Team:TU-Munich/Results/Recombinant
From 2013.igem.org
(→Activity determination of Luminescense) |
(→Structure of the Nano Luciferase) |
||
| Line 145: | Line 145: | ||
===Structure of the Nano Luciferase=== | ===Structure of the Nano Luciferase=== | ||
| - | [[File:TUM13_CD_nLuc.png|thumb|right| | + | [[File:TUM13_CD_nLuc.png|thumb|right|300px| Figure xx:]] |
| + | [[File:TUM13_Annimated_test.gif|thumb|right|300px| Figure xx:]] | ||
There is no structure availible for the NanoLuc in the [http://www.rcsb.org/pdb/home/home.do Protein Data Bank]. In our [https://2013.igem.org/Team:TU-Munich/Modeling/Protein_Predictions protein modelling] we used homolgy search and identified the structure [http://www.rcsb.org/pdb/explore/explore.do?structureId=3PPT 3ppt_A] as the solved structure with the highest homology to the NanoLuc which has only 21% identify with a similarity of only 0.359. The result of the homology search is shown as annimated gif in Figure xx (please see our [https://2013.igem.org/Team:TU-Munich/Results/How_To How To] for an introduction). The protein was dialysed against 1x CD-buffer and subsequently a circular dichroism spectroscopy was tanken (learn about [https://2013.igem.org/Team:TU-Munich/Notebook/Methods#Circular_Dichroism_Spectroscopy CD spectroscopy]). The CD spectrum was used to predict the secondary structure content of the NanoLuc which could be determined to 35.1% helix, 27.6% b-strand, 18.5% turn and 18.8% random. As there is only a poor homology present, a detailed comparison of the determined and the predicted secondary structure is not possible. But it can be stated that both show a balanced content of different secondary structures and that the produced protein is present in an folded conformation. The mixed secondary structure content is also in consistance with the predicted secondary structure shown in the [https://2013.igem.org/Team:TU-Munich/Results/Software AutoAnnotator] sequence window (click on show). | There is no structure availible for the NanoLuc in the [http://www.rcsb.org/pdb/home/home.do Protein Data Bank]. In our [https://2013.igem.org/Team:TU-Munich/Modeling/Protein_Predictions protein modelling] we used homolgy search and identified the structure [http://www.rcsb.org/pdb/explore/explore.do?structureId=3PPT 3ppt_A] as the solved structure with the highest homology to the NanoLuc which has only 21% identify with a similarity of only 0.359. The result of the homology search is shown as annimated gif in Figure xx (please see our [https://2013.igem.org/Team:TU-Munich/Results/How_To How To] for an introduction). The protein was dialysed against 1x CD-buffer and subsequently a circular dichroism spectroscopy was tanken (learn about [https://2013.igem.org/Team:TU-Munich/Notebook/Methods#Circular_Dichroism_Spectroscopy CD spectroscopy]). The CD spectrum was used to predict the secondary structure content of the NanoLuc which could be determined to 35.1% helix, 27.6% b-strand, 18.5% turn and 18.8% random. As there is only a poor homology present, a detailed comparison of the determined and the predicted secondary structure is not possible. But it can be stated that both show a balanced content of different secondary structures and that the produced protein is present in an folded conformation. The mixed secondary structure content is also in consistance with the predicted secondary structure shown in the [https://2013.igem.org/Team:TU-Munich/Results/Software AutoAnnotator] sequence window (click on show). | ||
Revision as of 22:02, 28 September 2013
Characterization of recombinant effector proteins
Text. See proteinbiochemical methods for further informatons.
| Protein | BioBrick | RFC | Affinity tag | Size [kDa] | Disulphid bridges | comment |
|---|---|---|---|---|---|---|
| Eryhtromycin esterase (EreB) | <partinfo>BBa_K1159000</partinfo> | RFC25 | c-term. Streptag II | |||
| Laccase | <partinfo>BBa_K1159002</partinfo> | RFC25 | c-term. Streptag II | |||
| Nano Luciferase | <partinfo>BBa_K1159001</partinfo> | RFC25 | c-term. Streptag II | |||
| XylE | <partinfo>BBa_E0040</partinfo> | |||||
| PP1 | <partinfo>BBa_K1159004</partinfo> | |||||
| YFP_TEV_CFP | <partinfo>BBa_K1159112</partinfo> |
Eryhtromycin Esterase
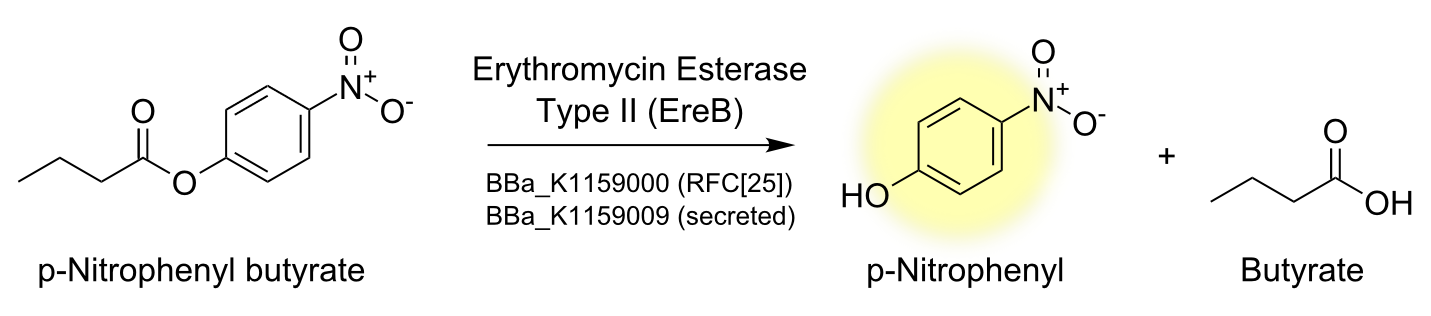
[...] description [...] reaction
| Protein data table for BioBrick BBa_ automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 25, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein) ATGGCCGGCAGGTTCGAA ... GTTTATGAAACCGGT ORF from nucleotide position -8 to 1260 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
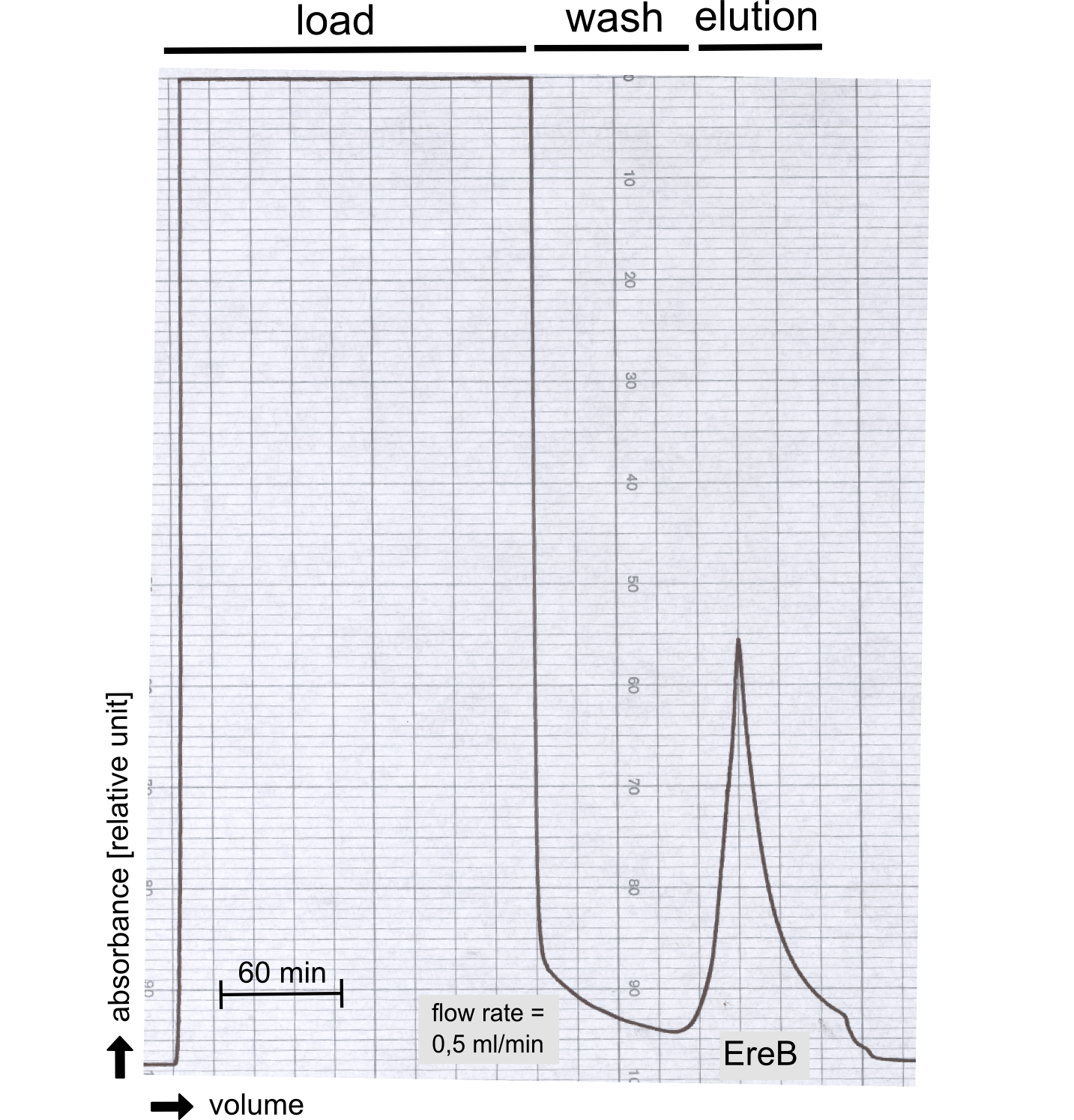
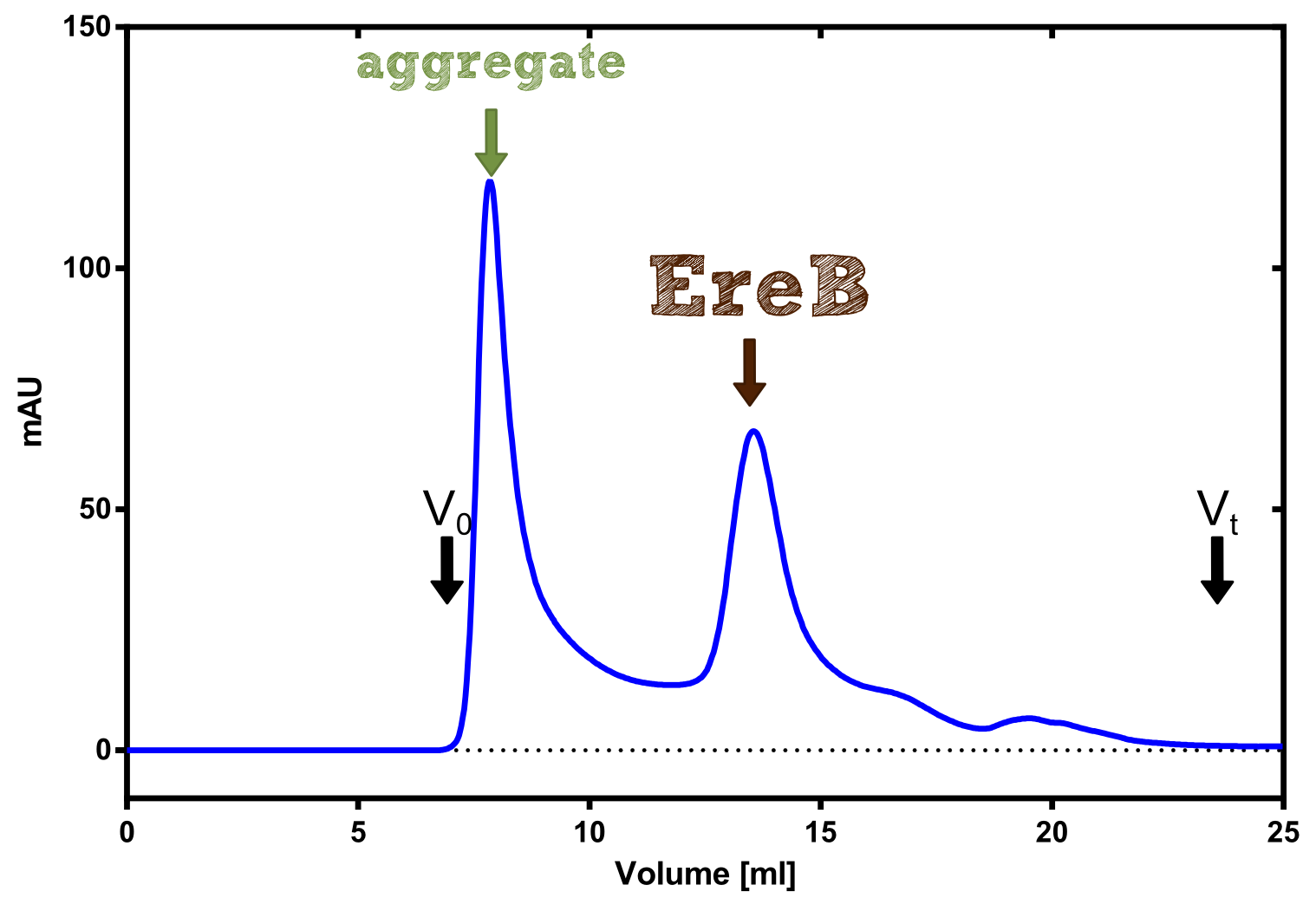
Analytical preparation
Degradation of a chromogenic esterase substrate: 4-Nitrophenyl butyrate
Degradation of Erythromycin
Reaction conditions HPLC Kirby Bauer-Assay
[...] Characterization
Laccase
[...] description [...] reaction [...] production
| Protein data table for BioBrick BBa_ automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 25, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein) ATGGCCGGCAACCTAGAA ... GATATCATCACCGGT ORF from nucleotide position -8 to 1530 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
Laccase a is a secreted enzyme
Analytical präparation
[...] Characterization
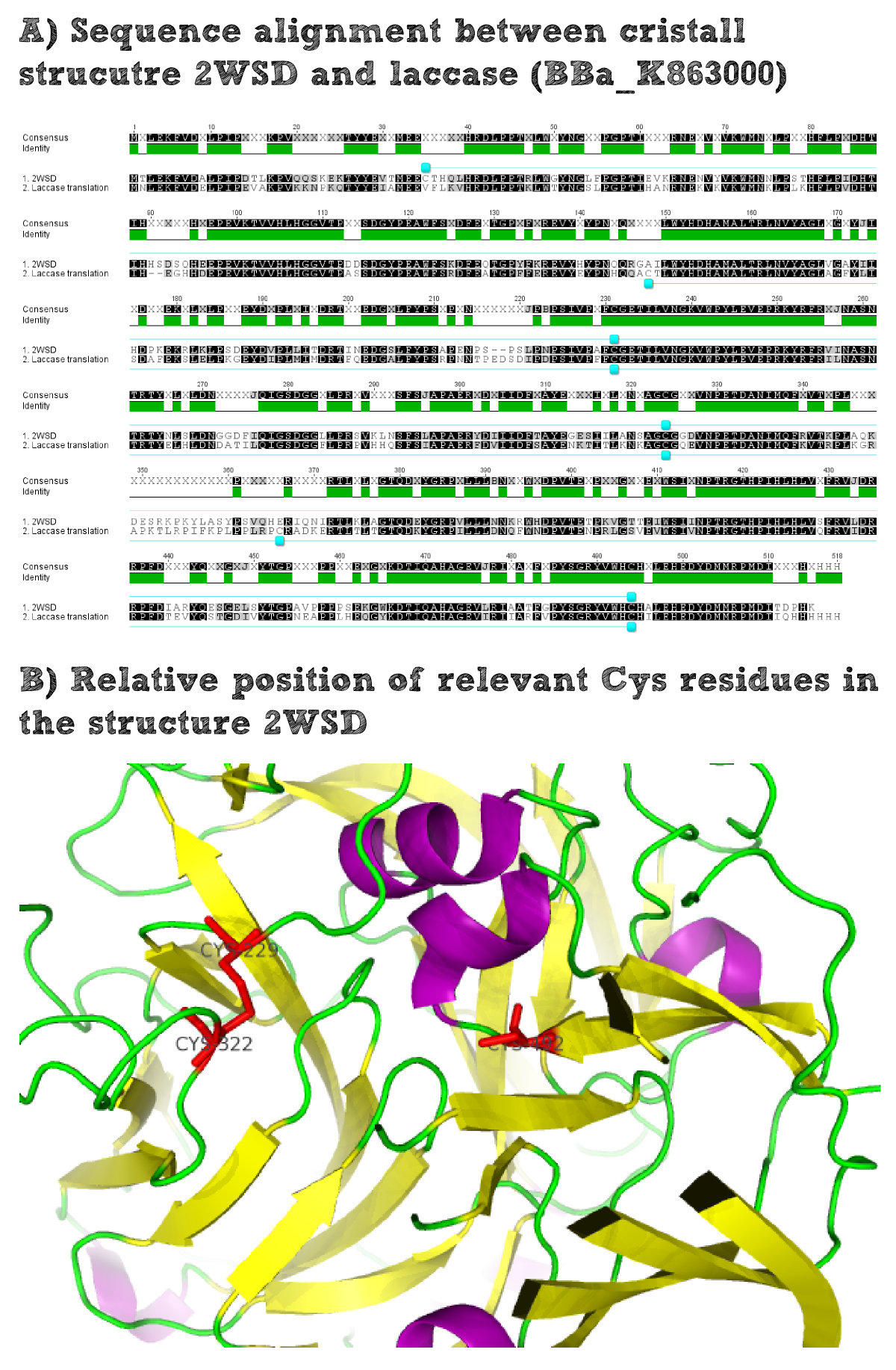
Structural consideration of Laccase from B. pumilus
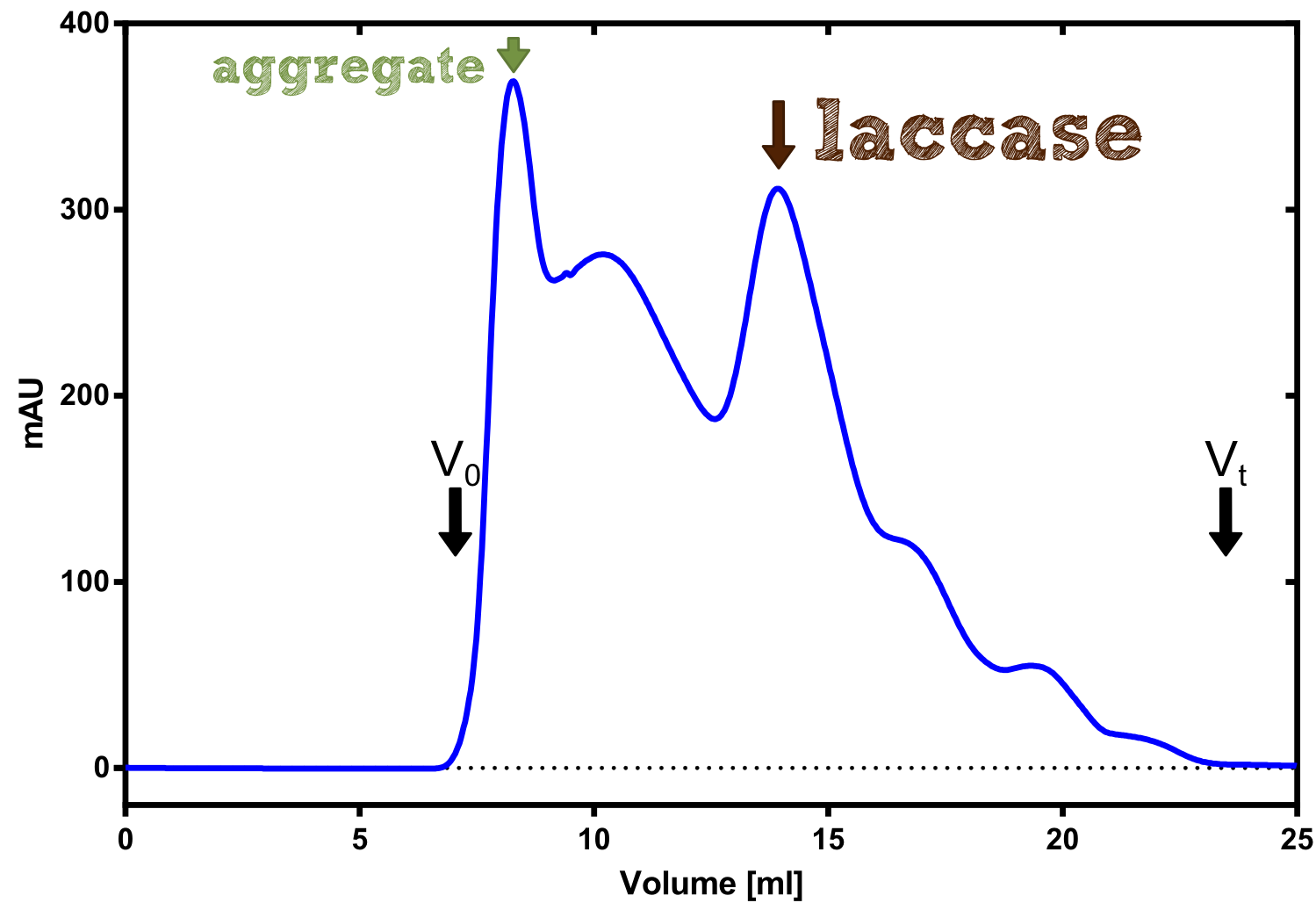
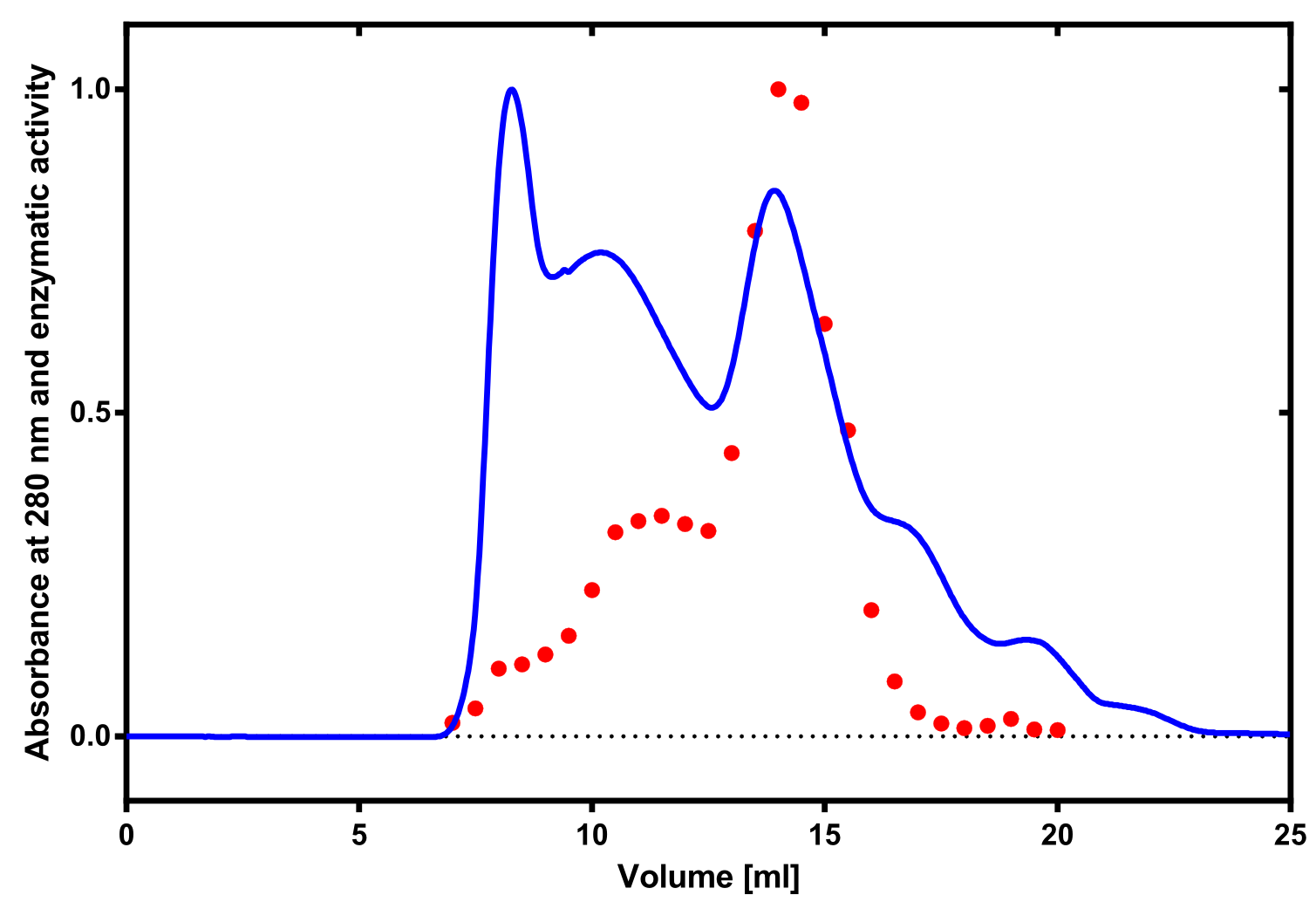
Activity determination using ABTS
The enzymatic activity of the purified laccae was determined by the ABTS-assay. In a first pre experiment the appropriate dilution factor was determined to 100-fold. The elution fractions obtained from size exclusion chromatography were diluted 1:100 in PBS and in an ELISA plate 100 µl of the enzyme and 100 µl of ABTS substrate were mixed and a kinetic measurement at 405 nm was performed. The absorption at 280 nm in the SEC chromatogramm (blue) identifies three main protein peaks, with a first peak corresponding to aggregated protein, a shoulder which also corresponds to higher molecular protein and a single peak which was proposed to be the monomeric laccase. The relative activity obtained for the different elution fractions was plotted in the same diagramm and shows a clear peak which matches the laccase peak in the SEC. Beside this major peak a second smaller peak of active fraction was visible which appeared in earlier elution fractions and might correspond to dimerized laccase. As the laccase is a secreted enzyme which also bears disluphide bonds it was produced in the cytoplasm and subsequently it was oxidized to form the proper disulphide bond. As this process might be only partial there is a possiblity for the formation of disulphid dimers. Never the less the fractions 14 to 17 were pooled for further experiments as they showed the highest enzymatic activity. The protein concentration of the pooled fraction was determined to 0.48 mg/ml after SEC.
Oxidation of relevant xenobiotics
Diclofenac
Estradiol
Nano Luciferase
The Nano Luciferase (NanoLuc) which was introduced in 2013 by Promega is a new member of the luciferase reporter gene/protein familiy and shows some advantages compared to the other family members. The NanoLuc is very small (19 kDa) compared to the firefly luciferase (61 kDa) and the Renilla luciferase (36 kDa). On the other hand it is also said that the specific activity of the NanoLuc is about 150-fold stronger compared to conventional luciferases and the background caused by autoluminescense of the substrate shel be smaller.
| Protein data table for BioBrick BBa_K1159001 automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 10: (underlined part encodes the protein) ATGGCCGGC ... GCTACCGGTTAA ORF from nucleotide position 1 to 525 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC 25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
| Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges | ||||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
Alignments (obtained from PredictProtein.org)
| ||||||||||||||||||||||||||||||||||||||||||||||
| Predictions (obtained from PredictProtein.org) | ||||||||||||||||||||||||||||||||||||||||||||||
Subcellular Localization (reliability in brackets)
| Gene Ontology (reliability in brackets)
| |||||||||||||||||||||||||||||||||||||||||||||
Predicted features:
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
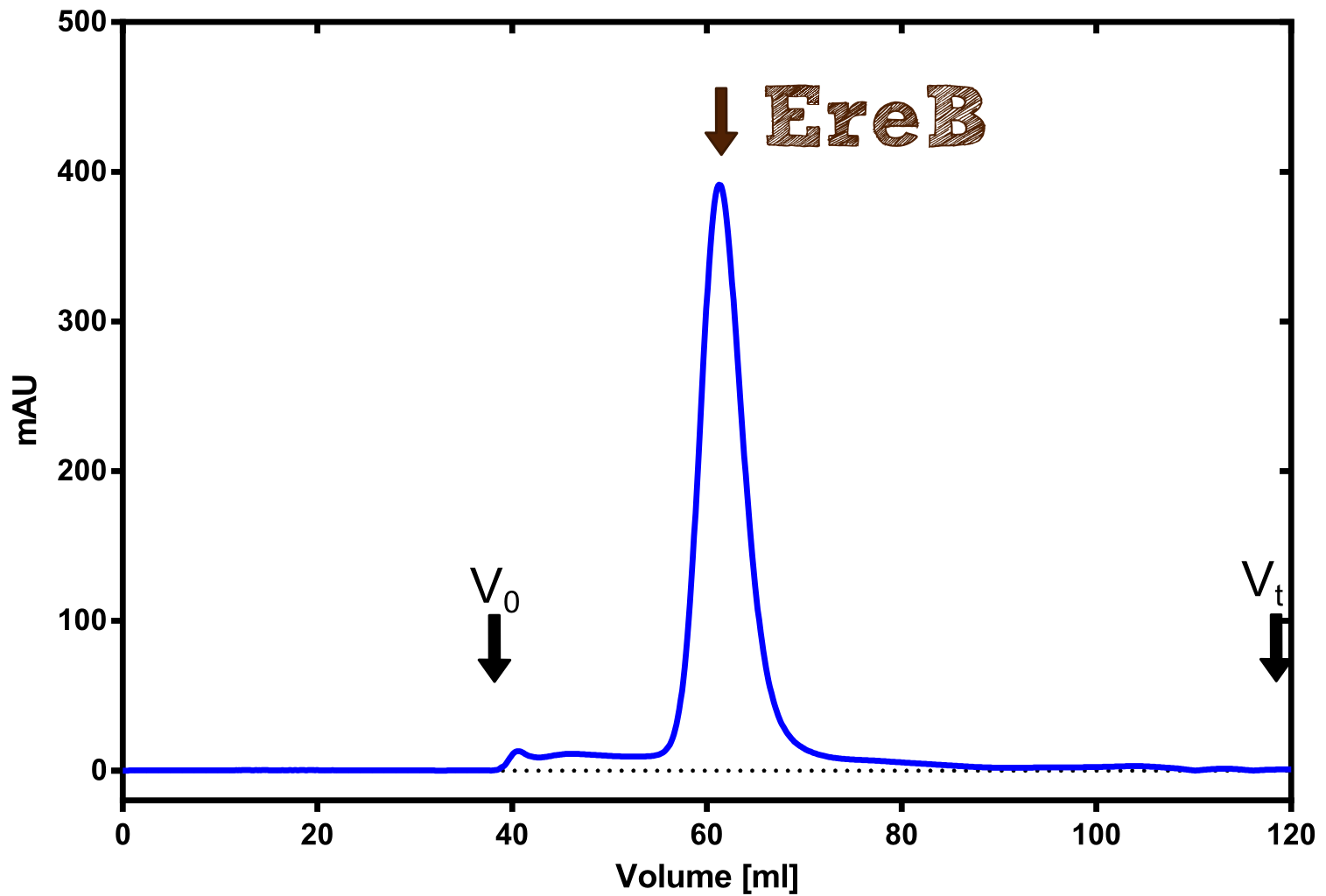
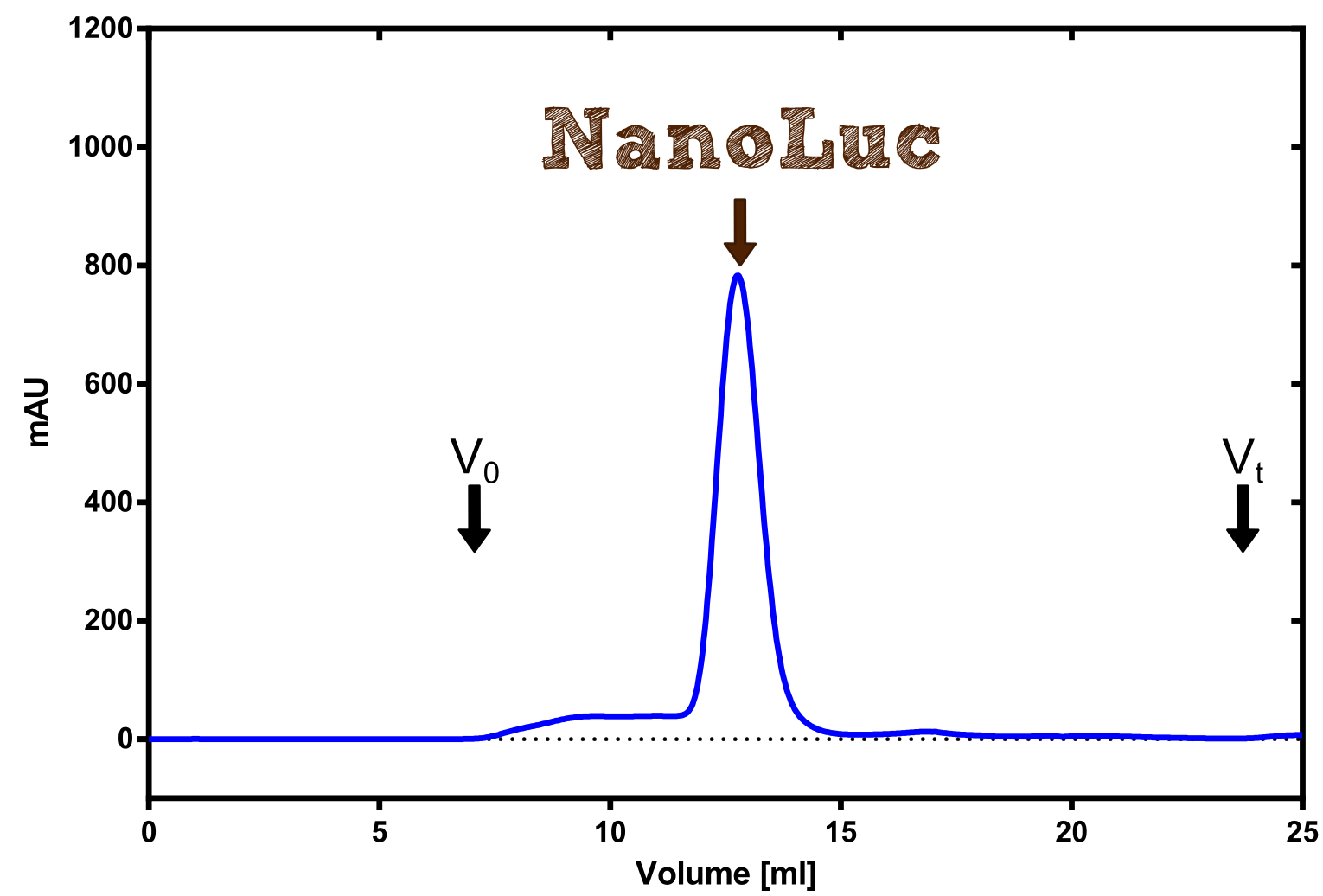
Production in E. coli and purification
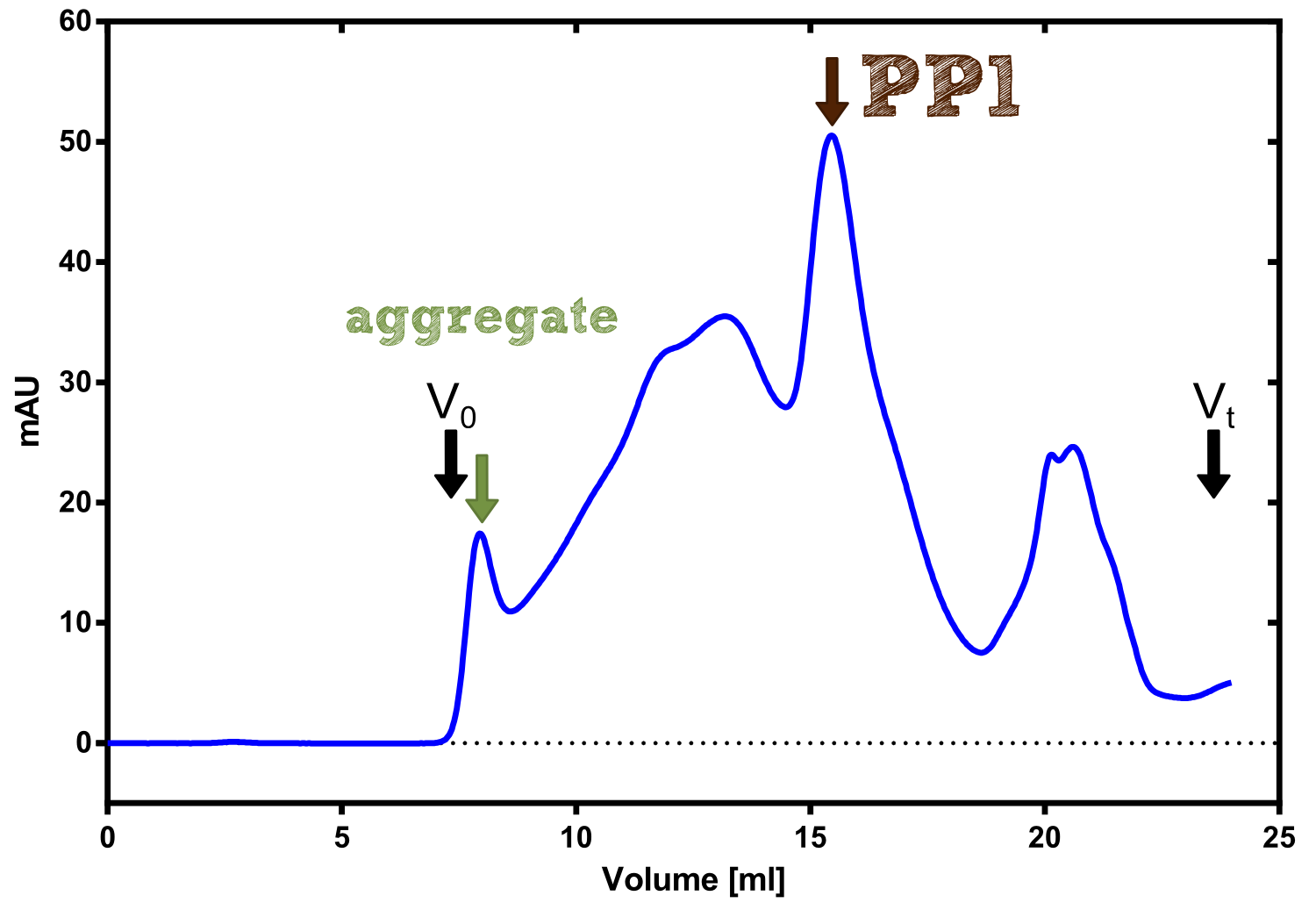
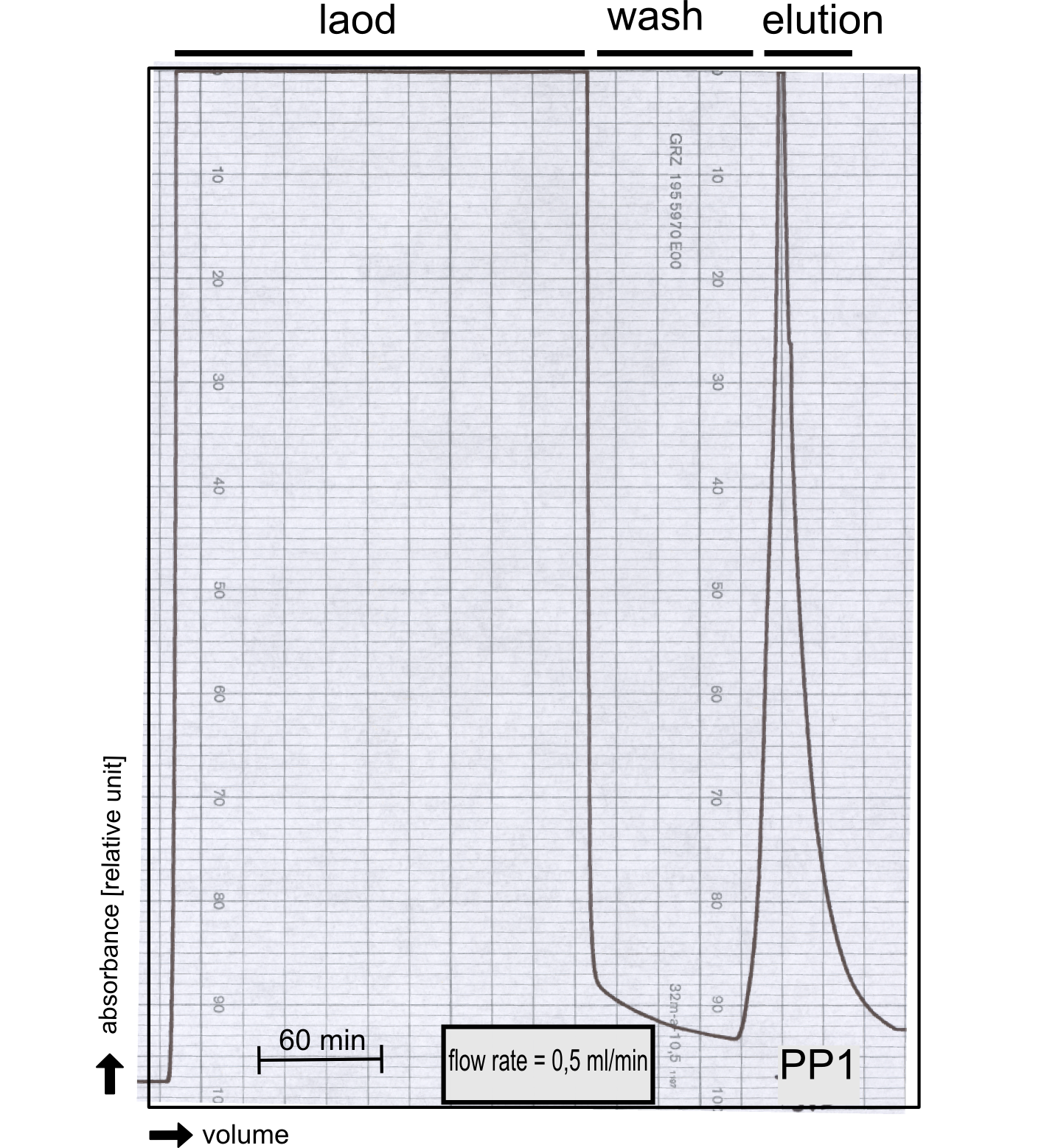
Therefore the NanoLuc was synthesized as a BioBrick in RFC[25] and was produced in E. coli using the pBad expression system with a C-terminal Strep-tag. After the production (2 l of LB-media for analytical and 12 l for preparative preparations) the cells were disrupted using sonification and the lysate was dialysed against 5 l of 1x SA-buffer. Afterwards the lysate was applied to a Streptavidin-Affinity (SA) column and was subsequently washed using SA-Buffer until a baseline was reached and the protein was then eluted using 5 mM of biotin (Attention: These are special columns which are not availible commercially. If you are using commercial colum material you have to use d-Desthiobiotin because usual biotin will elute your protein but you will not be able to regenerate the column after your chromatography). After the SA-chromatography the protein was concentrated using centrifugal concentration units (MWCO: 10 kDa). The concentrated protein was then applied on a Superdex S200/75 size exclusion chromatography.
Structure of the Nano Luciferase
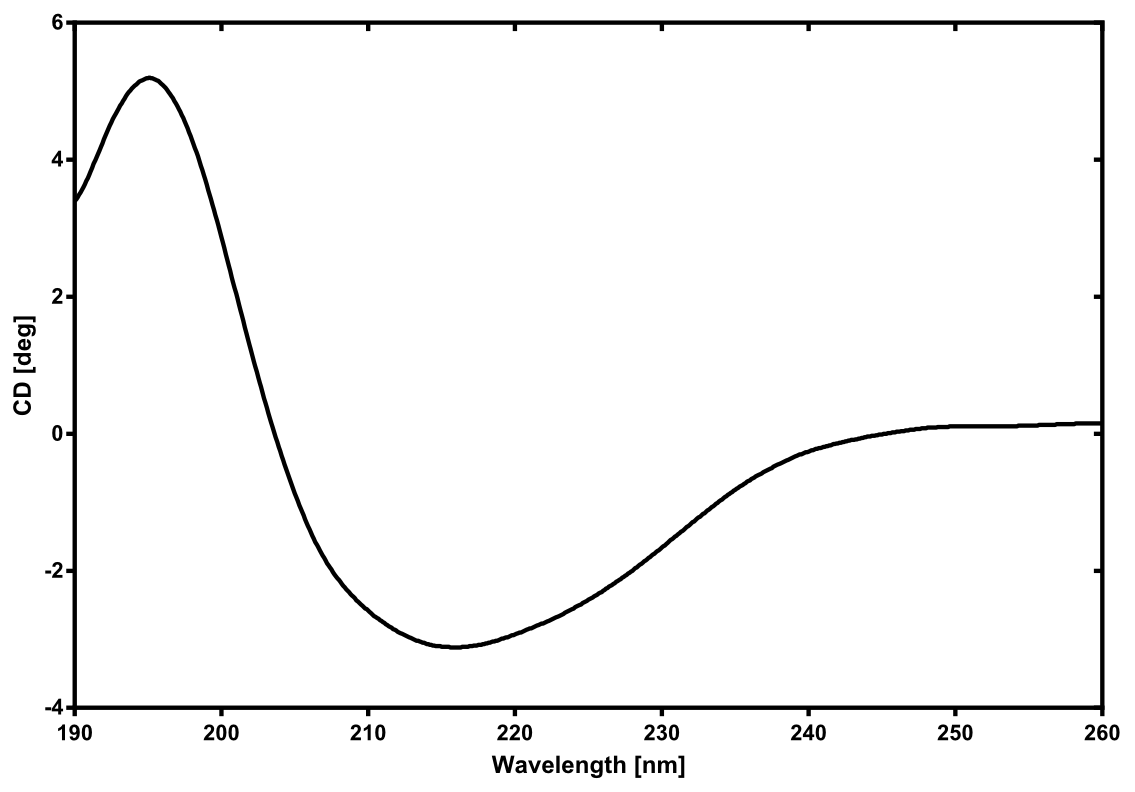
There is no structure availible for the NanoLuc in the [http://www.rcsb.org/pdb/home/home.do Protein Data Bank]. In our protein modelling we used homolgy search and identified the structure [http://www.rcsb.org/pdb/explore/explore.do?structureId=3PPT 3ppt_A] as the solved structure with the highest homology to the NanoLuc which has only 21% identify with a similarity of only 0.359. The result of the homology search is shown as annimated gif in Figure xx (please see our How To for an introduction). The protein was dialysed against 1x CD-buffer and subsequently a circular dichroism spectroscopy was tanken (learn about CD spectroscopy). The CD spectrum was used to predict the secondary structure content of the NanoLuc which could be determined to 35.1% helix, 27.6% b-strand, 18.5% turn and 18.8% random. As there is only a poor homology present, a detailed comparison of the determined and the predicted secondary structure is not possible. But it can be stated that both show a balanced content of different secondary structures and that the produced protein is present in an folded conformation. The mixed secondary structure content is also in consistance with the predicted secondary structure shown in the AutoAnnotator sequence window (click on show).
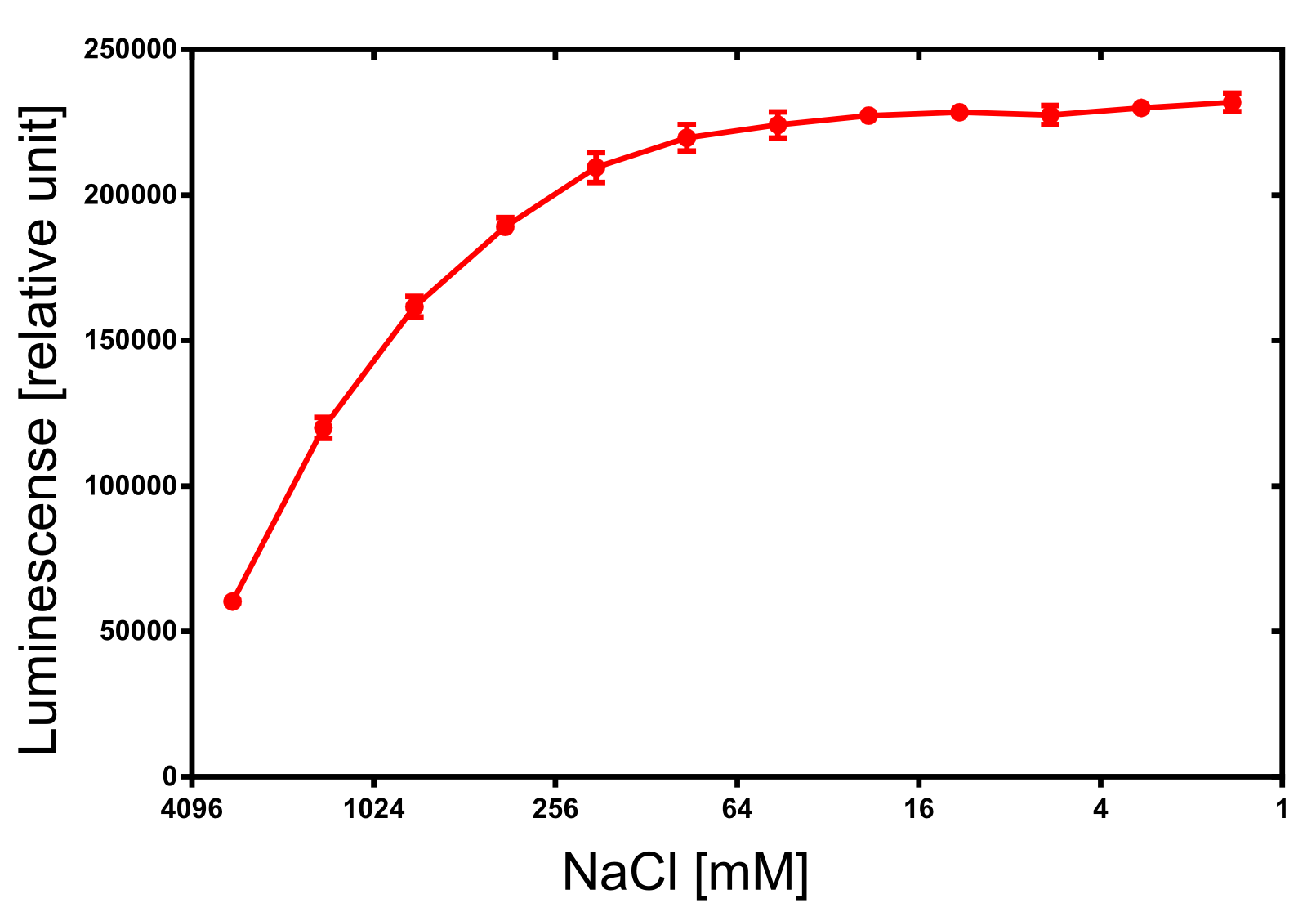
Activity determination of Luminescense
The activity of the produced NanoLuc was investigated by its luminescense. The luminescense
XylE
[...] description [...] reaction [...] production
| Protein data table for BioBrick BBa_ automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 25, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein) ATGGCCGGCAACAAAGGT ... GTGCTGACCACCGGT ORF from nucleotide position -8 to 924 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
Analytical präparation
[...] Characterization
DDT-Dehydrochlorinase
[...] description [...] reaction [...] production
| Protein data table for BioBrick BBa_ automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 10: (underlined part encodes the protein) ATGGACTTT ... TTCCTGAGCTAGTAG ORF from nucleotide position 1 to 627 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
Analytical präparation
[...] Characterization
PP1
SpyCatcher & SpyTag
[...] description [...] reaction [...] production
| Protein data table for BioBrick BBa_ automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 25, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein) ATGGCCGGCGTTGATACC ... GCTCATATTACCGGT ORF from nucleotide position -8 to 345 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
Analytical präparation
[...] Characterization
References:
http://www.ncbi.nlm.nih.gov/pubmed/6327079 Edens et al., 1984
- http://www.ncbi.nlm.nih.gov/pubmed/6327079 Edens et al., 1984 Edens, L., Bom, I., Ledeboer, A. M., Maat, J., Toonen, M. Y., Visser, C., and Verrips, C. T. (1984). Synthesis and processing of the plant protein thaumatin in yeast. Cell, 37(2):629–33.
- http://udel.edu/~gshriver/pdf/Pimenteletal1997.pdf Pmentel et al., 1997 Pimentel, D., Wilson, C., McCullum, C., Huang, R., Dwen, P., Flack, J. Tran, Q., Saltman, T., Cliff, T. (1997). Economic and environmental benefits of biodiversity. BioScience, Vol. 47, No. 11., pp. 747-757.
 "
"







AutoAnnotator:
Follow us:
Address:
iGEM Team TU-Munich
Emil-Erlenmeyer-Forum 5
85354 Freising, Germany
Email: igem@wzw.tum.de
Phone: +49 8161 71-4351