Team:Bielefeld-Germany/Labjournal/May
From 2013.igem.org
(Difference between revisions)
| Line 28: | Line 28: | ||
===Milestones=== | ===Milestones=== | ||
| + | *Starting lab work with the first successful PCR: GldA with Pre- and Suffix overlaps could be amplified from ''Escherichia coli'' genome. | ||
<br><br><br><br> | <br><br><br><br> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| Line 58: | Line 53: | ||
===2.Week=== | ===2.Week=== | ||
| + | |||
| + | |||
====Organization==== | ====Organization==== | ||
*Distribution of our team in various subgroups for best work efficiency. | *Distribution of our team in various subgroups for best work efficiency. | ||
*We’ve organized a working group barbecue to thank the AG of Dr. Jörn Kalinowski in the CeBiTec for the appropriation of space and support in the laboratory. | *We’ve organized a working group barbecue to thank the AG of Dr. Jörn Kalinowski in the CeBiTec for the appropriation of space and support in the laboratory. | ||
| + | |||
====MFC==== | ====MFC==== | ||
| Line 69: | Line 67: | ||
===3.Week=== | ===3.Week=== | ||
| + | |||
====Organization==== | ====Organization==== | ||
*Planning the trip to the European jamboree in Lyon from 11.-13. October 2013. | *Planning the trip to the European jamboree in Lyon from 11.-13. October 2013. | ||
| + | |||
====MFC==== | ====MFC==== | ||
| Line 82: | Line 82: | ||
**Successful PCR on the GldA gene of ''Escherichia coli'' KRX strain. | **Successful PCR on the GldA gene of ''Escherichia coli'' KRX strain. | ||
| - | |||
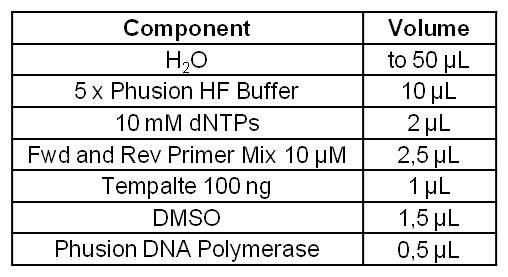
| + | [[Image:IGEM_Bielefeld_Standard_Phusion_PCR_Master_MixLRO.jpg|300px|thumb|left| <p align="justify">'''Table 1: Standard Phusion PCR Master Mix. '''</p>]] | ||
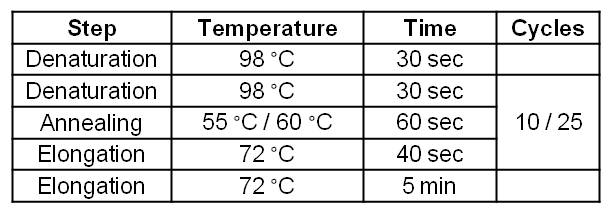
[[Image:IGEM_Bielefeld_Standard_Phu_PCR_GldA_OprF.jpg|300px|thumb|center| <p align="justify">'''Table 2: Two step standard Phusion PCR programm for GldA amplification. '''</p>]] | [[Image:IGEM_Bielefeld_Standard_Phu_PCR_GldA_OprF.jpg|300px|thumb|center| <p align="justify">'''Table 2: Two step standard Phusion PCR programm for GldA amplification. '''</p>]] | ||
| + | |||
**GldA PCR product was isolated by Agarose gel electrophorese and [[Team:Bielefeld-Germany/Labjournal/Molecular#Used Kits| purificated]]. | **GldA PCR product was isolated by Agarose gel electrophorese and [[Team:Bielefeld-Germany/Labjournal/Molecular#Used Kits| purificated]]. | ||
**Bands are at expected size of 1050 bp. | **Bands are at expected size of 1050 bp. | ||
| + | |||
[[Image:IGEM_Bielefeld_GldA_PCR_Gel.jpg|200px|thumb|left| <p align="justify">'''Figure 1: Agarosegel from PCR on the GldA gene of ''Escherichia coli'' KRX strain with forward and reverse primer GldA. For Ladder we used [http://www.thermoscientificbio.com/nucleic-acid-electrophoresis/generuler-1-kb-dna-ladder-ready-to-use-250-to-10000-bp GeneRuler™ 1 kb DNA Ladder fromThermo Scientific]. '''</p>]] | [[Image:IGEM_Bielefeld_GldA_PCR_Gel.jpg|200px|thumb|left| <p align="justify">'''Figure 1: Agarosegel from PCR on the GldA gene of ''Escherichia coli'' KRX strain with forward and reverse primer GldA. For Ladder we used [http://www.thermoscientificbio.com/nucleic-acid-electrophoresis/generuler-1-kb-dna-ladder-ready-to-use-250-to-10000-bp GeneRuler™ 1 kb DNA Ladder fromThermo Scientific]. '''</p>]] | ||
| Line 95: | Line 97: | ||
<br><br><br><br> | <br><br><br><br> | ||
===4.Week=== | ===4.Week=== | ||
| - | |||
| + | |||
| + | ====Organization==== | ||
*We will also take part at the congress "Next generation of biotechnological processes 2020+" in Berlin at 27. June 2013. | *We will also take part at the congress "Next generation of biotechnological processes 2020+" in Berlin at 27. June 2013. | ||
*Having a second presentation at Merck KGaA head office in Darmstadt for explaining our project in detail. We are happy that Merck will be our next supporter in 2013. | *Having a second presentation at Merck KGaA head office in Darmstadt for explaining our project in detail. We are happy that Merck will be our next supporter in 2013. | ||
| + | |||
====MFC==== | ====MFC==== | ||
Revision as of 22:58, 30 September 2013
May
Milestones
- Starting lab work with the first successful PCR: GldA with Pre- and Suffix overlaps could be amplified from Escherichia coli genome.
Week 1
Organization
- We were already able to find sponsors for our team: IIT BIEKUBA, IIT Biotech, Evonik, PlasmidFactory and FisherScientific will support us.
MFC
Mediators
- Glycerol dehydrogenase
- Primerdesign for isolation of GldA gene from Escherichia coli KRX strain, with overlaps for Biobrick Prefix and Suffix:
- Forward Primer GldA (43 bp): GAATTCGCGGCCGCTTCTAGATGGACCGCATTATTCAATCACC
- Reverse Primer GldA (45 bp): CTGCAGCGGCCGCTACTAGTATTATTCCCACTCTTGCAGGAAACG
2.Week
Organization
- Distribution of our team in various subgroups for best work efficiency.
- We’ve organized a working group barbecue to thank the AG of Dr. Jörn Kalinowski in the CeBiTec for the appropriation of space and support in the laboratory.
MFC
3.Week
Organization
- Planning the trip to the European jamboree in Lyon from 11.-13. October 2013.
MFC
Mediators
- Glycerol dehydrogenase
- Starting first cultivation of Escherichia coli KRX strain for complete genome isolation.
- Successful genome isolation of Escherichia coli KRX.
- Successful PCR on the GldA gene of Escherichia coli KRX strain.
- GldA PCR product was isolated by Agarose gel electrophorese and purificated.
- Bands are at expected size of 1050 bp.

Figure 1: Agarosegel from PCR on the GldA gene of Escherichia coli KRX strain with forward and reverse primer GldA. For Ladder we used GeneRuler™ 1 kb DNA Ladder fromThermo Scientific.
4.Week
Organization
- We will also take part at the congress "Next generation of biotechnological processes 2020+" in Berlin at 27. June 2013.
- Having a second presentation at Merck KGaA head office in Darmstadt for explaining our project in detail. We are happy that Merck will be our next supporter in 2013.
MFC
 "
"