Team:Bielefeld-Germany/Labjournal/July
From 2013.igem.org
(Difference between revisions)
| Line 80: | Line 80: | ||
*Glycerol dehydrogenase | *Glycerol dehydrogenase | ||
**Further work on GldA Biobrick is needed: Plasmid restriction analysis and colony PCR could not give sufficient results. | **Further work on GldA Biobrick is needed: Plasmid restriction analysis and colony PCR could not give sufficient results. | ||
| - | **Successful PCR (standard Phusion PCR) on the BBa_J04450 Biobrick plasmid using Forward Primer pSB1C3 and Reverse Primer pSB1C3 for later Gibson Assembly. | + | **Successful PCR (standard Phusion PCR) on the <bbpart>BBa_J04450</bbpart> Biobrick plasmid using Forward Primer pSB1C3 and Reverse Primer pSB1C3 for later Gibson Assembly. |
**Gibson Assembly with GldA PCR product and pSB1C3 PCR product. | **Gibson Assembly with GldA PCR product and pSB1C3 PCR product. | ||
**Screening of colonies from Gibson Assembly with colony PCR and Plasmid restriction analysis shows religated pSB1C3, as already seen while using NEB Assembly Kit. | **Screening of colonies from Gibson Assembly with colony PCR and Plasmid restriction analysis shows religated pSB1C3, as already seen while using NEB Assembly Kit. | ||
Revision as of 16:38, 28 September 2013
July
Milestones
9.Week
Organization
- We’ve presented us at the congress ‘Next generation of biotechnological processes 2020+’ in Berlin at 27. June 2013. There we have met an expert (Dr. Falk Harnisch) on the topic of MFC and we have decided to participate the German Synthetic Biology Day.
MFC
Mediators
Cytochromes
- Design and order of new Primers
- The mtrCAB fragment has two illegal PstI restriction sites at x xbp and yy bp, so we had to design new primers to remove them. We replaced one base in each restriction site, without affecting the coding triplett, by respective primer overlaps . We ended up with three different fragments, which will be ligated back together via Gibson Assembly.
- mtrCAB_Frag1_rev:
- mtrCAB_Frag2_fwd:
- mtrCAB_Frag2_rev:
- mtrCAB_Frag3_fwd:
Biosafety
Porines
10.Week
Organization
- We can present new sponsors of iGEM-Team Bielefeld: Stockmeier, Baxter, Promega, BIO.NRW and Applichem will support our team.
MFC
Mediators
- Glycerol dehydrogenase
- Further work on GldA Biobrick is needed: Plasmid restriction analysis and colony PCR could not give sufficient results.
- Successful PCR (standard Phusion PCR) on the <bbpart>BBa_J04450</bbpart> Biobrick plasmid using Forward Primer pSB1C3 and Reverse Primer pSB1C3 for later Gibson Assembly.
- Gibson Assembly with GldA PCR product and pSB1C3 PCR product.
- Screening of colonies from Gibson Assembly with colony PCR and Plasmid restriction analysis shows religated pSB1C3, as already seen while using NEB Assembly Kit.
Cytochromes
Biosafety
Porines
- We will try to use Gibson Assembly for cloning OprF into pSB1C3.
- Gibson Assembly with OprF PCR product and pSB1C3 PCR product. Screening of colonies from Gibson Assembly with colony PCR and Plasmid restriction analysis shows religated pSB1C3, as already seen while using NEB Assembly Kit.
11.Week
Organization
- We’ve presented us at the congress “BioNRW pHD Student Convention” in Düsseldorf at 13. July 2013 with a short presentation about iGEM and our project.
- We proudly present our [http://ekvv.uni-bielefeld.de/blog/uniaktuell/entry/mit_bakterien_batterie_strom_erzeugen first press release.] It was amazing to see how often our press release was picked up.
MFC
Mediators
- Glycerol dehydrogenase
- Repeating Gibson Assembly shows every time a religated pSB1C3, a Gibson overlap of Prefix and Suffix with 20 bp seems to be to short, furthermore Prefix and Suffix have too many homologous regions.
Cytochromes
Biosafety
Porines
- Repeating Gibson Assembly shows every time a religated pSB1C3, a Gibson overlap of Prefix and Suffix with 20 bp seems to be to short, furthermore Prefix and Suffix have too many homologous regions.
12.Week
Organization
- Dr. Falk Harnisch will have a presentation at CeBiTec colloquium as an expert for our team.
- Preparing experiments for the ‘Day of Synthetic Biology’. Our ideas are to have different experiments like DNA isolation from fruit and vegetables, pipetting of bright colors, chromatography with markers, a potato battery or microscopy.
- The TV station WDR would like to contribute with us on our topic
MFC
Mediators
- Glycerol dehydrogenase
- New Primerdesign for pSB1C3 with longer overlaps individually for each part in order to stop the problem of religated pSB1C3:
- Forward Primer pSB1C3 GldA (43 bp): TCCTGCAAGAGTGGGAATAATACTAGTAGCGGCCGCTGCAGTC
- Reverse Primer pSB1C3 GldA (43 bp): GATTGAATAATGCGGTCCATCTAGAAGCGGCCGCGAATTCCAG
Cytochromes
- Amplification of Fragment 1
- Size: 1800 bp
- Program: PhusionPCR
- Gradient: 54°C - 71°C over 8 steps
- Primer: mtrC_fwd & mtr_Frag1_rev
- Template: S. oneidensis PCR Template from genomic DNA
- Notes: Annealing temperature has no significant impact on the PCR
- Amplification of Fragment 2
- Size: 330 bp
- Program: PhusionPCR
- Primer: mtr_Frag2_fwd & mtr_Frag2_rev
- Template: S. oneidensis PCR Template from genomic DNA
- Notes:
- Amplification of Fragment 3
- Size: 3000 bp
- Program: PhusionPCR
- Gradient: 54°C - 71°C over 8 steps
- Primer: mtr_Frag3_fwd & mtrB_rev
- Template: S. oneidensis PCR Template from genomic DNA
- Notes: Annealing temperature has no significant impact on the PCR
- Amplification of ccmAH cluster
- Size:6311 bp
- Program: PhusionPCR
- Gradient: 51.5°C - 68°C over 8 steps
- Primer: ccmAH_fwd & ccmAH_rev
- Template: E. coli PCR Template from genomic DNA
- Notes: A high annealing temperature of 68°C or more increases yield as well as reduces unspecific bands and is therefore recommended.
- Gelextraction and cleanup of Fragment 1, 2, 3 and ccmAH
- Gelextraction and clean up with the AnalytikJena GelExtraction-Kit
- Elution buffer was preheated to 50°C
- We eluted each column twice, with 20 ul each and combined it afterwards
- Measurement of nucleid acid concentration via NanoDrop
- Fragment1: 4-2607-304: 10.5 ng/ul</p>
- Fragment2: 4-2707-003: 44.6 ng/ul</p>
- Fragment2: 4-2707-004: 13.5 ng/ul</p>
- Fragment3: 4-2607-301: 11.1 ng/ul</p>
- Fragment3: 4-2607-302: 8.9 ng/ul</p>
- Fragment3: 4-2607-303: 9.2 ng/ul</p>
- ccmAH: 4-2807-302: 11.0 ng/ul</p>
- ccmAH: 4-2807-303: 11.5 ng/ul</p>
Cytochromes
Biosafety
Porines
- New Primerdesign for pSB1C3 with longer overlaps individually for OprF part in order to stop the problem of religated pSB1C3:
- Forward Primer pSB1C3 OprF (43 bp): TTGAAGCCCAAGCTAAGTAATACTAGTAGCGGCCGCTGCAGTC
- Reverse Primer pSB1C3 OprF (43 bp): AAGGTGTTTTTCAGTTTCATCTAGAAGCGGCCGCGAATTCCAG
13.Week
Organization
- Spontaneous visit of Radio Bielefeld in our laboratory. In addition to a radio interview a [http://www.radiobielefeld.de/programm/bei-uns-im-programm/studenten-entwickeln-biobatterie.html short video clip] was filmed.
MFC
Mediators
Cytochromes
Biosafety
- Amplification of different fragments:
- Recipe:
- PhusionBuffer: 5,5µL
- dNTPs: 1µL
- Primer1: 1µL
- Primer2: 1µL
- Template: 1,5 µL
- DMSO: 0,5 µL
- Phusion: 0,5 µL
- Dest. H2O: 39 µL
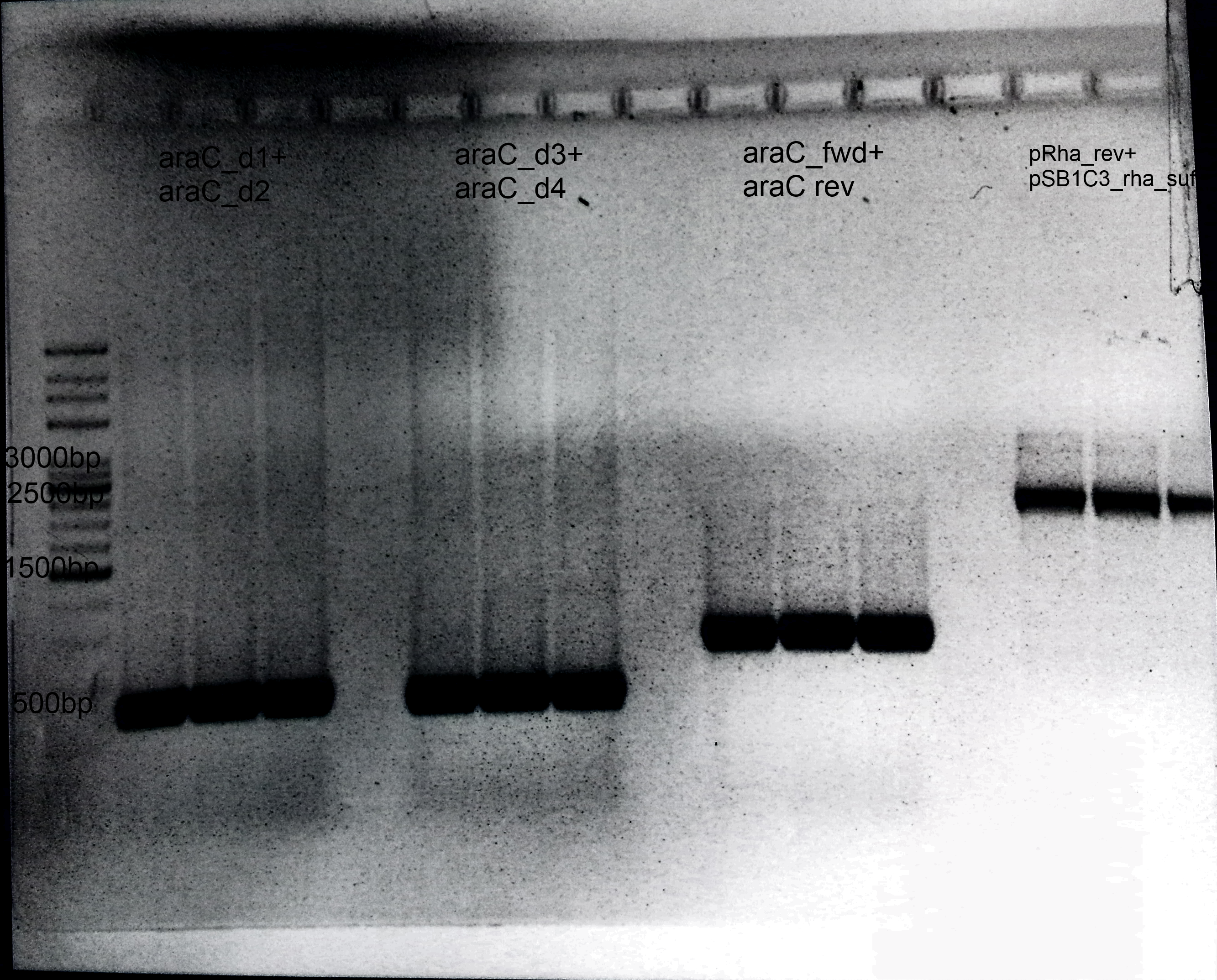
- 1.AraC:
- Primer: araC_d1+ araC_d2
- Annealing: 62°C
- Extension: 1min
- Template: E.coli Genome
- Size: 600 bp
- 2.Deletion of araC:
- Primer: araC_d3+ araC_d4
- Annealing= 62°C
- Extension: 1min
- Template: E.coli Genome
- Size: 600 bp
- 3.Amplification of araC into our Biosafety system:
- Primer: araC_fwd+ araC rev
- Annealing= 62°C
- Extension: 1min
- Template: BBa_I13458
- Size: 900 bp
- 4.Präparation of pSB1C3:
- Primer: pRha_rev+ pSB1C3_rha_suf
- Annealing: 62°C
- Extension: 1.30 min
- Template: BBa_I13541
- Size: 3 kb
- 5.Präparation of pSB1C3:
- Primer: pSB1C3_alr_fwd+ pSB1C3_alr_rev
- Annealing: 62°C
- Extension: 1 min
- Template: E.coli Genome
- Size: 1kb
- 6.Präparation of pSB1C3:
- Primer: pSB1C3_plac_alr+ pSB1C3_alr_rev
- Annealing: 62°C
- Extension: 1min
- Template: E.coli Genome
- Size: 1kb
- 7.Präparation of pSB1C3:
- Primer: pSB1C3_alr_pre+ pSB1C3_alr_suf
- Annealing: 62°C
- Extension: 1.30 min
- Template: pSB1C3
- Size: 2kb
- 8.Präparation of pSB1C3:
- Primer: pSB1C3_plac_pre+ pSB1C3_alr_suf
- Annealing: 62°C
- Extension: 1.30min
- Template: pSB1C3
- Size: 2kb
- Analysis by agarosegel
- Result:failed
- New PCR:
- Program: 06 phy_cyt mit 62°C Annealing
- 10µL 5xHF-Buffer
- 4µL dNTPs
- 0,5µL Primer1
- 0,5µL Primer2
- 1µL Template
- 1,5µL DMSO
- 1µL Enzyme
- 31,5µL dest. H2O
- Analysis by agarosegel:
- PCR-Purifikation:
- BS1a (9-38-451) 113,6ng/µL
- BS1b (9-38-452) 75,1 ng/µL
- BS2a (9-38-453) 85,5 ng/µL
- BS3a (9-38-454) 107,6 ng/µL
- BS4a (9-38-455) 14,4 ng/µL
- BS5a (9-38-456) 44,5 ng/µL
- BS5b (9-38-457) 49,0 ng/µL
- BS6a (9-38-458) 63,6 ng/µL
- BS7a (9-38-459) 6,9 ng/µL
- BS8a (9-38-460) 8,6 ng/µL
- BS1a (9-38-451) 113,6ng/µL
Porines
 "
"