Ba. First the transesterification by the Glu-73 residue is performed and then this cyclic intermediate is hydrolyzed by the His-102 of the Barnase.]] |
<br> | <br> | ||
| - | |||
<p align="justify"> | <p align="justify"> | ||
| - | In ''Bacillus amyloliquefaciens'' the activity | + | In ''Bacillus amyloliquefaciens'', the activity of the Barnase (RNase Ba) is inhibited intracellular by an inhibitor called barstar. Barstar consists of only 89 amino acids and binds with a high affinity to the toxic Barnase. This prevents the cleavage of the intracellular RNA in the host organism ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Paddon ''et al.'', 1989]). Therefore the Barnase normally acts only outside the cell and is translocated under natural conditions. For the Biosafety-System araCtive we modified the enzyme by cloning only the sequence responsible for the cleavage of the RNA, leaving out the N-terminal signal peptide part. |
<br> | <br> | ||
| - | As shown in | + | As shown in Figure 8 below, the transcription of the DNA, which encodes the Barnase produces a 474 nt RNA. The translation of the RNA starts about 25 nucleotides downstream from the transcription start and can be divided into two parts. The first part (colored in orange) is translated into a signal peptide at the amino-terminus of the Barnase coding RNA. This part is responsible for the extracellular translocation of the RNase Ba, while the peptide sequence for the active Barnase starts 142 nucleotides downstream from the transcription start (colored in red).<br> |
| - | For the Biosafety-System araCtive we only used the | + | For the Biosafety-System araCtive, we only used the part (<bbpart>BBa_K1172904</bbpart>) of the Barnase encoding the catalytic domain without the extracellular translocation signal of the toxic gene product. Translation of the barnase gene leads to rapid cell death if the expression of the Barnase is not repressed by the repressor AraC of our Biosafety-System.</p> |
<br> | <br> | ||
| - | [[Image:Team-Bielefeld-Biosafety_Barnase_Sequence.png|600px|thumb|center|'''Figure | + | [[Image:Team-Bielefeld-Biosafety_Barnase_Sequence.png|600px|thumb|center|'''Figure 8:''' Sequence of the signal peptide amino terminal of the RNase Ba (Barnase). The Biobrick <bbpart>BBa_K1172904</bbpart> does not contain the signal sequence for the extracellular translocation, but only the coding sequence for the mature enzyme.]] |
| Line 182: | Line 176: | ||
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
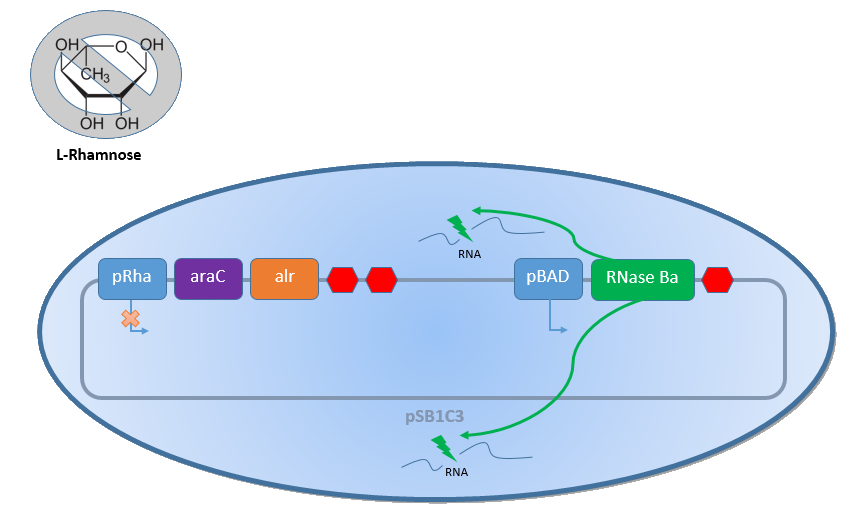
| - | Combining the gene described above with the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' results in a powerful device, allowing us to control the bacterial cell division. The control of the bacterial growth is possible either active or passive. Active by inducing the P<sub>BAD</sub> promoter with L-arabinose and passive by the induction of L-rhamnose. The passive control makes it possible to control the bacterial cell division in a defined closed environment, like the MFC, by | + | Combining the gene described above with the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' results in a powerful device, allowing us to control the bacterial cell division. The control of the bacterial growth is possible either active or passive. Active by inducing the P<sub>''BAD''</sub> promoter with L-arabinose and passive by the induction of L-rhamnose. The passive control makes it possible to control the bacterial cell division in a defined closed environment, like the MFC, by continuously adding L-rhamnose to the medium. As shown in the Figure 9 below, this leads to an expression of the essential alanine racemase (''alr'') and the AraC regulator, so that the expression of the RNase Ba is repressed. </p><br> |
| - | [[File:IGEM Bielefeld 2013 Biosafety System S+ 2.png|600px|thumb|center|'''Figure | + | [[File:IGEM Bielefeld 2013 Biosafety System S+ 2.png|600px|thumb|center|'''Figure 9:''' Biosafety-System araCtive in the presence of L-rhamnose. The essential alanine racemase (Alr) and the repressor AraC are expressed, resulting in a repression of the expression of the RNAse Ba. Consequently the bacteria show normal growth behaviour.]] |
<br> | <br> | ||
| - | |||
<p align="justify"> | <p align="justify"> | ||
| - | In the event that bacteria exit the defined environment of the MFC or L-rhamnose is not added to the medium any more, both the expression of the alanine | + | In the event that bacteria exit the defined environment of the MFC or L-rhamnose is not added to the medium any more, both the expression of the alanine racemase (Alr) and the AraC regulatordecrease, so that the expression of the toxic RNase Ba (Barnase) begins. The cleavage of the intracellular RNA by the Barnase and the lack of synthesized D-alanine, caused by the repressed alanine racemase inhibit the cell division and makes sure that the bacteria can only grow in the defined environment or the device of choice respectively. </p><br> |
| - | [[File:IGEM Bielefeld 2013 Biosafety System S ohne Rhamnose 2.png|600px|thumb|center|'''Figure | + | [[File:IGEM Bielefeld 2013 Biosafety System S ohne Rhamnose 2.png|600px|thumb|center|'''Figure 10:''' Active Biosafety-System araCtive outside of a defined environment lacking L-rhamnose. Both the expression of the alanine racemase (Alr) and AraC repressor are reduced and ideally completely shutdown. In contrast, the expression of the RNase ba (Barnase) is turned on, leading to cell death by RNA cleavage.]] |
<br> | <br> | ||
| Line 198: | Line 191: | ||
==Results== | ==Results== | ||
<br> | <br> | ||
| - | ==='''Characterization of the arabinose promoter | + | ==='''Characterization of the arabinose promoter P<sub>''BAD''</sub>'''=== |
<p align="justify"> | <p align="justify"> | ||
| + | First of all, the arabinose promoter P<sub>''BAD''</sub> was characterized to get a first impression of its basal transcription rate. Therefore the bacterial growth was investigated under the pressure of the unrepressed P<sub>''BAD''</sub> promoter on different carbon source using M9 minimal medium with either glucose or glycerol. The transcription rate was determined by fluorescence measurement of GFP <bbpart>BBa_E0040</bbpart> transcribed via the P<sub>''BAD''</sub> promoter using the BioBrick <bbpart>BBa_I13541</bbpart>.<br> | ||
| - | + | As shown in Figure 12 below, the bacteria grew better on glucose than on glycerol. This is caused by the fact that glucose is the better energy source of this two, because glycerol enters glycolysis at a later step and therefore delivers less energy (in addition to the additional ATP consumption to drive glycerol uptake). For the investigation of the basal transcription the fluorescence measurements, shown in Figure 13, is more interesting. <br> | |
| - | + | Both the wild type ''E. coli'' K-12 and the strain ''E. coli'' K-12 ∆''alr'' ∆''dadX'' ∆''araC'' show about the same fluorescence on glucose (red and orange curve) but differ on glycerol. This can be explained by the fact that growth on glucose does not cause an additional induction of the arabinose promoter P<sub>''BAD''</sub>, while glycerol does. In the presence of glucose, the intracellular concentration of cAMP is low. The absence of cAMP results in inactive CAP (carbon utilization activator protein) and thus no induction of inefficient catabolisms like the catabolism of arabinose. Therefore, glucose is catabolized by the bacteria first, resulting in an optimal growth. In the absence of glucose, the cAMP level increases which enhances via CAP-cAMP the transcription of most operons encoding enzymes for the catabolism of alternative carbon sources. Therefore, the expression of GFP under the control of the P<sub>BAD</sub> promoter increases on glycerol. Another difference can be seen between the wild type (black curve) and the strain K-12 ∆''alr'' ∆''dadX'' ∆''araC'' (blue curve). | |
| - | + | The Biosafety-Strain shows lower expression then the wild type, but has about the same growth rate, according to Figure 11. The reason for this characteristic is caused by the deleted AraC protein in the Biosafety-Strain. Because the AraC protein functions not only as are repressor but also as an activator the transcription rate decreases in the Biosafety-Strain.</p><br> | |
| - | As shown in Figure 12 below, the bacteria | + | |
| - | + | ||
| - | The Biosafety-Strain shows lower expression then the wild type, but has about the same growth rate, according to Figure | + | |
<br> | <br> | ||
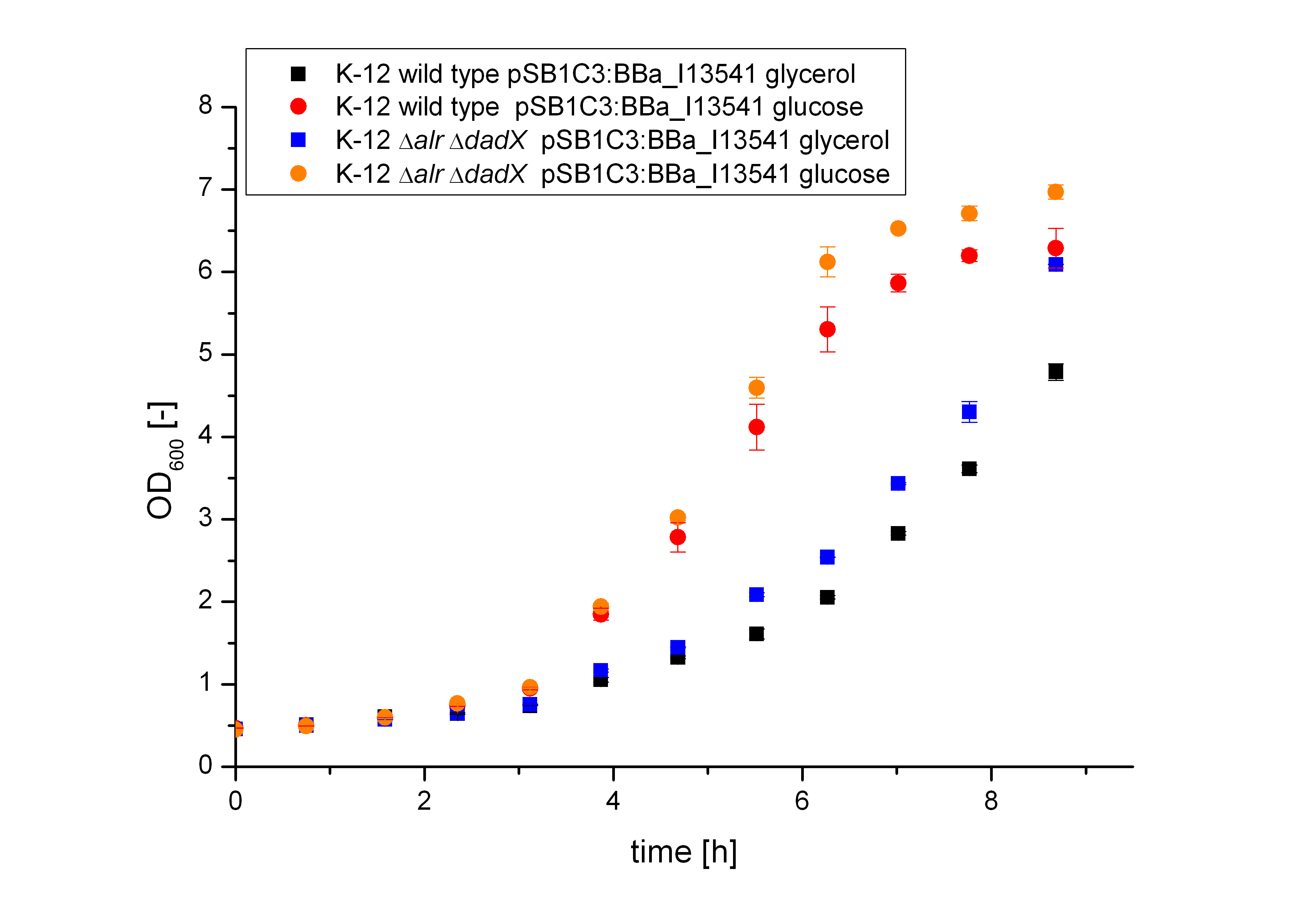
| - | [[File:Team-Bielefeld-Biosafety-System_araCtive_OD.jpg|600px|thumb|center|'''Figure | + | [[File:Team-Bielefeld-Biosafety-System_araCtive_OD.jpg|600px|thumb|center|'''Figure 11:''' Characterization of the bacterial growth of the strain K-12 ∆''alr'' ∆''dadX'' ∆''araC'' containing the plasmid <bbpart>BBa_I13541</bbpart> with GFP (<bbpart>BBa_E0040</bbpart>) under the control of the P<sub>''BAD''</sub> promoter. The M9 medium was supplemented with 5 mM D-alanine. It could be demonstrated, that the bacteria grow faster on M9 minimal medium containing glucose than on M9 minimal medium with glycerol.]] |
<br> | <br> | ||
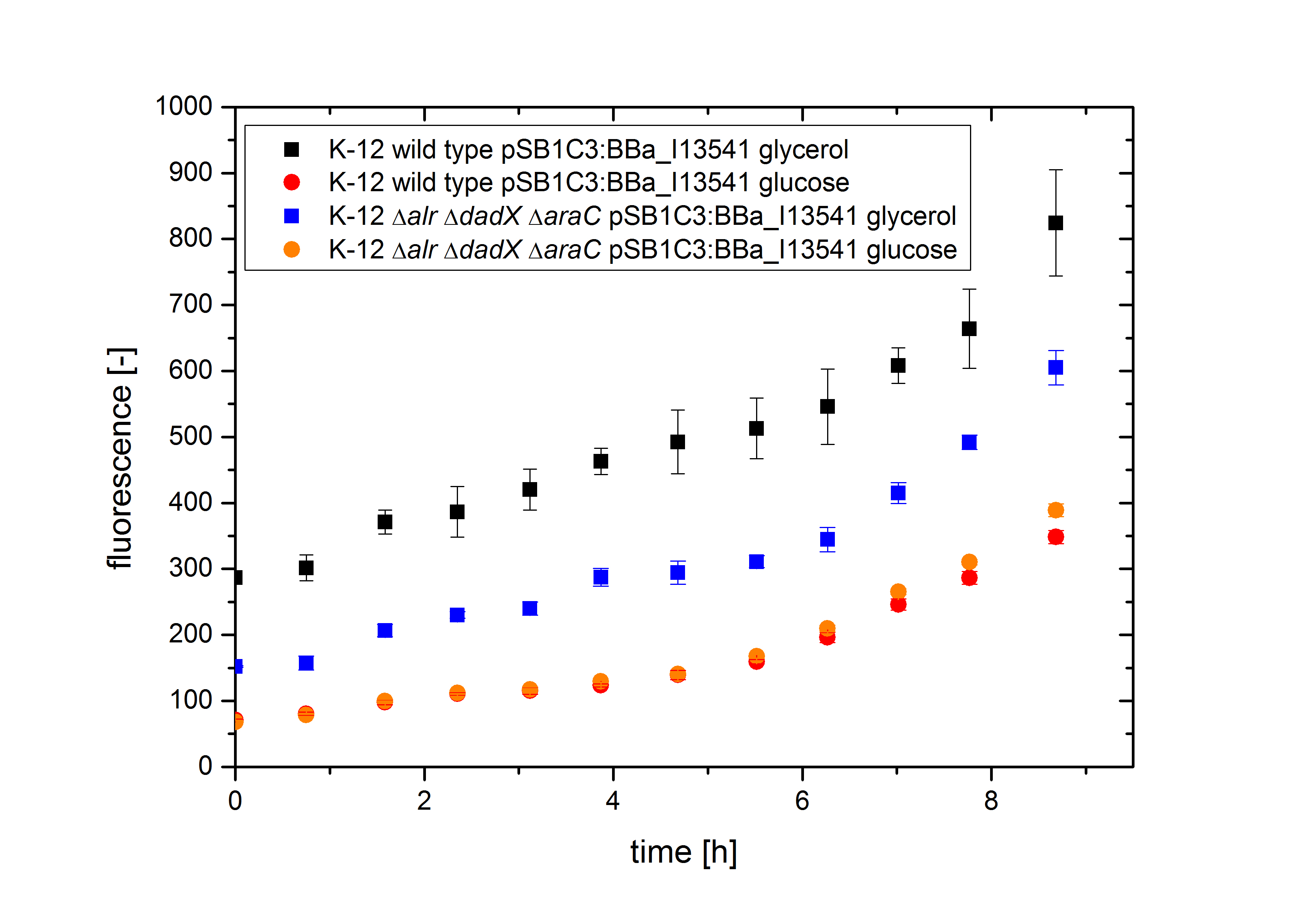
| - | [[File:Team-Bielefeld_Biosafety-System- | + | [[File:Team-Bielefeld_Biosafety-System-araCtive_fluorescence2.jpg|600px|thumb|center|'''Figure 12:''' Characterization of the fluorescence of the strain K-12 ∆''alr'' ∆''dadX'' ∆''araC'' containing the plasmid <bbpart>BBa_I13541</bbpart> with GFP (<bbpart>BBa_E0040</bbpart>) under control of the P<sub>''BAD''</sub> promoter. The Biosafety-Strain was cultivated on M9 minimal medium with 5 mM D-alanine supplemented.]] |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | The effect that glucose represses the transcription of the P<sub>BAD</sub> promoter becomes more | + | The effect that glucose represses the transcription of the P<sub>''BAD''</sub> promoter becomes more obvious by calculating of the specific production rates of GFP, as shown in Figure 13. The specific production rates were calculated via equation (1) :</p><br> |
[[File:IGEM Bielefeld 2013 Sepzifische Produktionsrate.png|600px|center|]] | [[File:IGEM Bielefeld 2013 Sepzifische Produktionsrate.png|600px|center|]] | ||
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
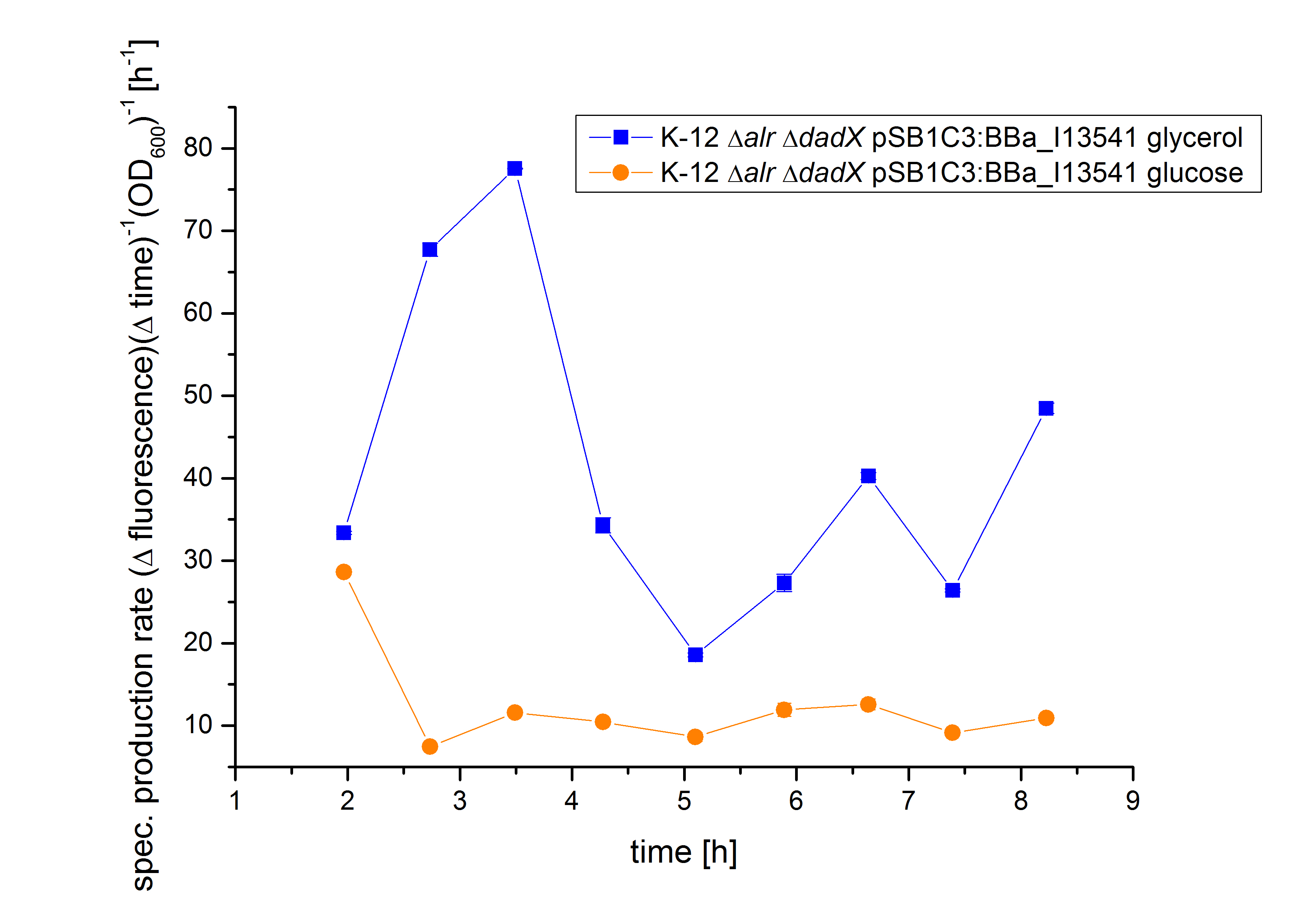
| - | With the specific production rate of GFP it can be demonstrated that | + | With the specific production rate of GFP, it can be demonstrated that GFP synthesis rates differ extremely between the cultivation on glucose and glycerol. The specific production rate is constantly very low when using glucose as carbon source, but shows a higher more unstable level in the cultivation with glycerol.<br> |
| - | Because the specific production rate was calculated between every single measurement point, the curve in Figure | + | Because the specific production rate was calculated between every single measurement point, the curve in Figure 13 is not smoothed and so the fluctuations have to be ignored, as they do not stand for are real fluctuations in the expression of GFP. They are caused by measurement variations in the growth curve and the fluorescence curve. And as neither of this curves are ideal, the fluctuations are the result. Nevertheless this graph shows clearly the difference between the two carbon sources.</p><br> |
| - | [[File:Team-Bielefeld-Biosafety-System-araCtive-pBAD- | + | [[File:Team-Bielefeld-Biosafety-System-araCtive-pBAD-GFProduct2.jpg|600px|thumb|center|'''Figure 13:''' Specific production rate of GFP transcribed via the P<sub>BAD</sub> promoter in dependence of different carbon sources.]] |
<br> | <br> | ||
| Line 230: | Line 221: | ||
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | The Biosafety-System araCtive was characterized on M9 minimal medium using glycerol as carbon source. As for the characterization of the pure arabinose promoter P<sub>BAD</sub> above, the bacterial growth and the fluorescence of GFP <bbpart>BBa_E0040</bbpart> was measured. Therefore the wild type and the Biosafety-Strain ''E. coli'' K-12 | + | The Biosafety-System araCtive was characterized on M9 minimal medium using glycerol as carbon source. As for the characterization of the pure arabinose promoter P<sub>''BAD''</sub> above, the bacterial growth and the fluorescence of GFP <bbpart>BBa_E0040</bbpart> was measured. Therefore, the wild type and the Biosafety-Strain ''E. coli'' K-12 ∆''alr'' ∆''dadX'', both containing the Biosafety-Plasmid <bbpart>BBa_K1172909</bbpart>, were cultivated once with the induction of 1% L-rhamnose and once only on glycerol.<br> |
| - | + | It becomes obvious (Figure 14) that the bacteria, induced with 1 % L-rhamnose (red and black curve) grow significantly slower than on pure glycerol (orange and blue curve). This is attributed to the high metabolic burden encountered by the induced bacteria. The expression of the repressor AraC and the alanine racemase (Alr) simultaneously causes a high strain on the cells, so that they grow slower than the uninduced cells, which express only GFP.<br> | |
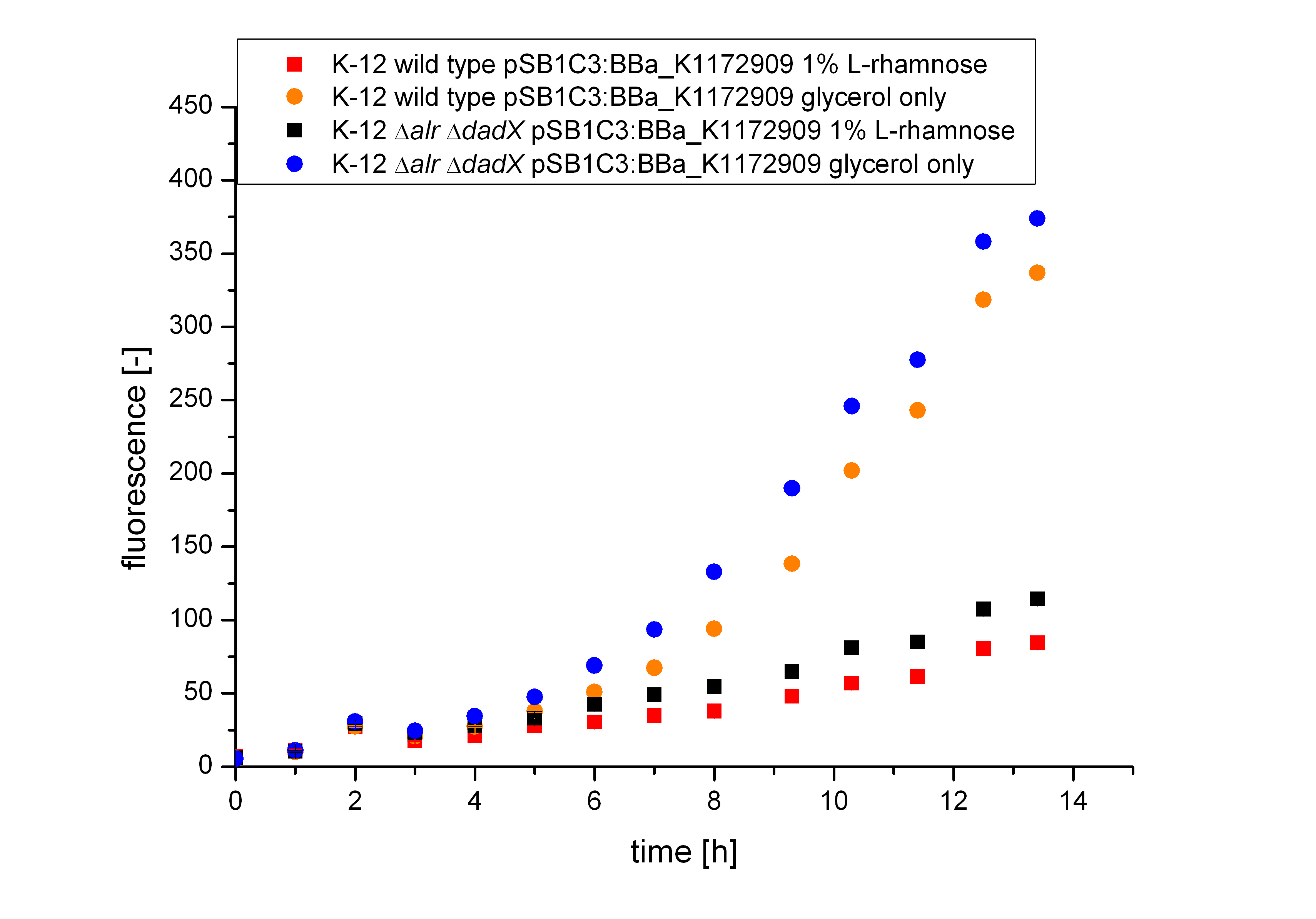
| - | Comparing the bacterial growth with the fluorescence in Figure | + | Comparing the bacterial growth with the fluorescence in Figure 15, it can be seen that the fluorescence seems to follow the same trend than the bacterial growth. The uninduced cells show approximately an exponential rise of fluorescence, while in comparision the fluorescence of the induced bacteria increases only slowly.</p><br> |
| - | [[File:Team-Bielefeld-Biosafety-System-araCtive-ODALL.jpg|600px|thumb|center|'''Figure | + | [[File:Team-Bielefeld-Biosafety-System-araCtive-ODALL.jpg|600px|thumb|center|'''Figure 14:''' Characterization of the bacterial growth of the Biosafety-System on M9 minimal medium with glycerol. The Figure compares the wild type K-12 and the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' containing the Biosaftey-Plasmid <bbpart>BBa_K1172909</bbpart> and the induction by 1% L-rhamnose to pure glycerol.]] |
<br> | <br> | ||
| - | [[File:Team-Bielefeld-Biosafety-System-araCtive-FlourescenceALL.jpg|600px|thumb|center|'''Figure | + | [[File:Team-Bielefeld-Biosafety-System-araCtive-FlourescenceALL.jpg|600px|thumb|center|'''Figure 15:''' Characterization of the fluorescence of the Biosafety-System araCtive. The Figure compares the wild type K-12 and the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' containing the Biosafety-Plasmid <bbpart>BBa_K1172909</bbpart> and the induction by 1% L-rhamnose to pure glycerol.]] |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | From the | + | From the data presented above it can not be determined if the expression of the repressor AraC does affect the transcription of GFP or not. The slower growth of the bacteria is a first indication that the repressor AraC and the alanine racemase (Alr) are highly expressed, but the growth of the bacteria shows nearly the same kinetics as the fluorescence. So it could be possible that the repressor does not affect the expression level of GFP under the control of the arabinose promoter P<sub>''BAD''</sub>. This becomes more clear by the calculation of the specific production rate of GFP by equation (1) . As shown in Figure 17 below the specific production rate differs clearly between the uninduced Biosafety-System and the Biosafety-System induced by 1% L-rhamnose. The production of GFP in the presence of L-rhamnose (red curve) is always lower than in its absence (orange curve), so that the expression of GFP is repressed in the presence of L-rhamnose.<br> |
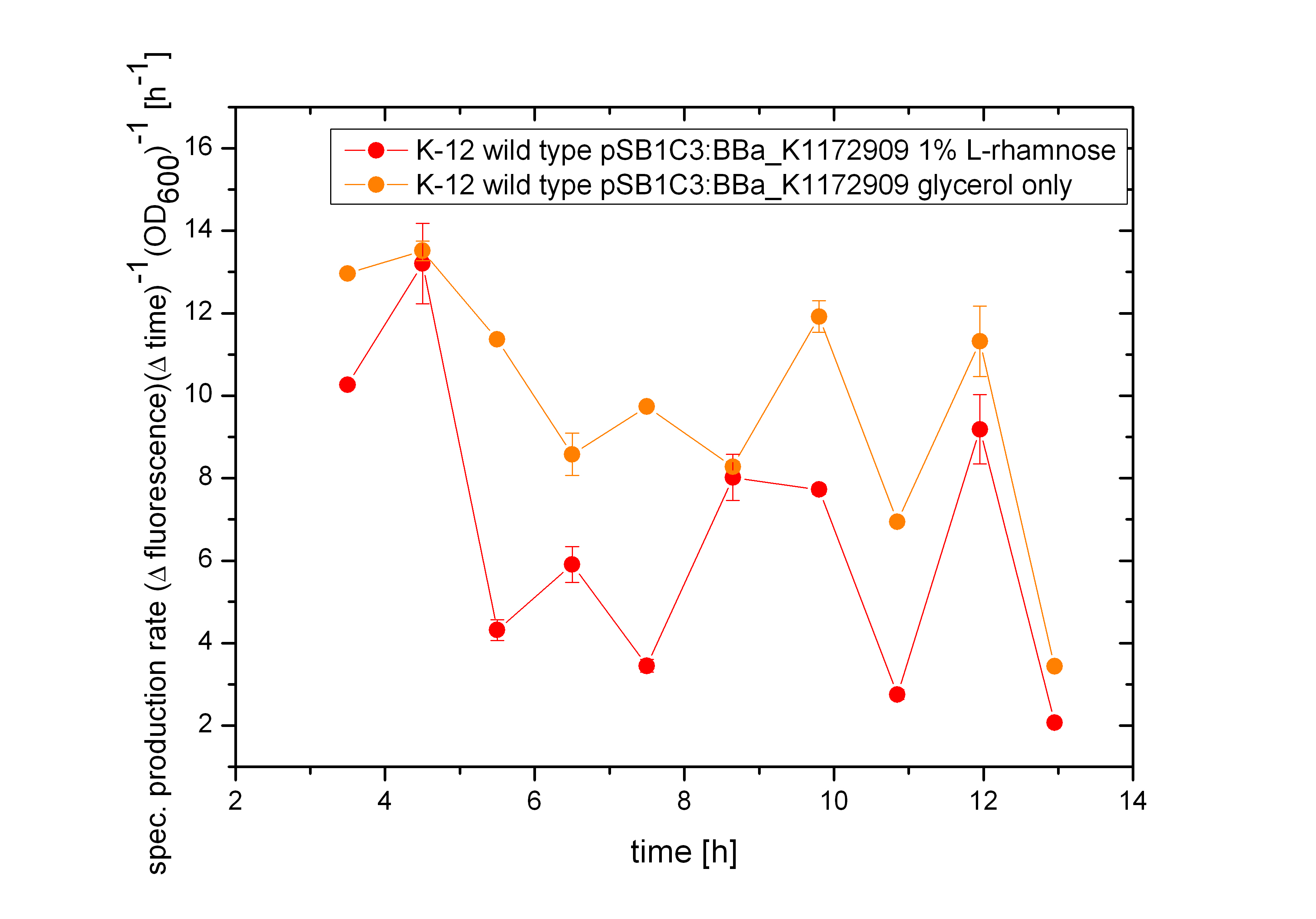
| - | Because the specific production rate of GFP was calculated between every single measurement point, the curve in Figure | + | Because the specific production rate of GFP was calculated between every single measurement point, the curve in Figure 16 is not smoothed and so the fluctuations have to be ignored, as they do not stand for are real fluctuations in the expression of GFP. They are caused by measurement variations in the growth curve and the fluorescence curve. But there is a clear tendency that the production of GFP is significantly lower when the bacteria are induced with 1% L-rhamnose. So the Biosafety-System araCtive works.</p><br> |
| - | [[File:Team-Bielefeld-Biosafety-System-araCtive-spezProductSara.jpg|600px|thumb|center|'''Figure | + | [[File:Team-Bielefeld-Biosafety-System-araCtive-spezProductSara.jpg|600px|thumb|center|'''Figure 16:''' Specific production rate of GFP for the Biosafety-System araCtive, calculated via equation (1). The production rate of GFP of the uninduced bacteria is significantly higher compared to the bacteria induced with 1% L-rhamnose. The Biosafety-System AraCtive works.]] |
| - | ===''' | + | ==='''Conclusions'''=== |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | + | It could be demonstrated that the Biosafety-System arActive works in principle. For example, the expression level of GFP is decreased in the presence of L-rhamnose, as can be seen in Figure 17, depicting the specific production rates of GFP after 7,5 hours. The induction of the Biosafety-System with L-rhamnose leads to a tight repression of the expression of GFP (red bar) when compared to the uninduced state (orange bar). Therefore the second part of the Biosafety-System, which contains normally the toxic RNase Ba <bbpart>BBa_K1172904</bbpart> can be used as a kill-switch regulated by the pentose L-rhamnose. In [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System#Results comparison] to the other Biosafety-Systems it becomes clear, that the Biosafety-System araCtive would be probably the best choice of our three Biosafety-Systems, because of its tight repression by the P<sub>''BAD''</sub> promoter of the second part.<br> | |
| + | But a number of problems remain to be fixed. First of all, the basal expression of AraC under uninduced conditions is too high. This becomes evident when comparing GFP expression in the uninduced Biosafety-System (orange bar) to the uninduced second part of the Biosafety-System <bbpart>BBa_I13541</bbpart> (purple bar). The strong reduction of GFP expression is likely caused by the basal transcription of the first part of the Biosafety-System containing the AraC regulator, which causes a tighter repression of the P<sub>''BAD''</sub> promoter, while there is no sufficient amount of AraC in the strain carrying the second part only.<br></p> | ||
| - | |||
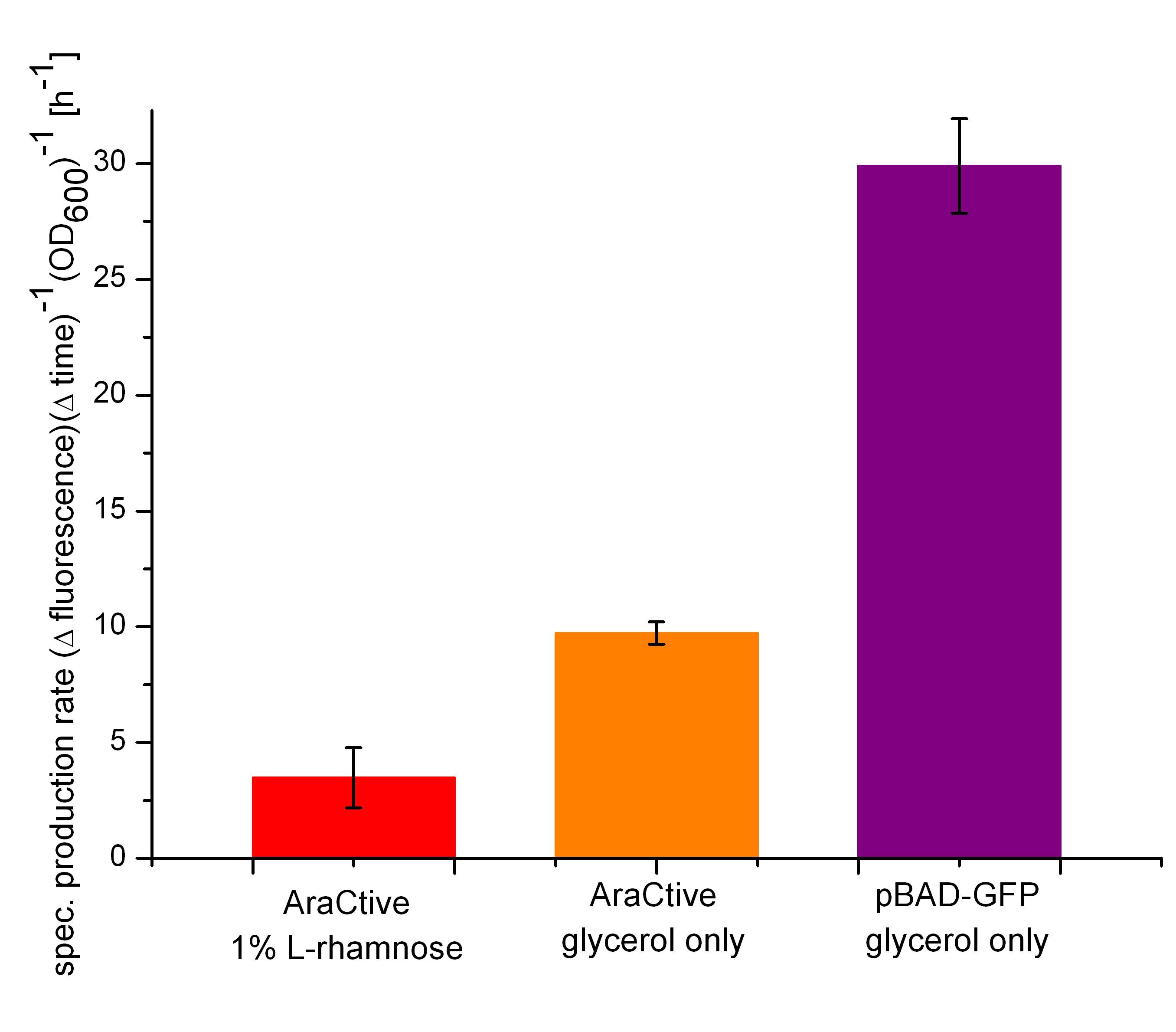
| + | [[File:Team-Bielefeld-Biosafety-System-araCtive-Resultbalken.jpg|600px|thumb|center|'''Figure 17:''' Comparision of the specific production rate of GFP. Shown are the induced (1% L-rhamnose) Biosafety-System araCtive, the uninduced Biosafety-System araCtive and the second part of the Biosafety-System (P<sub>''BAD''</sub> - GFP only).]] | ||
| + | <br> | ||
| + | <p align="justify"> | ||
| + | In [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System#Results comparison] to the other Biosafety-Systems it becomes clear, that the Biosafety-System araCtive would be probably the best choice of our three Biosafety-Systems, because of its tight repression by the P<sub>''BAD''</sub> promoter of the second part. And although the Biosafety-System works there are still some optimization remaining. | ||
| + | This high basal expression also causes problems for the double kill-switch. As can be seen in Figure 14, the Biosafety-Strain carrying the Biosafety-System araCtive can grow even without L-rhamnose induction. Thus, a tighter repression of the alanine racemase is needed. This was tried by cultivation in M9 minimal medium with glucose to obtain a tighter repression of the L-rhamnose promoter P<sub>''Rha''</sub>. In this cultivation a tighter repression of both parts of the Biosafety-System was observed, but did not significantly improve the double-kill switch mechanism. This might be due to supplementation of the preculture L-rhamnose and D-alanine. Although the cells were washed several times before cultivation, it is possible that the bacteria store D-alanine and L-rhamnose for some time, as observed for D-alanine in the [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#Characterization_of_the_D-alanine_auxotrophy D-alanine-auxotrophic strain]. Additional also the alanine racemase (Alr) could accumulate because of the induction. This could be reason for the survival of the Biosafety-Strain in the absence of L-rhamnose. A possible solution here might be the introduction of a SsrA [http://parts.igem.org/Protein_domains/Degradation degradation tag] at the C-terminus of Alr ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S#References Levchenko ''et. al'', 2003]).<br> | ||
| + | Furthermore, as glucose does not occur in this amounts in nature. Thus, a sufficiently low transcription of the rhamnose promoter P<sub>''Rha''</sub> should be independent of the carbon source and the corresponding intracellular cAMP concentration. So another improvement of the Biosafety-System could be to adjust the plasmid copy number. The plasmid pSB1C3 we used for the Biosafety-System araCtive <bbpart>BBa_K1172909</bbpart> is a high copy plasmid, resulting in 100 – 300 copies of the plasmid in each cell. Therefore it could be worthwhile to clone the Biosafety-System into a medium (15 – 20 copies) or even a low copy plasmid (2 – 12 copies) to take advantage of the double-kill switch on one hand, and to reduce the metabolic pressure triggered by the plasmid on the other hand. Moreover the lower plasmid copy number might simplify the integration of the Barnase instead of GFP. Nevertheless the Biosafety-System araCtive we constructed can already be used as a kill-switch Biosafety-System and is characterized by a higher plasmid stability compared to common Biosafety-Systems.</p><br> | ||
| Line 258: | Line 255: | ||
*Baldoma L and Aguilar J (1988) Metabolism of L-Fucose and L-Rhamnose in Escherichia coli: Aerobic-Anaerobic Regulation of L-Lactaldehyde Dissimilation [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210658/pdf/jbacter00179-0434.pdf|''Journal of Bacteriology 170: 416 - 421.'']. | *Baldoma L and Aguilar J (1988) Metabolism of L-Fucose and L-Rhamnose in Escherichia coli: Aerobic-Anaerobic Regulation of L-Lactaldehyde Dissimilation [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210658/pdf/jbacter00179-0434.pdf|''Journal of Bacteriology 170: 416 - 421.'']. | ||
*Carafa, Yves d'Aubenton Brody, Edward and Claude (1990) Thermest Prediction of Rho-independent Escherichia coli Transcription Terminators - A Statistical Analysis of their RNA Stem-Loop Structures [http://ac.els-cdn.com/S0022283699800059/1-s2.0-S0022283699800059-main.pdf?_tid=ede07e2a-2a92-11e3-b889-00000aab0f6c&acdnat=1380629809_2d1a59e395fc69c8608ab8b5aea842f7|''Journal of molecular biology 216: 835 - 858'']. | *Carafa, Yves d'Aubenton Brody, Edward and Claude (1990) Thermest Prediction of Rho-independent Escherichia coli Transcription Terminators - A Statistical Analysis of their RNA Stem-Loop Structures [http://ac.els-cdn.com/S0022283699800059/1-s2.0-S0022283699800059-main.pdf?_tid=ede07e2a-2a92-11e3-b889-00000aab0f6c&acdnat=1380629809_2d1a59e395fc69c8608ab8b5aea842f7|''Journal of molecular biology 216: 835 - 858'']. | ||
| + | *Casali, N., Preston A. (2003) E. coli Plasmid Vectors - Methods and Applications [http://www.springerprotocols.com/BookToc/doi/10.1385/1592594093 ''Methods in Molecular Biology 235]. | ||
*Cass, Laura G. and Wilcoy, Gary (1988) Novel Activation of araC Expression and a DNA Site Required for araC Autoregulation in Escherichia coli B/r [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC211425/pdf/jbacter00187-0394.pdf|''Journal of Bacteriology 170: 4174 - 4180'']. | *Cass, Laura G. and Wilcoy, Gary (1988) Novel Activation of araC Expression and a DNA Site Required for araC Autoregulation in Escherichia coli B/r [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC211425/pdf/jbacter00187-0394.pdf|''Journal of Bacteriology 170: 4174 - 4180'']. | ||
*Hamilton, Eileen P. and Lee, Nancy (1988) Three binding sites for AraC protein are required for autoregulation of araC in Escherichia coli [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC279856/pdf/pnas00258-0029.pdf|''Proc Natl Acad Sci U S A 85: 1749 - 53'']. | *Hamilton, Eileen P. and Lee, Nancy (1988) Three binding sites for AraC protein are required for autoregulation of araC in Escherichia coli [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC279856/pdf/pnas00258-0029.pdf|''Proc Natl Acad Sci U S A 85: 1749 - 53'']. | ||
| + | *Levchenko I., Grant R., Wah D., Sauer R., Baker T. (2003) Structure of a delivery protein for an AAA+ protease in complex with a peptide degradation tag. [http://ac.els-cdn.com/S109727650300323X/1-s2.0-S109727650300323X-main.pdf?_tid=4fe9541a-3fed-11e3-a15a-00000aab0f01&acdnat=1382977603_a779ff84590dc557902de5503c7e433a|''Molecular cell 12:365 - 72.] | ||
*Mossakowska, Danuta E. Nyberg, Kerstin and Fersht, Alan R. (1989) Kinetic Characterization of the Recombinant Ribonuclease from Bacillus amyloliquefaciens (Barnase) and Investigation of Key Residues in Catalysis by Site-Directed Mutagenesis [http://pubs.acs.org/doi/pdf/10.1021/bi00435a033|''Biochemistry 28: 3843 - 3850.'']. | *Mossakowska, Danuta E. Nyberg, Kerstin and Fersht, Alan R. (1989) Kinetic Characterization of the Recombinant Ribonuclease from Bacillus amyloliquefaciens (Barnase) and Investigation of Key Residues in Catalysis by Site-Directed Mutagenesis [http://pubs.acs.org/doi/pdf/10.1021/bi00435a033|''Biochemistry 28: 3843 - 3850.'']. | ||
*Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|''Journal of Bacteriology 171: 1185 - 1187.'']. | *Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|''Journal of Bacteriology 171: 1185 - 1187.'']. | ||
| Line 267: | Line 266: | ||
*Wickstrum, J.R., Santangelo, T.J., and Egan, S.M. (2005) Cyclic AMP receptor protein and RhaR synergistically activate transcription from the L-rhamnose-responsive rhaSR promoter in Escherichia coli. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251584/?report=reader|''Journal of Bacteriology 187: 6708 – 6719.'']. | *Wickstrum, J.R., Santangelo, T.J., and Egan, S.M. (2005) Cyclic AMP receptor protein and RhaR synergistically activate transcription from the L-rhamnose-responsive rhaSR promoter in Escherichia coli. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251584/?report=reader|''Journal of Bacteriology 187: 6708 – 6719.'']. | ||
| - | + | <br><br><br> | |
| - | + | ||
| - | + | ||
</div> | </div> | ||
Latest revision as of 02:46, 29 October 2013
Biosafety System AraCtive
Overview
The Biosafety-System araCtive <bbpart>BBa_K1172909</bbpart> is an improvement of the BioBrick <bbpart>BBa_K914014</bbpart> by replacing the first promoter by the rhamnose promoter PRha, integration of the alanine racemase <bbpart>BBa_K1172901</bbpart> and utilization of the regulator AraC to control the transcription of the Barnase behind the PBAD promoter. Because of the tight repression of this promoter, this system has the lowest basal transcription of the Barnase and is therefore the most active and attractive one.
Genetic Approach
Rhamnose promoter PRha
The promoter PRha (<bbpart>BBa_K914003</bbpart>) naturally regulates the catabolism of the hexose L-rhamnose. The advantage to use this promoter is its solely positive regulation. The regulon consists of the gene rhaT for the L-rhamnose transporter and the two operons rhaSR and rhaBAD.
The operon rhaSR encodes the two transcriptional activators RhaS and RhaR, who are responsible for the positive activation of the L-rhamnose catabolism, while the operon rhaBAD encodes the genes for the direct catabolism of L-rhamnose.
When L-rhamnose is present, it acts as an inducer by binding to the regulatory protein RhaR. RhaR regulates its own expression and the expression of the regulatory gene rhaS by repressing or, in the presence of L-rhamnose, activating the operon rhaSR. Normally, the expression level is modest, but it can be enhanced by a higher level of intracellular cAMP, which increases in the absence of glucose. So, in the presence of L-rhamnose and a high concentration of intracellular cAMP, the activator protein RhaR is expressed on higher level, resulting in an activation of the promoter PrhaT for an efficient L-rhamnose uptake and an activation of the operon rhaBAD. The L-rhamnose is than broken down into dihydroxyacetone phosphate and lactate aldehyde by the enzymes encoded by rhaBAD (Wickstrum et al., 2005).
A brief schematic summary of the regulation is shown in Figure 1.
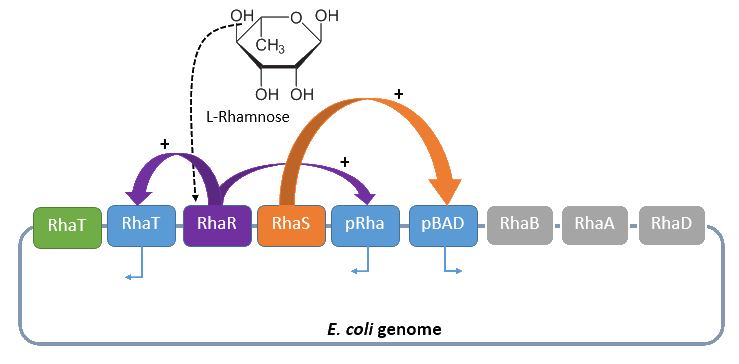
Dihydroxyacetone phosphate can be metabolized in the glycolysis pathway, while lactate aldehyde is oxidized to lactate under aerobic conditions and reduced to L-1,2,-propandiol under anaerobic conditions.
The degradation of L-rhamnose can be separated in three steps. In the first step the L-rhamnose is turned into L-rhamnulose by an isomerase (gene rhaA). L-rhamnulose is in turn phosphorylated to L-rhamnulose-1-phosphate by a kinase (gene rhaB) and the sugar phosphate is finally hydrolyzed by an aldolase (gene rhaD) to dihydroxyacetone phosphate and lactate aldehyde (Baldoma et al., 1988).
For our Safety-System araCtive, the rhamnose promoter PRha is used to control the expression of the repressor AraC and the essential alanine racemase, because this promoter has an even lower basal transcription then the arabinose promoter PBAD. This is needed to tightly repress the expression of the alanine racemase (alr) and thereby take advantage of the double-kill switch. Although the rhamnose promoter PRha is characterized by a very low basal transcription it can not be used for the control of the toxic RNase Ba (Barnase), which ist needed for the alanine racemase (alr), the other part of the double-kill switch. In conclusion this promoter is the best choice for the first part of the Biosafety-System, as it is tightly repressed in the absence of L-rhamnose, but activated in its presence.
Regulator AraC
The araC gene is naturally found in E. coli and is coding for the 32 kDa AraC protein, which regulates the expression of the genes required for the uptake and catabolism of the pentose L-arabinose. The genes for the catabolism of arabinose are under the control of the arabinose promoter PBAD, which is both positively and negatively regulated by AraC. Naturally in the presence of L-arabinose the expression of those genes is activated, while it is repressed in its absence. In addition the AraC protein regulates its own transcription under the control of the so called PC promoter. Compared to the PBAD promoter the PC promoter and the associated araC gene are therefore transcribed in opposite direction (Schleif, 2010).
Naturally in the absence of L-arabinose the AraC protein is found in its repressor conformation, binding to the O2-region and I1-region. Binding of two AraC proteins to these domains results in a protein-protein-interaction of AraC forming a dimer. This AraC dimer is responsible for the tight repression of the PC and the PBAD promoter by forming a DNA-loop to inhibit the initiation of the RNA-polymerase (see Figure 3 above).
When L-arabinose is present in the media the operon changes to the active form. Following this conditions the AraC protein functions as activator by binding to the I1- and I2-region of the operator region. This causes transcription of the genes behind the PBAD and the PC promoter. Depending on the intracellular level of cAMP the transcription of both the genes behind the promoters PBAD and PC can even be enhanced. In the absence of glucose the cell reaches a high level of cAMP as a signal molecule. cAMP binds as the typically CRP-cAMP complex binds in front of the AraC regulator and increases the binding affinity of the RNA-polymerase, so that the genes behind the PBAD promoter are expressed on higher levels. Besides the transcription affinity of the PBAD promoter, the AraC protein also regulates the genes araE and the operon consisting the genes araFGH, which are necessary for an efficient uptake of L-arabinose (Cass, 1988).
The auto-regulation of the araC gene itself works both in the presence and absence of L-arabinose. In presence of L-arabinose, the DNA-loop which prevents also the transcription of the PBAD promoter also inhibits the transcription of araC. When L-arabinose is present and the AraC concentration is high enough, it auto-regulates its own transcription by dimerization between the araO1- and the araO2-region. This leads to an other DNA-loop, inhibiting solely the transcription of the genes behind the PC promoter (Hamilton, 1988).
For our Biosafety-System we decided to use the arabinose PBAD promoter, because this promoter is tightly regulated by AraC and shows no basal transcription in the complete absence of AraC. Additionally this promoter is repressed by glucose and basal transcription is activated in the presence of cAMP.
Alanine racemase Alr
The alanine racemase Alr (EC 5.1.1.1) from the Gram-negative enteric bacteria Escherichia coli is a racemase, which catalyses the reversible conversion of L-alanine into the enantiomer D-alanine (see Figure 4). For this reaction, the cofactor pyridoxal-5'-phosphate (PLP) is necessary. The constitutively expressed alanine racemase (alr) is naturally responsible for the accumulation of D-alanine. This compound is an essential component of the bacterial cell wall, because it is used for the cross-linkage of peptidoglycan (Walsh, 1989).
The usage of D-alanine instead of a typically L-amino acid prevents cleavage by peptidases. However, a lack of D-alanine causes to a bacteriolytic characteristics. In the absence of D‑alanine dividing cells will lyse rapidly. This fact is used for our Biosafety-Strain, a D-alanine auxotrophic mutant (K-12 ∆alr ∆dadX). The Biosafety-Strain grows only with a plasmid containing the alanine racemase (<bbpart>BBa_K1172901</bbpart>) to complement the D-alanine auxotrophy. Consequently the alanine racemase is essential for bacterial cell division. This approach guarantees a high plasmid stability, which is extremely important when the plasmid contains a toxic gene like the Barnase. In addition this construction provides the possibility for the implementation of a double kill-switch system. Because if the expression of the alanine racemase is repressed and there is no D-alanine supplementation in the medium, cells will not grow.
Terminator
Terminators are essential to terminate transcription of an operon. In procaryotes, two types of terminators exist. The rho-dependent and the rho-independent terminator. Rho-independent terminators are characterized by their stem-loop forming sequence. In general, the terminator-region can be divided into four regions. The first region is GC-rich and constitutes one half of the stem. This region is followed by the loop-region and another GC-rich region that makes up the opposite part of the stem. The terminator closes with a poly uracil region, which destabilizes the binding of the RNA-polymerase. The stem-loop of the terminator causes a distinction of the DNA and the translated RNA. Consequently the binding of the RNA-polymerase is cancelled and the transcription ends after the stem-loop (Carafa et al., 1990).
For our Bioafety-System araCtive the terminator is necessary to avoid that the expression of the genes under control of the rhamnose promoter PRha, like the Repressor AraC and the alanine racemase (alr) results in the transcription of the genes behind the arabinose promoter PBAD, which contains the toxic Barnase <bbpart>BBa_K1172904</bbpart> and would lead to cell death.
Arabinose promoter PBAD
The arabinose promoter PBAD controls the expression of the genes, which are necessary for the catabolism of the pentose arabinose. The expression is regulated by the bifunctional regulator AraC. The genes of this operon convert the pentose to xylose, which can be converted into fructose-1,6-diphosphate and subsequently be fed into glycolysis. The conversion of L-arabinose is realized in three steps. First the isomerase (AraA) catalyzes the reaction from L-arabinose to the isomer L-ribulose. In the second step the ribulose kinase (AraB) phosphorylates the L-ribulose to L-ribulose-phosphate. For this reaction ATP is needed. Last, the L-ribulose-phosphate is converted to D-xylose-5-phosphat by L-ribulose-5-phosphate-epimerase (AraD, Schleif, 2010).
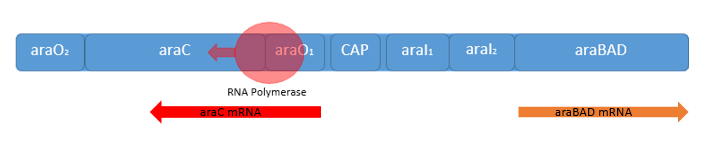
For our Safety-System we used the arabinose promoter PBAD for the regulation of the toxic gene product Barnase. As this promoter has very low basal transcription and as the expression of the genes behind this promoter is strict depending on the presence of AraC for activating the transcription, it is possible to use the arabinose promoter PBAD to control the expression of a toxic gene product without causing cell death.
RNase Ba (Barnase)
The Barnase (EC 3.1.27) is a 12 kDa extracellular microbial ribonuclease, which is naturally found in the Gram-positive soil bacteria Bacillus amyloliquefaciens and consists of a single chain of 110 amino acids. The Barnase (RNase Ba) catalyses the cleavage of single stranded RNA, preferentially behind Gs. In the first step of the RNA-degradation a cyclic intermediate is formed by transesterification and afterwards this intermediate is hydrolyzed yielding in a 3'-nucleotide (Mossakowska et al., 1989).
In Bacillus amyloliquefaciens, the activity of the Barnase (RNase Ba) is inhibited intracellular by an inhibitor called barstar. Barstar consists of only 89 amino acids and binds with a high affinity to the toxic Barnase. This prevents the cleavage of the intracellular RNA in the host organism (Paddon et al., 1989). Therefore the Barnase normally acts only outside the cell and is translocated under natural conditions. For the Biosafety-System araCtive we modified the enzyme by cloning only the sequence responsible for the cleavage of the RNA, leaving out the N-terminal signal peptide part.
As shown in Figure 8 below, the transcription of the DNA, which encodes the Barnase produces a 474 nt RNA. The translation of the RNA starts about 25 nucleotides downstream from the transcription start and can be divided into two parts. The first part (colored in orange) is translated into a signal peptide at the amino-terminus of the Barnase coding RNA. This part is responsible for the extracellular translocation of the RNase Ba, while the peptide sequence for the active Barnase starts 142 nucleotides downstream from the transcription start (colored in red).
For the Biosafety-System araCtive, we only used the part (<bbpart>BBa_K1172904</bbpart>) of the Barnase encoding the catalytic domain without the extracellular translocation signal of the toxic gene product. Translation of the barnase gene leads to rapid cell death if the expression of the Barnase is not repressed by the repressor AraC of our Biosafety-System.
Biosafety-System araCtive
Combining the gene described above with the Biosafety-Strain K-12 ∆alr ∆dadX results in a powerful device, allowing us to control the bacterial cell division. The control of the bacterial growth is possible either active or passive. Active by inducing the PBAD promoter with L-arabinose and passive by the induction of L-rhamnose. The passive control makes it possible to control the bacterial cell division in a defined closed environment, like the MFC, by continuously adding L-rhamnose to the medium. As shown in the Figure 9 below, this leads to an expression of the essential alanine racemase (alr) and the AraC regulator, so that the expression of the RNase Ba is repressed.
In the event that bacteria exit the defined environment of the MFC or L-rhamnose is not added to the medium any more, both the expression of the alanine racemase (Alr) and the AraC regulatordecrease, so that the expression of the toxic RNase Ba (Barnase) begins. The cleavage of the intracellular RNA by the Barnase and the lack of synthesized D-alanine, caused by the repressed alanine racemase inhibit the cell division and makes sure that the bacteria can only grow in the defined environment or the device of choice respectively.
Results
Characterization of the arabinose promoter PBAD
First of all, the arabinose promoter PBAD was characterized to get a first impression of its basal transcription rate. Therefore the bacterial growth was investigated under the pressure of the unrepressed PBAD promoter on different carbon source using M9 minimal medium with either glucose or glycerol. The transcription rate was determined by fluorescence measurement of GFP <bbpart>BBa_E0040</bbpart> transcribed via the PBAD promoter using the BioBrick <bbpart>BBa_I13541</bbpart>.
As shown in Figure 12 below, the bacteria grew better on glucose than on glycerol. This is caused by the fact that glucose is the better energy source of this two, because glycerol enters glycolysis at a later step and therefore delivers less energy (in addition to the additional ATP consumption to drive glycerol uptake). For the investigation of the basal transcription the fluorescence measurements, shown in Figure 13, is more interesting.
Both the wild type E. coli K-12 and the strain E. coli K-12 ∆alr ∆dadX ∆araC show about the same fluorescence on glucose (red and orange curve) but differ on glycerol. This can be explained by the fact that growth on glucose does not cause an additional induction of the arabinose promoter PBAD, while glycerol does. In the presence of glucose, the intracellular concentration of cAMP is low. The absence of cAMP results in inactive CAP (carbon utilization activator protein) and thus no induction of inefficient catabolisms like the catabolism of arabinose. Therefore, glucose is catabolized by the bacteria first, resulting in an optimal growth. In the absence of glucose, the cAMP level increases which enhances via CAP-cAMP the transcription of most operons encoding enzymes for the catabolism of alternative carbon sources. Therefore, the expression of GFP under the control of the PBAD promoter increases on glycerol. Another difference can be seen between the wild type (black curve) and the strain K-12 ∆alr ∆dadX ∆araC (blue curve).
The Biosafety-Strain shows lower expression then the wild type, but has about the same growth rate, according to Figure 11. The reason for this characteristic is caused by the deleted AraC protein in the Biosafety-Strain. Because the AraC protein functions not only as are repressor but also as an activator the transcription rate decreases in the Biosafety-Strain.
The effect that glucose represses the transcription of the PBAD promoter becomes more obvious by calculating of the specific production rates of GFP, as shown in Figure 13. The specific production rates were calculated via equation (1) :
With the specific production rate of GFP, it can be demonstrated that GFP synthesis rates differ extremely between the cultivation on glucose and glycerol. The specific production rate is constantly very low when using glucose as carbon source, but shows a higher more unstable level in the cultivation with glycerol.
Because the specific production rate was calculated between every single measurement point, the curve in Figure 13 is not smoothed and so the fluctuations have to be ignored, as they do not stand for are real fluctuations in the expression of GFP. They are caused by measurement variations in the growth curve and the fluorescence curve. And as neither of this curves are ideal, the fluctuations are the result. Nevertheless this graph shows clearly the difference between the two carbon sources.
Characterization of the Biosafety-System araCtive
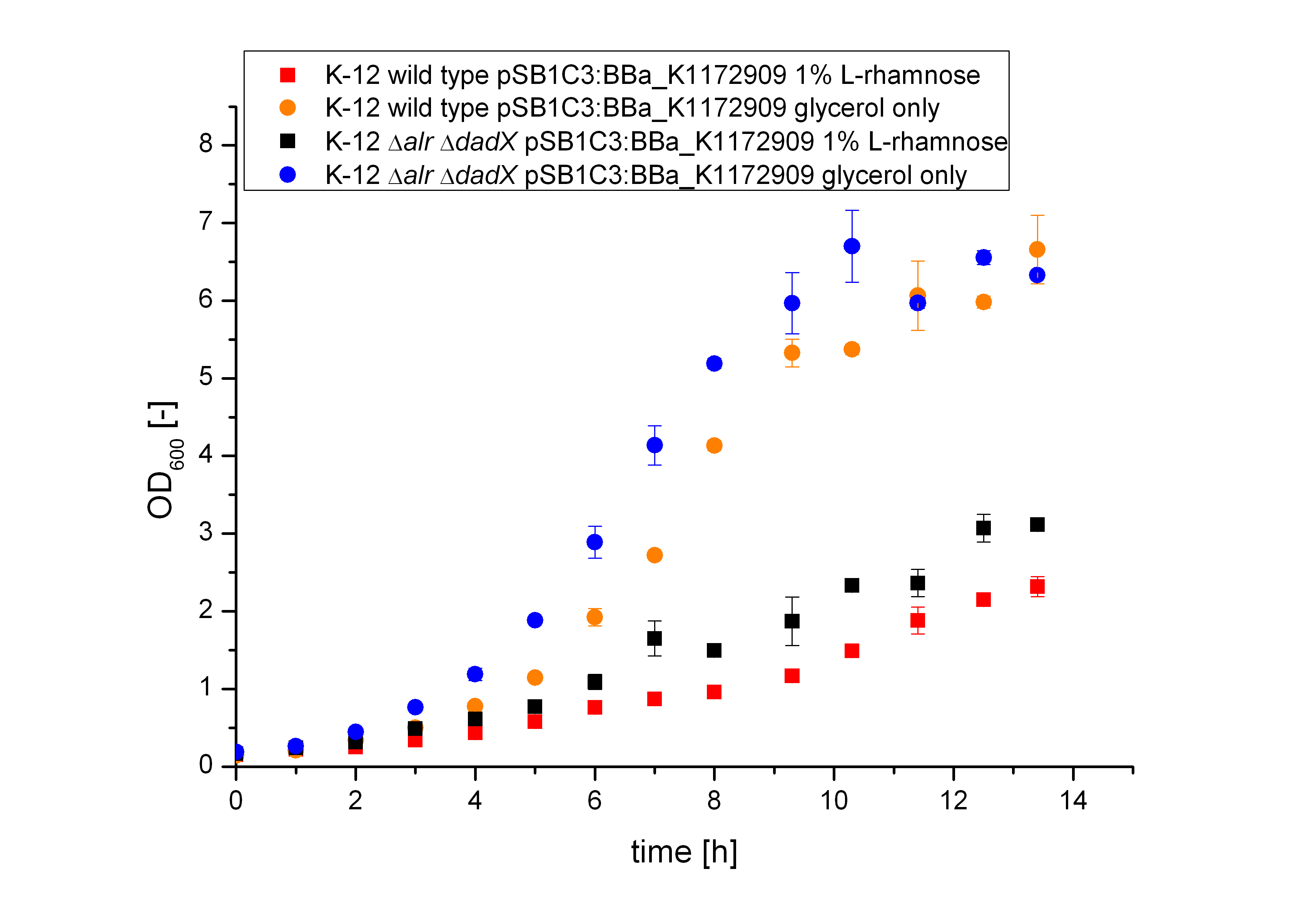
The Biosafety-System araCtive was characterized on M9 minimal medium using glycerol as carbon source. As for the characterization of the pure arabinose promoter PBAD above, the bacterial growth and the fluorescence of GFP <bbpart>BBa_E0040</bbpart> was measured. Therefore, the wild type and the Biosafety-Strain E. coli K-12 ∆alr ∆dadX, both containing the Biosafety-Plasmid <bbpart>BBa_K1172909</bbpart>, were cultivated once with the induction of 1% L-rhamnose and once only on glycerol.
It becomes obvious (Figure 14) that the bacteria, induced with 1 % L-rhamnose (red and black curve) grow significantly slower than on pure glycerol (orange and blue curve). This is attributed to the high metabolic burden encountered by the induced bacteria. The expression of the repressor AraC and the alanine racemase (Alr) simultaneously causes a high strain on the cells, so that they grow slower than the uninduced cells, which express only GFP.
Comparing the bacterial growth with the fluorescence in Figure 15, it can be seen that the fluorescence seems to follow the same trend than the bacterial growth. The uninduced cells show approximately an exponential rise of fluorescence, while in comparision the fluorescence of the induced bacteria increases only slowly.
From the data presented above it can not be determined if the expression of the repressor AraC does affect the transcription of GFP or not. The slower growth of the bacteria is a first indication that the repressor AraC and the alanine racemase (Alr) are highly expressed, but the growth of the bacteria shows nearly the same kinetics as the fluorescence. So it could be possible that the repressor does not affect the expression level of GFP under the control of the arabinose promoter PBAD. This becomes more clear by the calculation of the specific production rate of GFP by equation (1) . As shown in Figure 17 below the specific production rate differs clearly between the uninduced Biosafety-System and the Biosafety-System induced by 1% L-rhamnose. The production of GFP in the presence of L-rhamnose (red curve) is always lower than in its absence (orange curve), so that the expression of GFP is repressed in the presence of L-rhamnose.
Because the specific production rate of GFP was calculated between every single measurement point, the curve in Figure 16 is not smoothed and so the fluctuations have to be ignored, as they do not stand for are real fluctuations in the expression of GFP. They are caused by measurement variations in the growth curve and the fluorescence curve. But there is a clear tendency that the production of GFP is significantly lower when the bacteria are induced with 1% L-rhamnose. So the Biosafety-System araCtive works.
Conclusions
It could be demonstrated that the Biosafety-System arActive works in principle. For example, the expression level of GFP is decreased in the presence of L-rhamnose, as can be seen in Figure 17, depicting the specific production rates of GFP after 7,5 hours. The induction of the Biosafety-System with L-rhamnose leads to a tight repression of the expression of GFP (red bar) when compared to the uninduced state (orange bar). Therefore the second part of the Biosafety-System, which contains normally the toxic RNase Ba <bbpart>BBa_K1172904</bbpart> can be used as a kill-switch regulated by the pentose L-rhamnose. In comparison to the other Biosafety-Systems it becomes clear, that the Biosafety-System araCtive would be probably the best choice of our three Biosafety-Systems, because of its tight repression by the PBAD promoter of the second part.
But a number of problems remain to be fixed. First of all, the basal expression of AraC under uninduced conditions is too high. This becomes evident when comparing GFP expression in the uninduced Biosafety-System (orange bar) to the uninduced second part of the Biosafety-System <bbpart>BBa_I13541</bbpart> (purple bar). The strong reduction of GFP expression is likely caused by the basal transcription of the first part of the Biosafety-System containing the AraC regulator, which causes a tighter repression of the PBAD promoter, while there is no sufficient amount of AraC in the strain carrying the second part only.
In comparison to the other Biosafety-Systems it becomes clear, that the Biosafety-System araCtive would be probably the best choice of our three Biosafety-Systems, because of its tight repression by the PBAD promoter of the second part. And although the Biosafety-System works there are still some optimization remaining.
This high basal expression also causes problems for the double kill-switch. As can be seen in Figure 14, the Biosafety-Strain carrying the Biosafety-System araCtive can grow even without L-rhamnose induction. Thus, a tighter repression of the alanine racemase is needed. This was tried by cultivation in M9 minimal medium with glucose to obtain a tighter repression of the L-rhamnose promoter PRha. In this cultivation a tighter repression of both parts of the Biosafety-System was observed, but did not significantly improve the double-kill switch mechanism. This might be due to supplementation of the preculture L-rhamnose and D-alanine. Although the cells were washed several times before cultivation, it is possible that the bacteria store D-alanine and L-rhamnose for some time, as observed for D-alanine in the D-alanine-auxotrophic strain. Additional also the alanine racemase (Alr) could accumulate because of the induction. This could be reason for the survival of the Biosafety-Strain in the absence of L-rhamnose. A possible solution here might be the introduction of a SsrA [http://parts.igem.org/Protein_domains/Degradation degradation tag] at the C-terminus of Alr (Levchenko et. al, 2003).
Furthermore, as glucose does not occur in this amounts in nature. Thus, a sufficiently low transcription of the rhamnose promoter PRha should be independent of the carbon source and the corresponding intracellular cAMP concentration. So another improvement of the Biosafety-System could be to adjust the plasmid copy number. The plasmid pSB1C3 we used for the Biosafety-System araCtive <bbpart>BBa_K1172909</bbpart> is a high copy plasmid, resulting in 100 – 300 copies of the plasmid in each cell. Therefore it could be worthwhile to clone the Biosafety-System into a medium (15 – 20 copies) or even a low copy plasmid (2 – 12 copies) to take advantage of the double-kill switch on one hand, and to reduce the metabolic pressure triggered by the plasmid on the other hand. Moreover the lower plasmid copy number might simplify the integration of the Barnase instead of GFP. Nevertheless the Biosafety-System araCtive we constructed can already be used as a kill-switch Biosafety-System and is characterized by a higher plasmid stability compared to common Biosafety-Systems.
References
- Baldoma L and Aguilar J (1988) Metabolism of L-Fucose and L-Rhamnose in Escherichia coli: Aerobic-Anaerobic Regulation of L-Lactaldehyde Dissimilation [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210658/pdf/jbacter00179-0434.pdf|Journal of Bacteriology 170: 416 - 421.].
- Carafa, Yves d'Aubenton Brody, Edward and Claude (1990) Thermest Prediction of Rho-independent Escherichia coli Transcription Terminators - A Statistical Analysis of their RNA Stem-Loop Structures [http://ac.els-cdn.com/S0022283699800059/1-s2.0-S0022283699800059-main.pdf?_tid=ede07e2a-2a92-11e3-b889-00000aab0f6c&acdnat=1380629809_2d1a59e395fc69c8608ab8b5aea842f7|Journal of molecular biology 216: 835 - 858].
- Casali, N., Preston A. (2003) E. coli Plasmid Vectors - Methods and Applications [http://www.springerprotocols.com/BookToc/doi/10.1385/1592594093 Methods in Molecular Biology 235].
- Cass, Laura G. and Wilcoy, Gary (1988) Novel Activation of araC Expression and a DNA Site Required for araC Autoregulation in Escherichia coli B/r [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC211425/pdf/jbacter00187-0394.pdf|Journal of Bacteriology 170: 4174 - 4180].
- Hamilton, Eileen P. and Lee, Nancy (1988) Three binding sites for AraC protein are required for autoregulation of araC in Escherichia coli [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC279856/pdf/pnas00258-0029.pdf|Proc Natl Acad Sci U S A 85: 1749 - 53].
- Levchenko I., Grant R., Wah D., Sauer R., Baker T. (2003) Structure of a delivery protein for an AAA+ protease in complex with a peptide degradation tag. [http://ac.els-cdn.com/S109727650300323X/1-s2.0-S109727650300323X-main.pdf?_tid=4fe9541a-3fed-11e3-a15a-00000aab0f01&acdnat=1382977603_a779ff84590dc557902de5503c7e433a|Molecular cell 12:365 - 72.]
- Mossakowska, Danuta E. Nyberg, Kerstin and Fersht, Alan R. (1989) Kinetic Characterization of the Recombinant Ribonuclease from Bacillus amyloliquefaciens (Barnase) and Investigation of Key Residues in Catalysis by Site-Directed Mutagenesis [http://pubs.acs.org/doi/pdf/10.1021/bi00435a033|Biochemistry 28: 3843 - 3850.].
- Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|Journal of Bacteriology 171: 1185 - 1187.].
- Schleif, Robert (2010) AraC protein, regulation of the L-arabinose operon in Escherichia coli, and the light switch mechanism of AraC action [http://gene.bio.jhu.edu/Ourspdf/127.pdf|FEMS microbial reviews 34: 779 - 796.].
- Voss, Carsten Lindau, Dennis and Flaschel, Erwin (2006) Production of Recombinant RNase Ba and Its Application in Downstream Processing of Plasmid DNA for Pharmaceutical Use [http://onlinelibrary.wiley.com/doi/10.1021/bp050417e/pdf|Biotechnology Progress 22: 737 - 744.].
- Walsh, Christopher (1989) Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. [http://www.jbc.org/content/264/5/2393.long|Journal of biological chemistry 264: 2393 - 2396.]
- Wickstrum, J.R., Santangelo, T.J., and Egan, S.M. (2005) Cyclic AMP receptor protein and RhaR synergistically activate transcription from the L-rhamnose-responsive rhaSR promoter in Escherichia coli. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251584/?report=reader|Journal of Bacteriology 187: 6708 – 6719.].
 "
"