Team:Bielefeld-Germany/Biosafety/Biosafety System L
From 2013.igem.org
| Line 215: | Line 215: | ||
*Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|''Journal of Bacteriology 171: 1185 - 1187.'']. | *Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|''Journal of Bacteriology 171: 1185 - 1187.'']. | ||
| + | |||
| + | *Voss, Carsten Lindau, Dennis and Flaschel, Erwin (2006) Production of Recombinant RNase Ba and Its Application in Downstream Processing of Plasmid DNA for Pharmaceutical Use [http://onlinelibrary.wiley.com/doi/10.1021/bp050417e/pdf|''Biotechnology Progress 22: 737 - 744.'']. | ||
| + | |||
| + | *Walsh, Christopher (1989) Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. [http://www.jbc.org/content/264/5/2393.long|''Journal of biological chemistry 264: 2393 - 2396.''] | ||
| + | |||
| + | *Wickstrum, J.R., Santangelo, T.J., and Egan, S.M. (2005) Cyclic AMP receptor protein and RhaR synergistically activate transcription from the L-rhamnose-responsive rhaSR promoter in Escherichia coli. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251584/?report=reader|''Journal of Bacteriology 187: 6708 – 6719.'']. | ||
Revision as of 13:06, 4 October 2013
Biosafety System Lac of Growth
Overview
comming soon... (will only take a couple of hours)
Genetic Approach
Rhamnose promoter
lacI
Naturally the lac operon regulates the catabolism of the disaccharide lactose (4-O-(β-D-Galactopyranosyl)-D-glucopyranose) in E. coli. The operon contains the lactose promoter (plac) and the genes for the catabolism of the Lactose to Glucose and Galactose. Upstream of the lac operator exists the coding sequence for the repressor lacI under the control of a weak promoter (Jacob et al., 1961).
Compared to the catabolism of the sugars L-Rhamnose or L-arabinose Lactose, as a disaccharide has a higher energy content and is therefore used more preferable. This is a reason, why the basal transcription of this promoter is even more higher. The leakiness of the lac promoter is caused by the fact that the lacY need to be expressed for an efficient Lactose uptake, while in the arabinose system the uptake is regulated separate (Görke et al., 2008).
In our Safety-System the lacI (<bbpart>BBa_C0012</bbpart>) is used for the repression of the lactose promoter (plac).
Alanine Racemase
Terminator
Lactose promoter (plac)
Naturally the lac operon regulates the catabolism of the disaccharide lactose (4-O-(β-D-Galactopyranosyl)-D-glucopyranose) in E. coli. The operon consists of a CAP-binding site, the lac promoter, the lac operator and the genes lacZ, lacY and lacA downstream of the promoter. The transcription of the lactose promoter is regulated by the lacI gene, which is found upstream of the operon under the control of a weak promoter. In the absence of lactose the transcription of the genes behind the lactose promoter is blocked caused by the binding of the lacI pressor. While in the presence of Lactose the repressor is released from the operator and the genes can be transcripted. Typically the transcription is enhanced by a high intracellular level of cAMP (Busby, 1999).

The lactose promoter thereby regulates the transcription of the genes lacZ, lacY and lacA. The lacZ gene encodes for the ß-Galactosidase a enzyme, who breaks down the lactose to glucose and galactose. The ß-Galactosidase catalyses additional the degradation from Lactose to Allolactose. By binding on the lacI repressor, it changes his conformation an is not any more able to bind on the operator sequnece and to block the transcription. As only one enzyme is necessary to gain a substrate of the glycolysis is becomes clear, why the degradation of Lactose is more preferable compared to L-arabinose or L-rhamnose.
To realize a preference of lactose, the transcription of the lactose promoter is not repressed as that strong. This is caused by the fact that the lacY gene, coding for the integral membrane protein lactose permease, is necessary for the lactose uptake and has to be transcripted on a low level.
The last gene of the lac operon, lacA, encodes for a Transacetylase, who acetylizes glycosides that can not be metabolized. The acetylated glycosides are transported outside the cell to avoid the accumulation of lactose.
In our Biosafety-System the lac-promoter is used for the regulation of GFP or the Barnase. As the lac promoter shows a high basal transcription, its might not ideal for the regulation of a toxic gene product, but the the Biosafety-System Lac of growth is ideal for comparison with the other Systems to measure the level of basal transcription under repressed and unrepressed conditions. Besides we improved the leakiness of the lactose promoter by adding a second lacI-binding site 12 nt downstream of the excisting bining site. As this distance corresponds to about one whorl of the double helix, this should allow an additional lacI repressor to bind on the other site of the DNA and tighten the repression of the lactose promoter. Unfortunately the improvement of the so called double lac promoter could not be quantified, because lac of time (Lewis, 2005).
Barnase
Biosafety-System Lac of growth
Together with the Biosafety-Strain K-12 ∆alr ∆dadX the Biosafety-System Lac of growth takes advantage of this genes by combine them to a powerful device, which allows to control the bacterial cell division. The control of the bacterial growth is thereby active and passive possible. Active by inducing the lactose promoter with IPTG or classically with Lactose and passive by the induction of L-Rhamnose. The passive control makes it possible to control the bacterial cell division in an defined environment, like the MFC by adding continously L-Rhamnose to the media. As shown in the figure below, this leads to an expression of the essential Alanine-Racemase (alr) and the lacI repressor, so that the expression of the RNase Ba is repressed.
When the bacteria exit the defined environment of the MFC or L-Rhamnose is not added any more to the media, the expression of the Alanine-Racemase (alr) and the lacI repressor decreases, so that the expression of the toxic RNase Ba (Barnase) is not inhibted so strong any more. The cleavage of the intracellular RNA by the Barnase and ideally also the lack of synthesized D-alanine, caused by the repressed Alanine-Racemase inhibits the cell division and makes sure that the bacteria can only grow in the defined area.

Results
Characterization of the lactose promoter plac
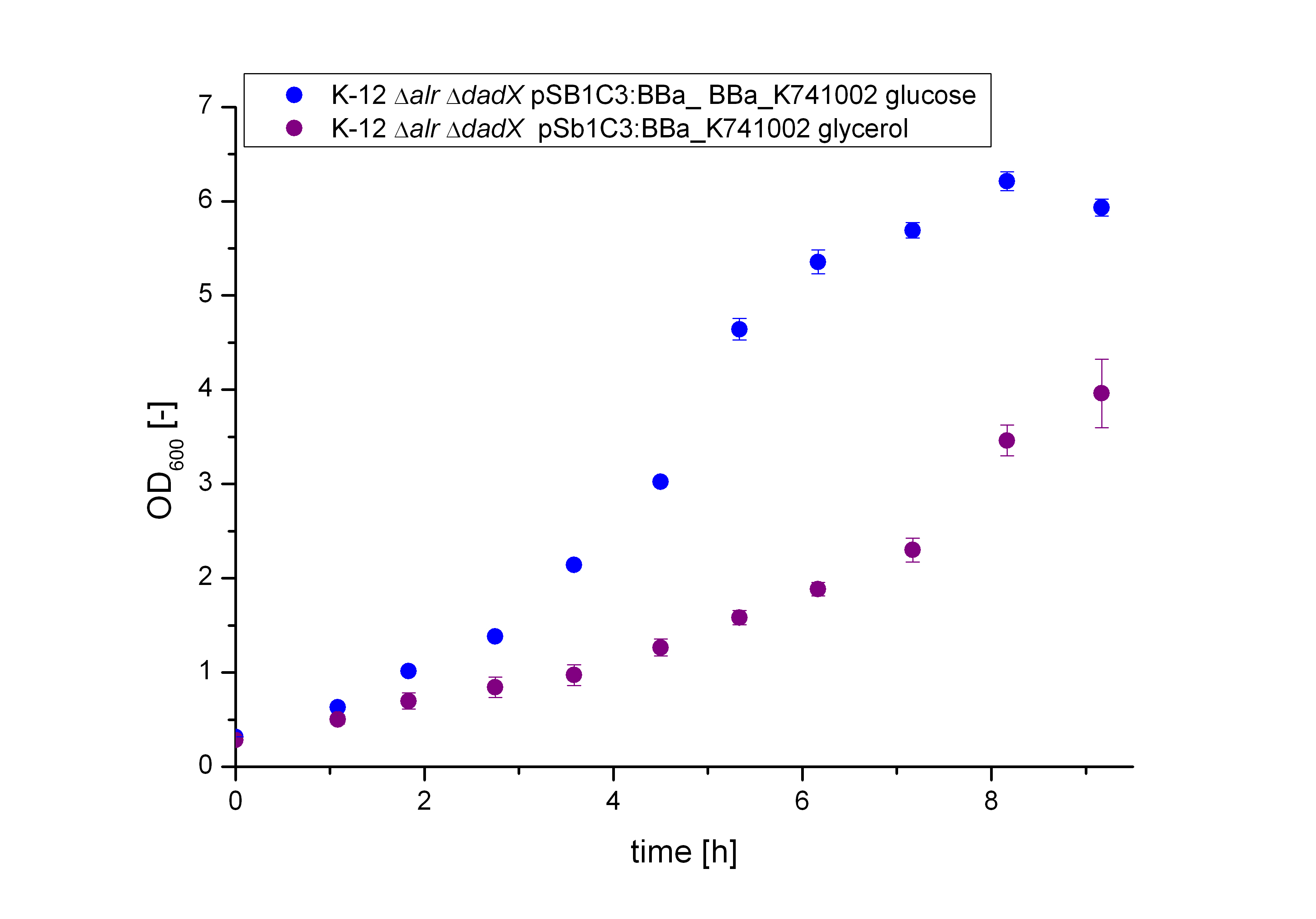
Characterization of the Biosafety-System Lac of growth
Conclusion of the Results
References
- Baldoma L and Aguilar J (1988) Metabolism of L-Fucose and L-Rhamnose in Escherichia coli: Aerobic-Anaerobic Regulation of L-Lactaldehyde Dissimilation [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210658/pdf/jbacter00179-0434.pdf|Journal of Bacteriology 170: 416 - 421.].
- Busby S, Ebright RH (1999) Transcription activation by catabolite activator protein (CAP).[http://ac.els-cdn.com/S0022283699931613/1-s2.0-S0022283699931613-main.pdf?_tid=a8f22b74-2cf4-11e3-9457-00000aab0f6c&acdnat=1380891687_f17fd24e5a3e15da96493226fdcaaa10|Journal of molecular biology 239: 199 - 213.].
- Carafa, Yves d'Aubenton Brody, Edward and Claude (1990) Thermest Prediction of Rho-independent Escherichia coli Transcription Terminators - A Statistical Analysis of their RNA Stem-Loop Structures [http://ac.els-cdn.com/S0022283699800059/1-s2.0-S0022283699800059-main.pdf?_tid=ede07e2a-2a92-11e3-b889-00000aab0f6c&acdnat=1380629809_2d1a59e395fc69c8608ab8b5aea842f7|Journal of molecular biology 3: 835 - 858].
- Görke, Boris and Stülke, Jörg (2008) Carbon catabolite repression in bacteria: many ways to make the most out of nutrients [http://www.nature.com/nrmicro/journal/v6/n8/full/nrmicro1932.html|Nature Reviews Microbiology 6: 613 - 624].
- Jacob, F Monod J (1961) Genetic regulatory mechanisms in the synthesis of proteins. [http://www.ncbi.nlm.nih.gov/pubmed/13718526|Journal of molecular biology 216: 318 - 356].
- Lewis, Mitchell (2005) The lac repressor [http://www.sciencedirect.com/science/article/pii/S1631069105000685|Comptes rendues biologies 328: 521 – 548].
- Mossakowska, Danuta E. Nyberg, Kerstin and Fersht, Alan R. (1989) Kinetic Characterization of the Recombinant Ribonuclease from Bacillus amyloliquefaciens (Barnase) and Investigation of Key Residues in Catalysis by Site-Directed Mutagenesis [http://pubs.acs.org/doi/pdf/10.1021/bi00435a033|Biochemistry 28: 3843 - 3850.].
- Paddon, C. J. Vasantha, N. and Hartley, R. W. (1989) Translation and Processing of Bacillus amyloliquefaciens Extracellular Rnase [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC209718/pdf/jbacter00168-0575.pdf|Journal of Bacteriology 171: 1185 - 1187.].
- Voss, Carsten Lindau, Dennis and Flaschel, Erwin (2006) Production of Recombinant RNase Ba and Its Application in Downstream Processing of Plasmid DNA for Pharmaceutical Use [http://onlinelibrary.wiley.com/doi/10.1021/bp050417e/pdf|Biotechnology Progress 22: 737 - 744.].
- Walsh, Christopher (1989) Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. [http://www.jbc.org/content/264/5/2393.long|Journal of biological chemistry 264: 2393 - 2396.]
- Wickstrum, J.R., Santangelo, T.J., and Egan, S.M. (2005) Cyclic AMP receptor protein and RhaR synergistically activate transcription from the L-rhamnose-responsive rhaSR promoter in Escherichia coli. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1251584/?report=reader|Journal of Bacteriology 187: 6708 – 6719.].
 "
"