Team:Bielefeld-Germany/Labjournal/September
From 2013.igem.org
| Line 34: | Line 34: | ||
| + | ===Week 18=== | ||
| + | ====Organization==== | ||
| - | |||
| - | + | *This week, we supported the [[Team:Bielefeld-Germany/Human Practice/Student Academy|CeBiTec student academy]] with supervising the ‘Synthetic Biology’ experiment. In addition we had a presentation about iGEM and the Bielefeld iGEM projects. | |
| - | *This week, we supported the CeBiTec student academy with supervising the ‘Synthetic Biology’ experiment. In addition we had a presentation about iGEM and the Bielefeld iGEM projects. | + | |
*Our team assigned the Track Food & Energy in this year, with the project title ‘Ecolectricity – currently available’. Project abstract, safety forms and team roster were added. | *Our team assigned the Track Food & Energy in this year, with the project title ‘Ecolectricity – currently available’. Project abstract, safety forms and team roster were added. | ||
| + | |||
====MFC==== | ====MFC==== | ||
| Line 47: | Line 48: | ||
====Mediators==== | ====Mediators==== | ||
| + | |||
| + | |||
*Glycerol dehydrogenase | *Glycerol dehydrogenase | ||
| - | **Repeating NADH-Assay with three times PBS buffer washing step shows good results with better NADH production for'' E. coli'' KRX with pSB1C3 <bbpart>BBa_K1172203</bbpart> and <bbpart>BBa_K1172204</bbpart> compared with ''E. coli'' KRX wildtyp. GldA with T7 promotor (<bbpart>BBa_K1172203</bbpart>) shows 0,5 μM NADH overproduction and GldA with Lac promotor (<bbpart>BBa_K1172204</bbpart>) shows 0,3 μM NADH overproduction. The data refer to a culture with OD600 = 3.0. | + | **Repeating [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]] with three times PBS buffer washing step shows good results with better NADH production for'' E. coli'' KRX with pSB1C3 <bbpart>BBa_K1172203</bbpart> and <bbpart>BBa_K1172204</bbpart> compared with ''E. coli'' KRX wildtyp. GldA with T7 promotor (<bbpart>BBa_K1172203</bbpart>) shows 0,5 μM NADH overproduction and GldA with Lac promotor (<bbpart>BBa_K1172204</bbpart>) shows 0,3 μM NADH overproduction. The data refer to a culture with OD600 = 3.0. |
| - | [[Image:IGEM_Bielefeld_Table5_Results_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 1: Results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] | + | [[Image:IGEM_Bielefeld_Table5_Results_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 1: Results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] |
| - | [[Image:IGEM_Bielefeld_Figure8_NADHAssay.jpg|300px|thumb|center|<p align="justify"> '''Figure 1: Column Chart with results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] | + | [[Image:IGEM_Bielefeld_Figure8_NADHAssay.jpg|300px|thumb|center|<p align="justify"> '''Figure 1: Column Chart with results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] |
| - | **Second successful optimized NADH-Assay with ''Escherichia coli'' KRX Wildtyp strain and ''E. coli'' KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart>, <bbpart>BBa_K1172204</bbpart>, and <bbpart>BBa_K1172205</bbpart>. | + | **Second successful optimized [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]] with ''Escherichia coli'' KRX Wildtyp strain and ''E. coli'' KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart>, <bbpart>BBa_K1172204</bbpart>, and <bbpart>BBa_K1172205</bbpart>. |
**GldA with T7 promotor (<bbpart>BBa_K1172203</bbpart>) shows 1,7 μM NADH overproduction and GldA with Lac promotor (<bbpart>BBa_K1172204</bbpart>) shows 0,2 μM NADH overproduction. The GldA expression by the Anderson promotor (<bbpart>BBa_K1172205</bbpart>) is too weak for an efficient NADH overproduction. The data refer to a culture with OD600 = 5.0. | **GldA with T7 promotor (<bbpart>BBa_K1172203</bbpart>) shows 1,7 μM NADH overproduction and GldA with Lac promotor (<bbpart>BBa_K1172204</bbpart>) shows 0,2 μM NADH overproduction. The GldA expression by the Anderson promotor (<bbpart>BBa_K1172205</bbpart>) is too weak for an efficient NADH overproduction. The data refer to a culture with OD600 = 5.0. | ||
| - | [[Image:IGEM_Bielefeld_Table6_Results_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 2: Results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172205</bbpart>, <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] | + | [[Image:IGEM_Bielefeld_Table6_Results_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 2: Results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172205</bbpart>, <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] |
| - | [[Image:IGEM_Bielefeld_Figure9_NADH.jpg|300px|thumb|center|<p align="justify"> '''Figure 2: Column Chart with results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172205</bbpart>, <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] | + | [[Image:IGEM_Bielefeld_Figure9_NADH.jpg|300px|thumb|center|<p align="justify"> '''Figure 2: Column Chart with results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172205</bbpart>, <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] |
| Line 392: | Line 395: | ||
====Mediators==== | ====Mediators==== | ||
*Glycerol dehydrogenase | *Glycerol dehydrogenase | ||
| - | **Glycerol dependent NADH-Assay with ''Escherichia coli'' KRX Wildtyp strain and ''E. coli'' KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart> by testing supplementation of different glycerol concentrations to the LB-medium. | + | **Glycerol dependent [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]] with ''Escherichia coli'' KRX Wildtyp strain and ''E. coli'' KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart> by testing supplementation of different glycerol concentrations to the LB-medium. |
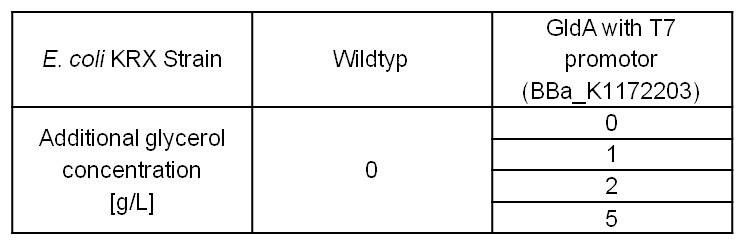
| - | [[Image:IGEM_Bielefeld_Table7_ExpDesign_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 4: Experimental design of the glycerol dependent NADH-Assay. Supplementation of different amounts of glycerol to LB-medium should show the effect of glycerol on NADH overproduction. '''</p>]] | + | [[Image:IGEM_Bielefeld_Table7_ExpDesign_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 4: Experimental design of the glycerol dependent [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Supplementation of different amounts of glycerol to LB-medium should show the effect of glycerol on NADH overproduction. '''</p>]] |
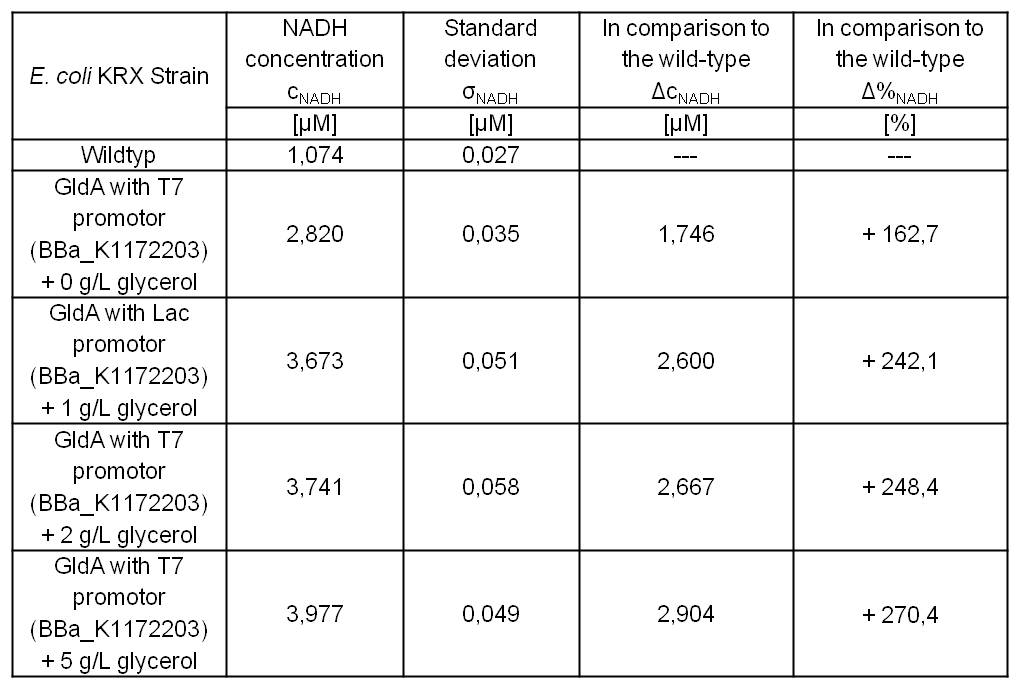
| - | [[Image:IGEM_Bielefeld_Table8_Results_NADH.jpg|300px|thumb|center|<p align="justify"> '''Table 5: Results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> supplemented with different amounts of glycerol to LB-medium. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] | + | [[Image:IGEM_Bielefeld_Table8_Results_NADH.jpg|300px|thumb|center|<p align="justify"> '''Table 5: Results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> supplemented with different amounts of glycerol to LB-medium. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] |
| - | **NADH-Assay shows a quiet good NADH overproduction for <bbpart>BBa_K1172203</bbpart> with increasing glycerol concentration. | + | **[[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]] shows a quiet good NADH overproduction for <bbpart>BBa_K1172203</bbpart> with increasing glycerol concentration. |
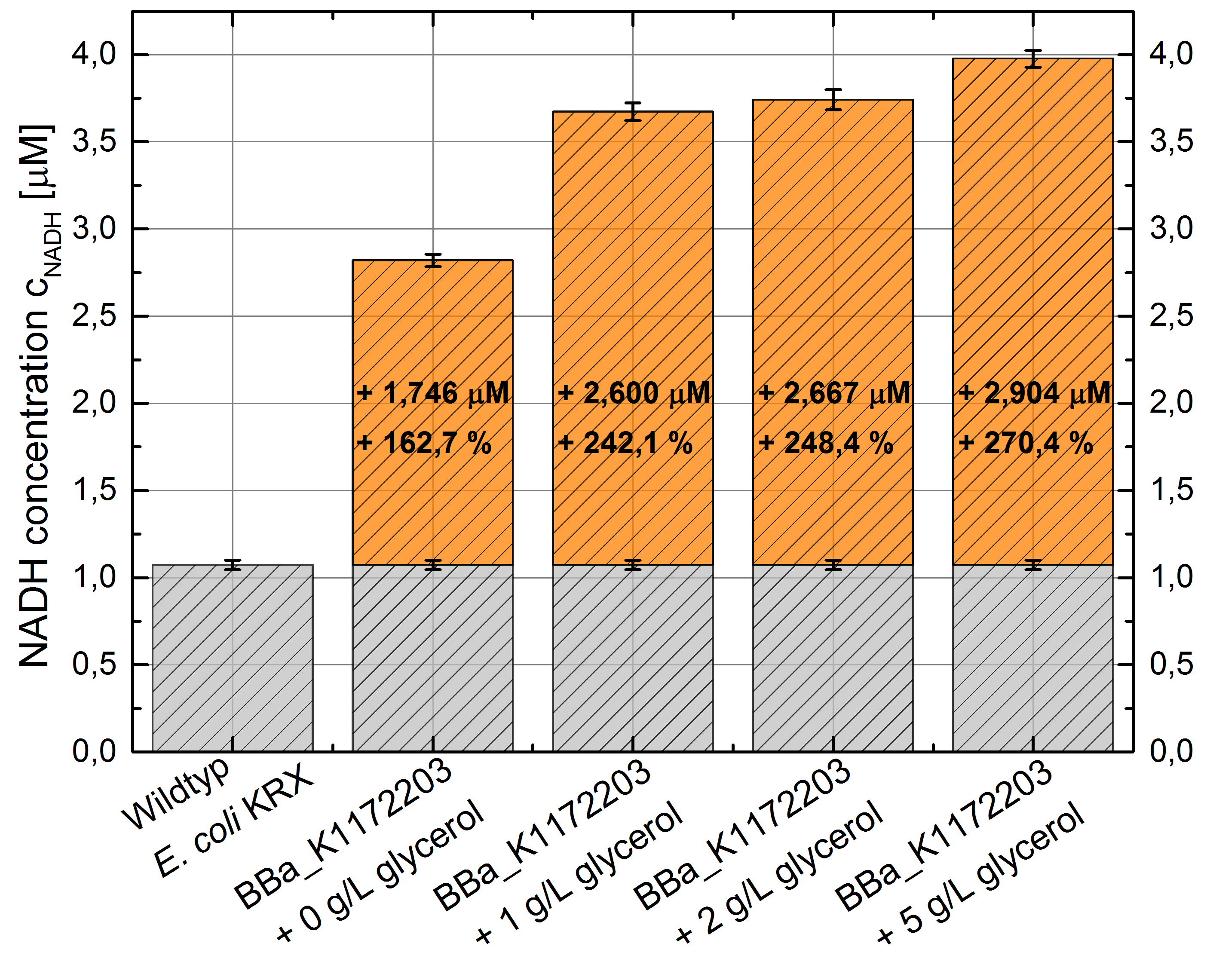
| - | [[Image:IGEM_Bielefeld_Figure10_Column_NADH.jpg|300px|thumb|left|<p align="justify"> '''Figure 8: Column Chart with results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> supplemented with different amounts of glycerol to LB-medium. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] | + | [[Image:IGEM_Bielefeld_Figure10_Column_NADH.jpg|300px|thumb|left|<p align="justify"> '''Figure 8: Column Chart with results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> supplemented with different amounts of glycerol to LB-medium. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown. '''</p>]] |
====Cytochromes==== | ====Cytochromes==== | ||
| Line 590: | Line 593: | ||
====Mediators==== | ====Mediators==== | ||
*Glycerol dehydrogenase | *Glycerol dehydrogenase | ||
| - | **Glycerol dependent NADH-Assay with ''Escherichia coli'' KRX Wildtyp strain and ''E. coli'' KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart> by testing different glycerol concentrations of M9-medium. | + | **Glycerol dependent [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]] with ''Escherichia coli'' KRX Wildtyp strain and ''E. coli'' KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart> by testing different glycerol concentrations of M9-medium. |
| - | [[Image:IGEM_Bielefeld_Table9_ExpDesign_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 6: Experimental design of the glycerol dependent NADH-Assay. Different concentrations of glycerol in M9-medium should show the effect of glycerol on NADH overproduction. '''</p>]] | + | [[Image:IGEM_Bielefeld_Table9_ExpDesign_NADH.jpg|300px|thumb|left|<p align="justify"> '''Table 6: Experimental design of the glycerol dependent [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Different concentrations of glycerol in M9-medium should show the effect of glycerol on NADH overproduction. '''</p>]] |
| - | [[Image:IGEM_Bielefeld_Table10_Results_NADH.jpg|300px|thumb|center|<p align="justify"> '''Table 7: Results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> grown in M9-medium with different concentrations of glycerol. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown for cell disruption and supernatant fraction. '''</p>]] | + | [[Image:IGEM_Bielefeld_Table10_Results_NADH.jpg|300px|thumb|center|<p align="justify"> '''Table 7: Results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> grown in M9-medium with different concentrations of glycerol. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown for cell disruption and supernatant fraction. '''</p>]] |
| - | **NADH-Assay shows an increasing intra- and extracellular NADH concentration for <bbpart>BBa_K1172203</bbpart> with increasing glycerol concentration in M9-medium. | + | **[[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]] shows an increasing intra- and extracellular NADH concentration for <bbpart>BBa_K1172203</bbpart> with increasing glycerol concentration in M9-medium. |
| - | [[Image:IGEM_Bielefeld_Figure11_Column_NADHAssay.jpg|300px|thumb|left|<p align="justify"> '''Figure 9: Column Chart with results of the NADH-Assay. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> grown in M9-medium with different concentrations of glycerol. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown for cell disruption (Intracellular) and supernatant (Extracellular) fraction. '''</p>]] | + | [[Image:IGEM_Bielefeld_Figure11_Column_NADHAssay.jpg|300px|thumb|left|<p align="justify"> '''Figure 9: Column Chart with results of the [[Team:Bielefeld-Germany/Labjournal/Molecular#NADH-Assay|NADH-Assay]]. Comparison between ''Escherichia coli'' KRX Wildtyp and ''Escherichia coli'' KRX with <bbpart>BBa_K1172203</bbpart> grown in M9-medium with different concentrations of glycerol. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown for cell disruption (Intracellular) and supernatant (Extracellular) fraction. '''</p>]] |
====Cytochromes==== | ====Cytochromes==== | ||
Revision as of 23:39, 2 October 2013
September
Milestones
Week 18
Organization
- This week, we supported the CeBiTec student academy with supervising the ‘Synthetic Biology’ experiment. In addition we had a presentation about iGEM and the Bielefeld iGEM projects.
- Our team assigned the Track Food & Energy in this year, with the project title ‘Ecolectricity – currently available’. Project abstract, safety forms and team roster were added.
MFC
Mediators
- Glycerol dehydrogenase
- Repeating NADH-Assay with three times PBS buffer washing step shows good results with better NADH production for E. coli KRX with pSB1C3 <bbpart>BBa_K1172203</bbpart> and <bbpart>BBa_K1172204</bbpart> compared with E. coli KRX wildtyp. GldA with T7 promotor (<bbpart>BBa_K1172203</bbpart>) shows 0,5 μM NADH overproduction and GldA with Lac promotor (<bbpart>BBa_K1172204</bbpart>) shows 0,3 μM NADH overproduction. The data refer to a culture with OD600 = 3.0.
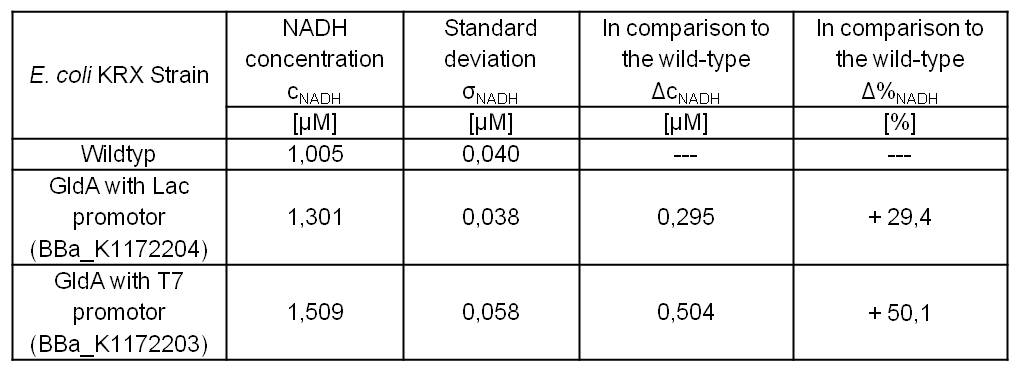
Table 1: Results of the NADH-Assay. Comparison between Escherichia coli KRX Wildtyp and Escherichia coli KRX with <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown.
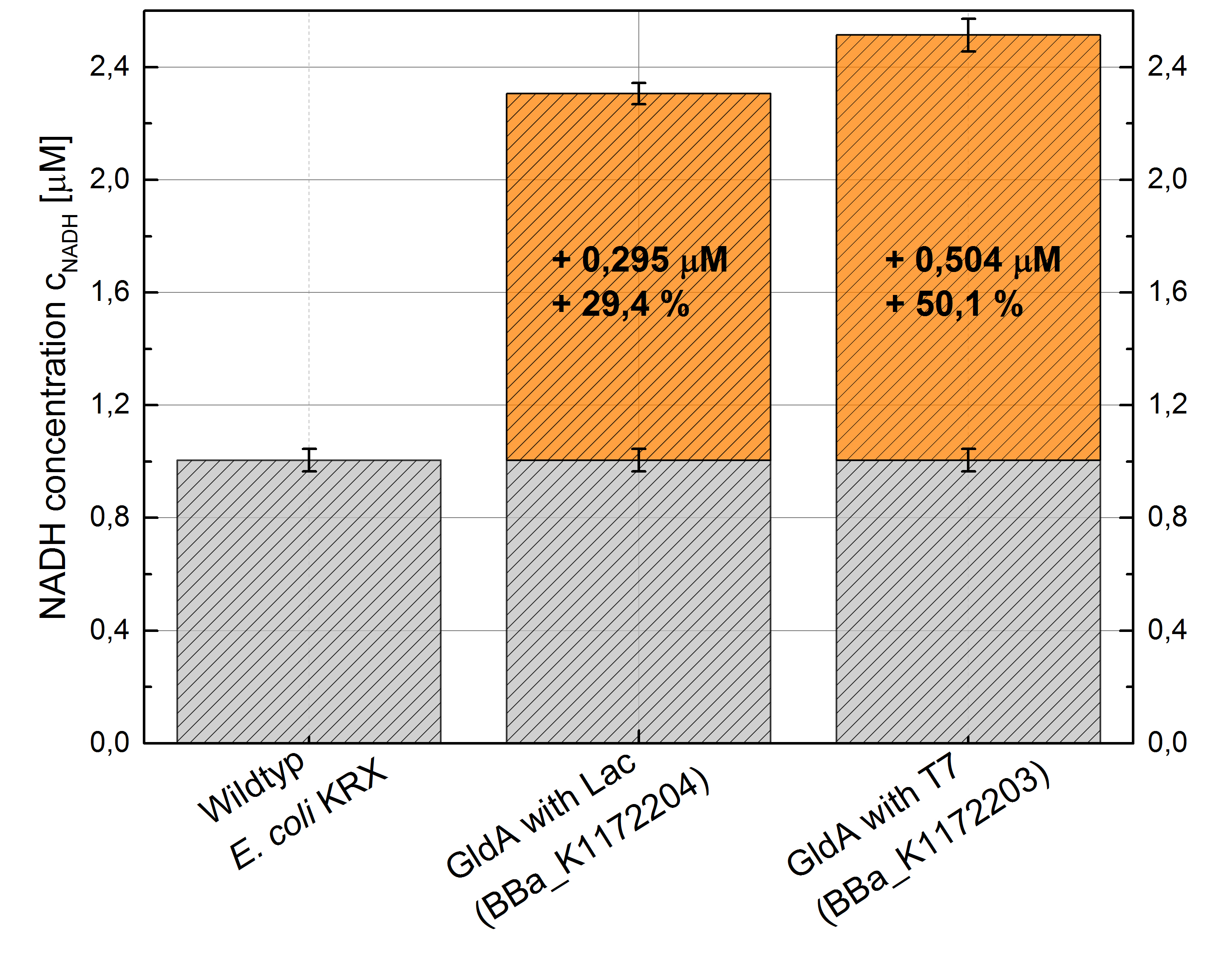
Figure 1: Column Chart with results of the NADH-Assay. Comparison between Escherichia coli KRX Wildtyp and Escherichia coli KRX with <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown.
- Second successful optimized NADH-Assay with Escherichia coli KRX Wildtyp strain and E. coli KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart>, <bbpart>BBa_K1172204</bbpart>, and <bbpart>BBa_K1172205</bbpart>.
- GldA with T7 promotor (<bbpart>BBa_K1172203</bbpart>) shows 1,7 μM NADH overproduction and GldA with Lac promotor (<bbpart>BBa_K1172204</bbpart>) shows 0,2 μM NADH overproduction. The GldA expression by the Anderson promotor (<bbpart>BBa_K1172205</bbpart>) is too weak for an efficient NADH overproduction. The data refer to a culture with OD600 = 5.0.
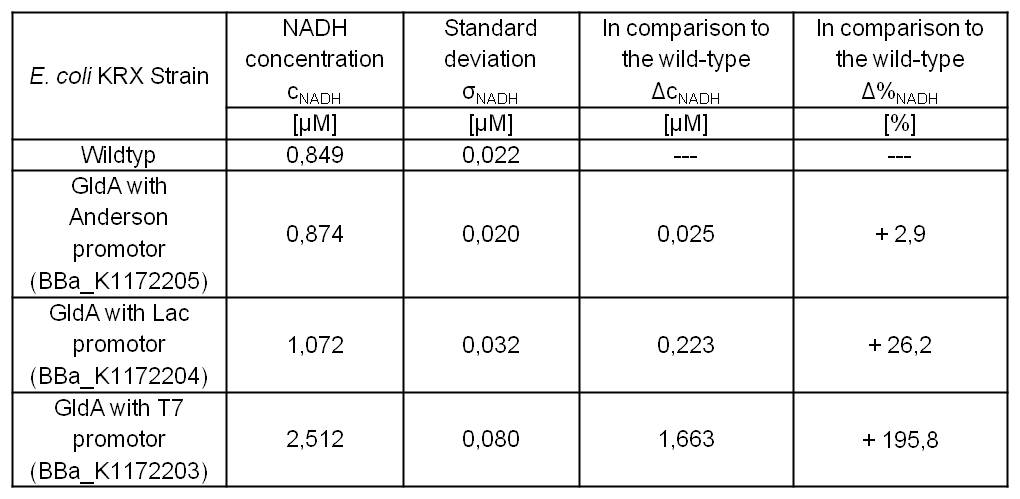
Table 2: Results of the NADH-Assay. Comparison between Escherichia coli KRX Wildtyp and Escherichia coli KRX with <bbpart>BBa_K1172205</bbpart>, <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown.
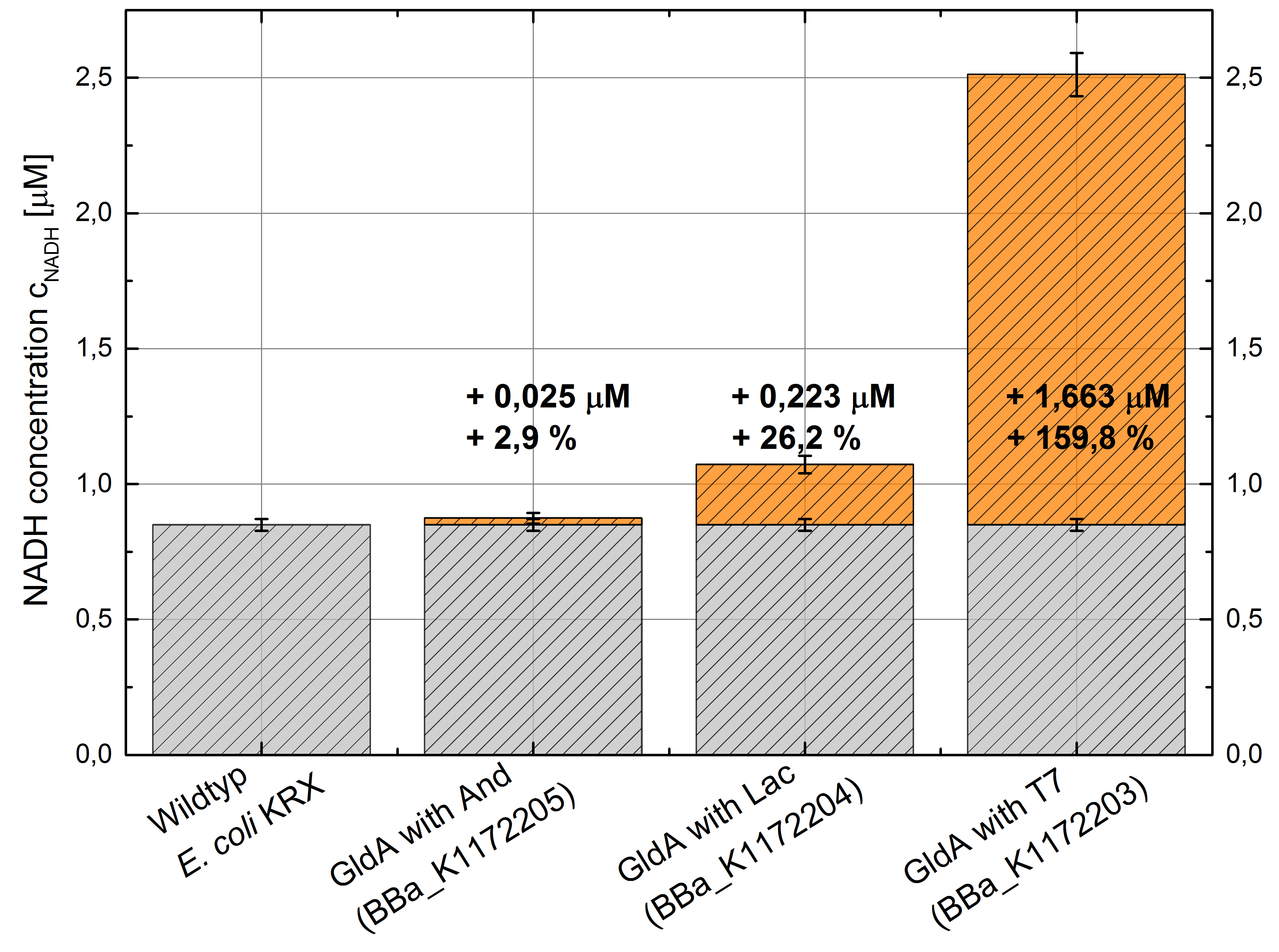
Figure 2: Column Chart with results of the NADH-Assay. Comparison between Escherichia coli KRX Wildtyp and Escherichia coli KRX with <bbpart>BBa_K1172205</bbpart>, <bbpart>BBa_K1172204</bbpart> and <bbpart>BBa_K1172203</bbpart>. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown.
- Overexpressed glycerol dehydrogenase (Figure. 4 Week 17) should be examined by MALDI-TOF MS/MS.
- Tryptic digest of gel lanes for analysis with MALDI-TOF.
- Glycerol dehydrogenase was examined by MALDI-TOF MS/MS with a Mascot Score of 266 against Escherichia coli database.
- Overexpressed glycerol dehydrogenase (Figure. 4 Week 17) should be examined by MALDI-TOF MS/MS.
- Riboflavin
- We tried to ligate our four DNA-Sequences in pSB1C3 using Gibson-Assembly.
- To further minimize the risk of false positive clones just carrying pSB1C3, we digested the pSB1C3-PCR-products with DpnI, prior to Gibson.
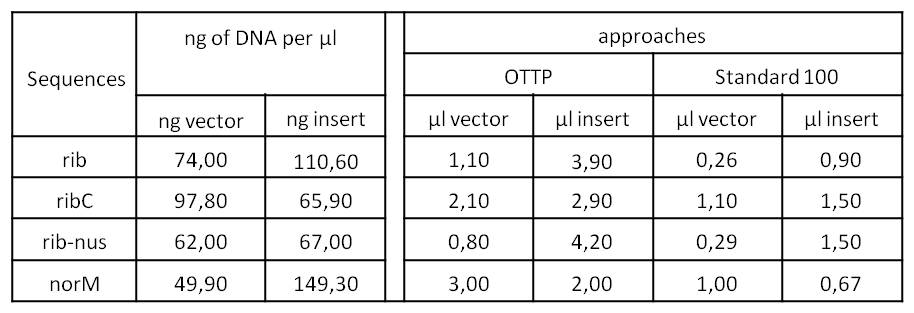
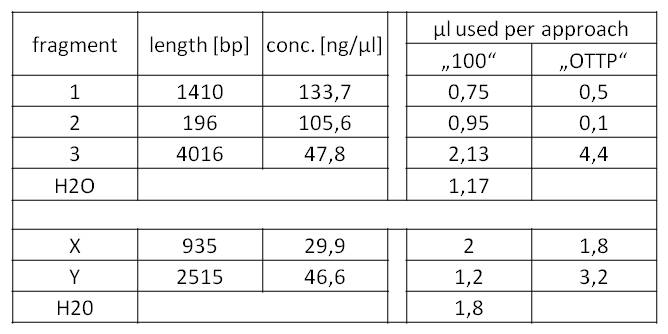
- We assembled every fragment with its specific vector six times. For each fragment, we used two different approaches: The “Standard-protocol” where you take 100ng insert in 3:1 molar excess compared to the vector and an “over the top-protocol” (OTTP). The OTTP uses the same molar excess, but a much higher amount of DNA.
- After picking colonies on ¼ plates we processed the plasmidisolation and test-digested the samples to confirm that the DNA-sequences, still with the illegal sites, were integrated in pSB1C3. Digestions were performed with 250 ng of plasmid-DNA + 2,5 µl of Cut-Smart-Buffer (NEB) + 0,5 µl of enzyme each.
- Due to the illegal restriction site EcoR1, we repeated the digestion of rib and rib-nus only with XbaI.
- We were succesfull in cloning rib, ribC and norM (with illegal sites) in pSB1C3.
- From this point on, we disclaimed rib-nus. It was the only dna sequence we were not able to obtain right away and further research indicated that it lacks significance for our project. So we decided to not waste our precious time on it.
- After the succesfull integration of the three most important sequences in pSB1C3, and the insofar completion of our first Biobrick BBa_K1172302 (ribC), we started with the elimination of the illegal restriction sites in rib and norM.
- Therefore we diluted one rib and one norM sample to 10 ng/µl and used it as PCR template.
- At the beginning of the week, we had started to design primers for this purpose:
- pst1-234 fwd (38 bp)
TGTCACCTTAGAACCCTGTAGCCATTATGGTCGTACGC - pst1-234 rev (32 bp)
CGACCATAATGGCTACAGGGTTCTAAGGTGAC - pst1-1644 fwd (30 bp)
GTGCCCCATACTGTAGGTGAAACCACGTTG - pst1-1644 rev (34 bp)
AACGTGGTTTCACCTACAGTATGGGGCACAATCG - EcoR1 fwd (43 bp)
GGCACTCAGTTCACTTAGGTATAGAATTTATAACAACAGTCAC - EcoR1 rev (43 bp)
GTGACTGTTGTTATAAATTCTATACCTAAGTGAACTGAGTGCC - pst1 norM fwd (38 bp)
GCGATCGTGCTGATATTTCTGCGGTGGTCGCCAAAGTC - pst1 norM rev (41 bp)
CAATAAGCCGACTTTGGCGACCACCGCAGAAATATCAGCAC
- pst1-234 fwd (38 bp)
- our initial phusion-gradient-pcr´s gave good results. The amplification of the fragments at fitting PCR conditions could start:
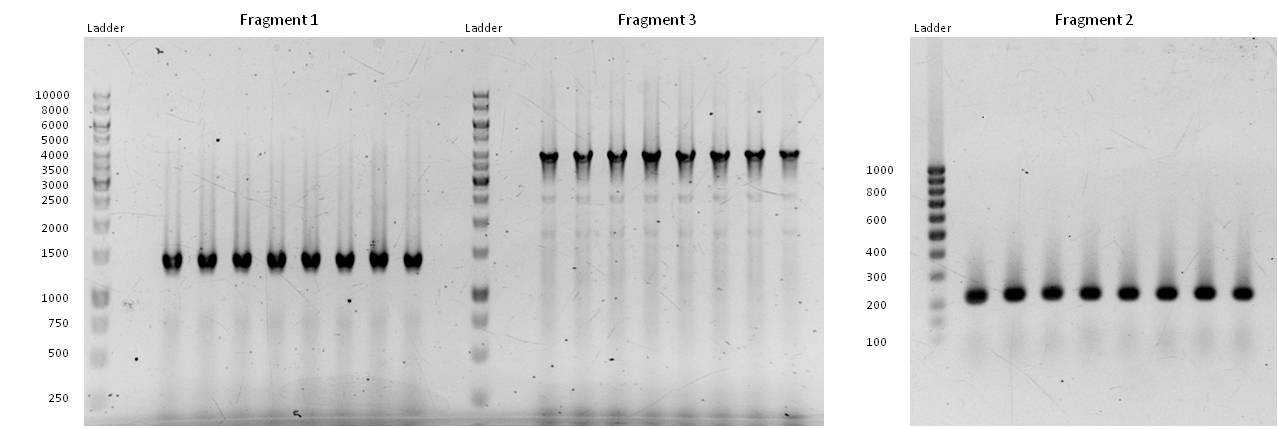
- Fragment 1 had a length of 1410 bp (+overlaps); Primers used: pst1-234 fwd + pst1-1644 rev; Annealing temp.: 69 C°; Elongation time: 22 sec.
Fragment 3 had a length of 4061 bp (+overlaps); Primers used: EcoR1 fwd + pst1-234 rev; Annealing temp.: 68 C°; Elongation time: 61 sec.
Fragment 2 had a length of 196 bp (+overlaps -> 225!); Primers used: pst1-1644 fwd + EcoR1 rev; Annealing temp.: 65,7 C°; Elongation time: 3 sec (worked well in a modern "Eppendorf Thermocycler").
- Fragment 1 had a length of 1410 bp (+overlaps); Primers used: pst1-234 fwd + pst1-1644 rev; Annealing temp.: 69 C°; Elongation time: 22 sec.
- our initial phusion-gradient-pcr´s gave good results. The amplification of the fragments at fitting PCR conditions could start:
- Fragment X had a length of 2515 bp (+overlaps); Primers used: pst1 norM fwd (see week 17) + pSB1C3_rev_norM; Annealing temp.: 72 C°; Elongation time: 30 + 38 sec. (this PCR was run as two-step PCR. The annealing and elongation were run at the same Temperature of 72 C°)
Fragment Y had a length of 935 bp (+overlaps); Primers used: norM_fwd (see week 17) + pst1 norM rev; Annealing temp.: 69 C°; Elongation time: 22 sec.
- Fragment X had a length of 2515 bp (+overlaps); Primers used: pst1 norM fwd (see week 17) + pSB1C3_rev_norM; Annealing temp.: 72 C°; Elongation time: 30 + 38 sec. (this PCR was run as two-step PCR. The annealing and elongation were run at the same Temperature of 72 C°)
Cytochromes
- New primers were ordered. pSB1C3 Gibson primer for the ccmAH cluster and a new reverse pSB1C3 primer for the mtrCAB cluster, which has a larger overlap and should therefore improve assembly efficiency.
- psB1C3_rev_ccm....(40nt) CTGGCTTCAAGCATACCCACCTAGAAGCGGCCGCGAATTC
- psB1C3_fwd_ccm....(41nt) CGCCGCAGGAGAGTAAATAATACTAGTAGCGGCCGCTGCAG
- psB1C3_rev_mtrCAB.(40nt) GATTTTTGTGCGTTCATCATCTAGAAGCGGCCGCGAATTC
- Repeated Amplification of psB1C3 with these new primers with corresponding Gibson-overlaps for mtrCAB and ccmAH.
- Phusion PCR
- Template: Plasmid pSB1C3::RFP (J04500) [5ng]
- Annealing: 60°C
- Elongation: 45s
- The PCR reaction was subsequently digested with DpnI to eliminate any remaining plasmid DNA.
- pSB1C3......4-0609-301......110ng/ul
- Repeated Amplification of fragment3 of the mtrCAB cluster.
- Frag3_fwd (30nt) :GTCTGCAATTATGGCATTAGTCGTCACACC
- mtrB_rev (41nt) :CTGCAGCGGCCGCTACTAGTATTAGAGTTTGTAACTCATGC
- Phusion PCR
- Template: S.oneidensis genomic DNA [5ng]
- Annealing: Gradient [60-70°C]
- Elongation: 50s
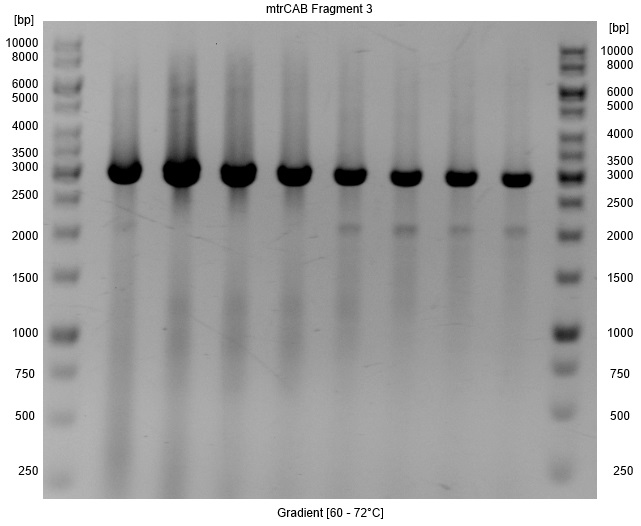
- A Gradient was applied to figure out the annealing temperature with the minimal false bands, so that a PCR clean up with higher yields instead of a gel extraction could be used for isolation. As Figure 4 shows the optimal annealing temperature is around 62-66°C.
- PCR clean up
- Fragment3.......4-0509-305......285ng/ul
- Repeated Amplification of the ccmAH cluster with Gibson primers.
- ccm_fwd (40nt) :GAATTCGCGGCCGCTTCTAGGTGGGTATGCTTGAAGCCAG
- ccm_rev (41nt) :CTGCAGCGGCCGCTACTAGTATTATTTACTCTCCTGCGGCG
- Phusion PCR
- Template: E.coli genomic DNA [5ng]
- Annealing: 68°C
- Elongation: 1:35
- The PCR was cleaned up and lyophilisated with the SpeedVac
- ccmAH......4-0509-301......157.0ng/ul
Biosafety
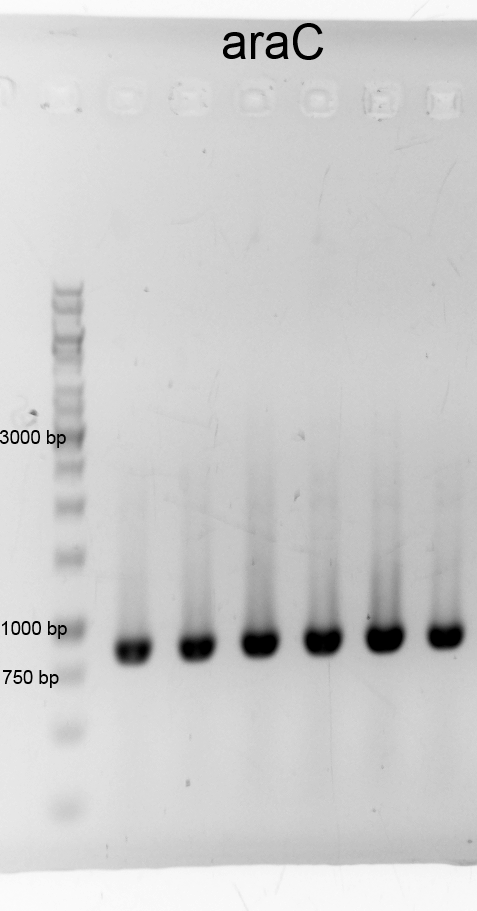
- We executed the characterization of araC by cultivation (200rpm) in shaking flasks with different carbon sources. We used the M9 minimal medium with glucose/arabinose/glycin to measure the growth and the fluorescence of our biosafety system. The fluorescence is caused by the protein GFP which is a replacement for Barnase which is applied for the characterization to draw conclusions from the theoretical expression of the RNase. In continious intervals samples were taken the growth of the cells were measured by absorbance and the fluorescence by a plate reader.
Porines
- Successful Hexadecan-Assay for characterization of hydrophobicity of the outer membrane. Using Escherichia coli KRX Wildtyp and comparing it with:
- <bbpart>BBa_K1172502</bbpart>: <bbpart>BBa_K1172501</bbpart> + <bbpart>BBa_K525998</bbpart>
- <bbpart>BBa_K1172503</bbpart>: <bbpart>BBa_K1172501</bbpart> + <bbpart>BBa_J04500</bbpart>
- <bbpart>BBa_K1172504</bbpart>: <bbpart>BBa_K1172501</bbpart> + <bbpart>BBa_K608007</bbpart>
- <bbpart>BBa_K1172505</bbpart>: <bbpart>BBa_K1172501</bbpart> + <bbpart>BBa_K608006</bbpart>
- <bbpart>BBa_K1172507</bbpart>: <bbpart>BBa_K1172501</bbpart> + <bbpart>BBa_K608002</bbpart>
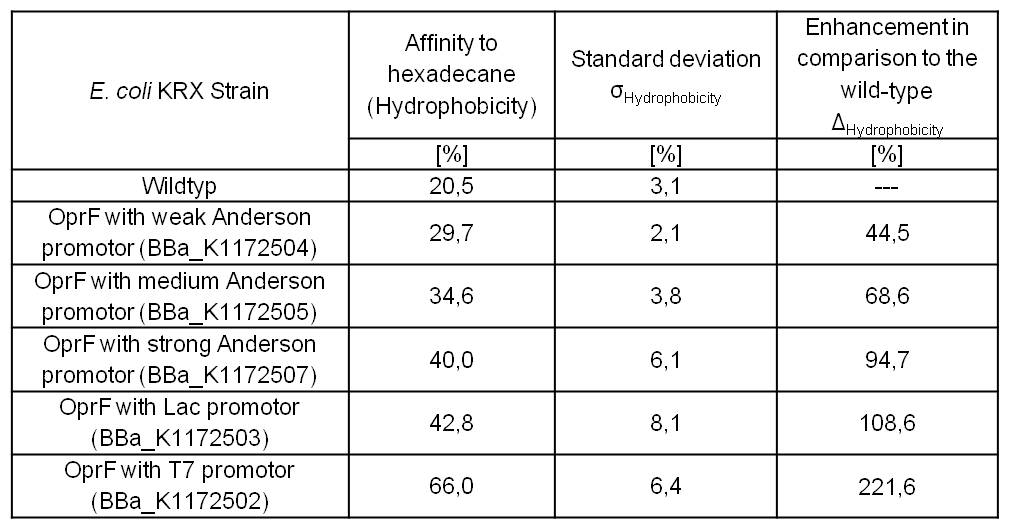
Table 3: Results of the Hexadecane-Hydrophobicity-Assay. Comparison of protein Hydrophobicity between Escherichia coli KRX wild type and Escherichia coli KRX with <bbpart>BBa_K1172502</bbpart>, <bbpart>BBa_K1172503</bbpart>, <bbpart>BBa_K1172504</bbpart>, <bbpart>BBa_K1172505</bbpart> and <bbpart>BBa_K1172507</bbpart>. Affinity to hexadecane (Hydrophobicity) with standard deviation and enhancement in comparison to the wild-type is shown.
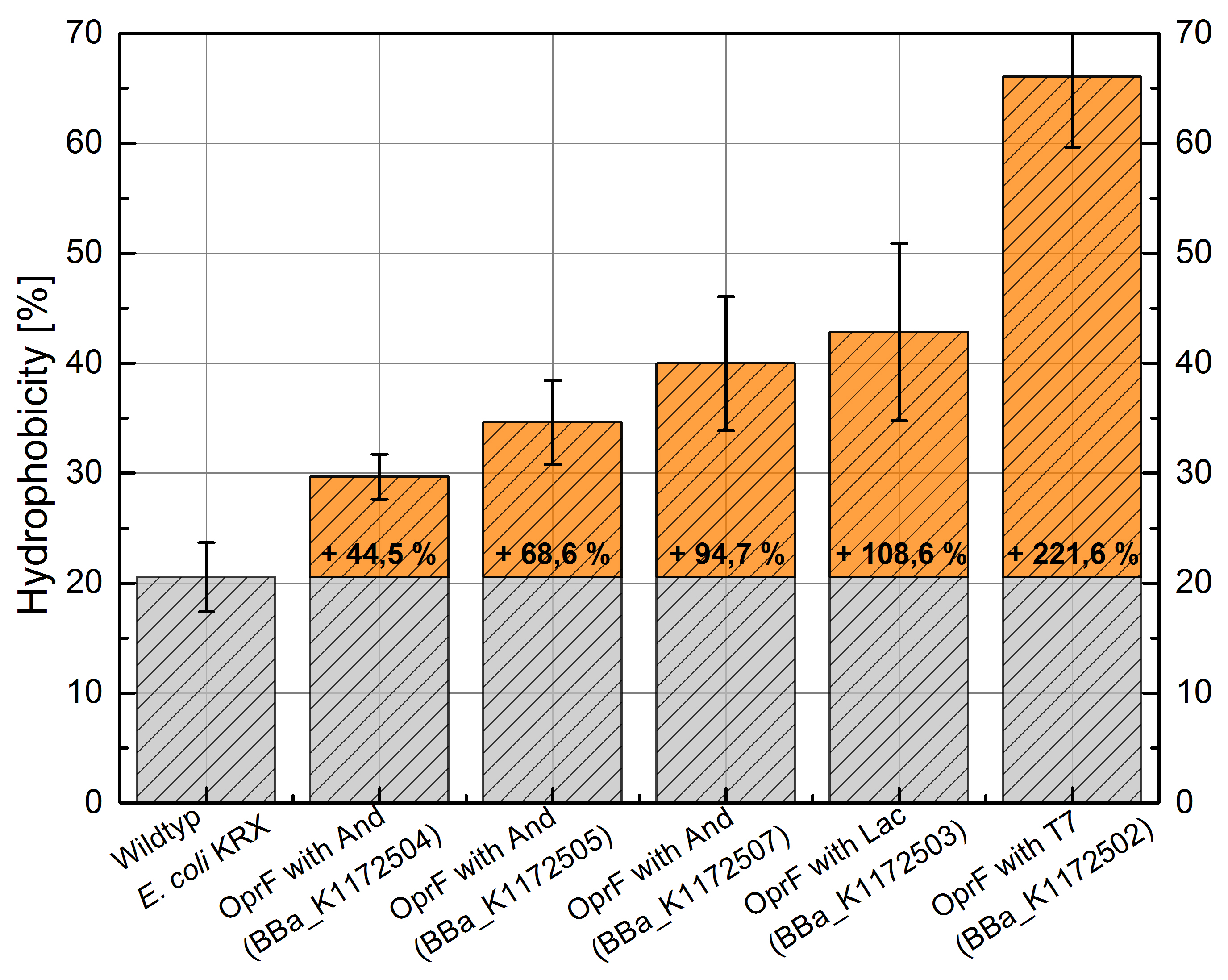
Figure 3: Results of the Hexadecane-Hydrophobicity-Assay. Comparison of protein Hydrophobicity between Escherichia coli KRX wild type and Escherichia coli KRX with <bbpart>BBa_K1172502</bbpart>, <bbpart>BBa_K1172503</bbpart>, <bbpart>BBa_K1172504</bbpart>, <bbpart>BBa_K1172505</bbpart> and <bbpart>BBa_K1172507</bbpart>. Affinity to hexadecane (Hydrophobicity) with standard deviation and enhancement in comparison to the wild-type is shown.
- The hydrophobicity increases continuously with increasing promoter strength up to 221% hydrophobizity in contrast to wildtyp strain.
- Successful ONPG and NPN uptake assays were performed. Membrane permeability is continuously increasing from weak to strong promoter strength.
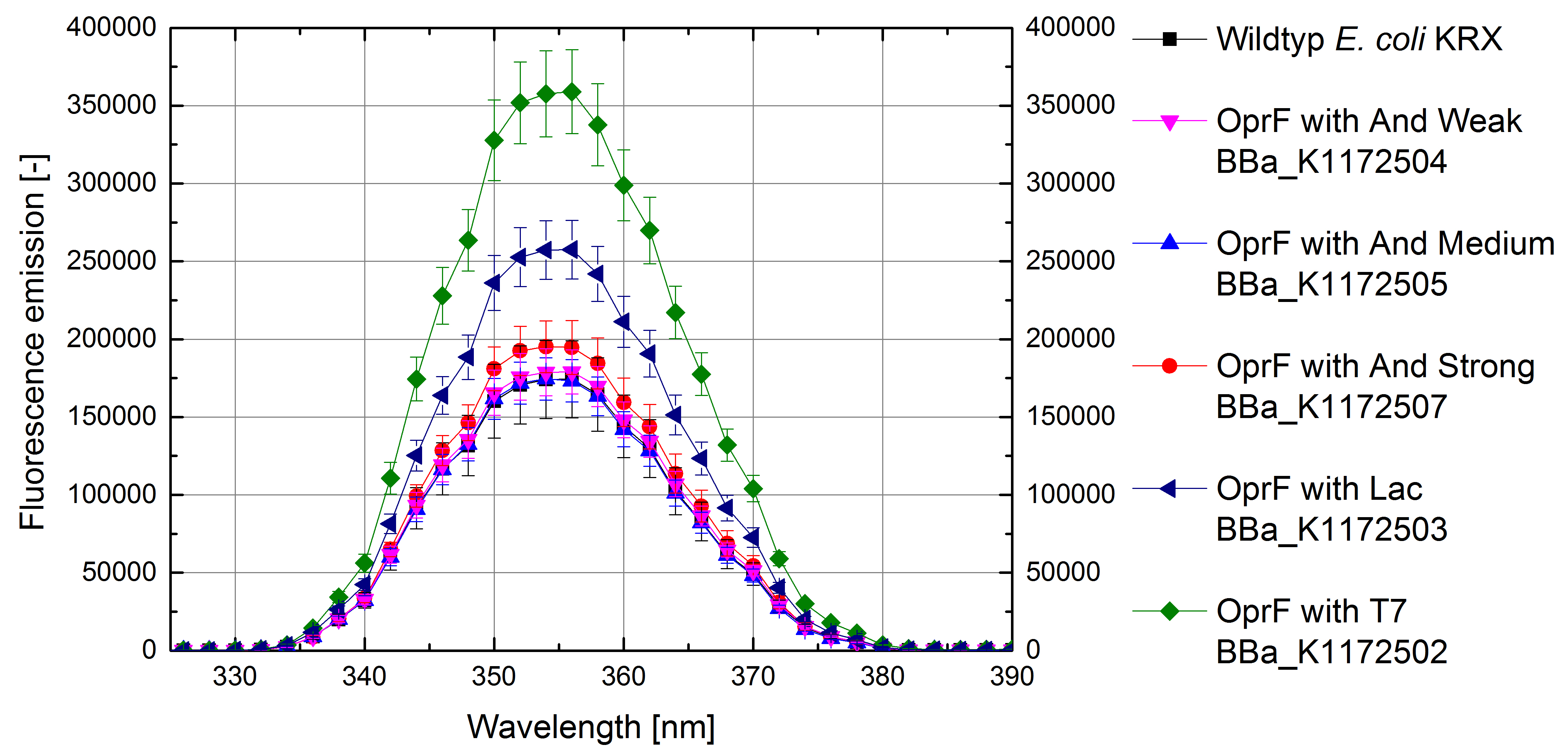
Figure 4: Results of the NPN-uptake-assay. Comparison of fluorescence emission between Escherichia coli KRX wild type and Escherichia coli KRX with <bbpart>BBa_K1172502</bbpart>, <bbpart>BBa_K1172503</bbpart>, <bbpart>BBa_K1172504</bbpart>, <bbpart>BBa_K1172505</bbpart> and <bbpart>BBa_K1172507</bbpart>. Fluorescence emission scan from 320 up to 390 nm wavelength with standard deviation is shown.
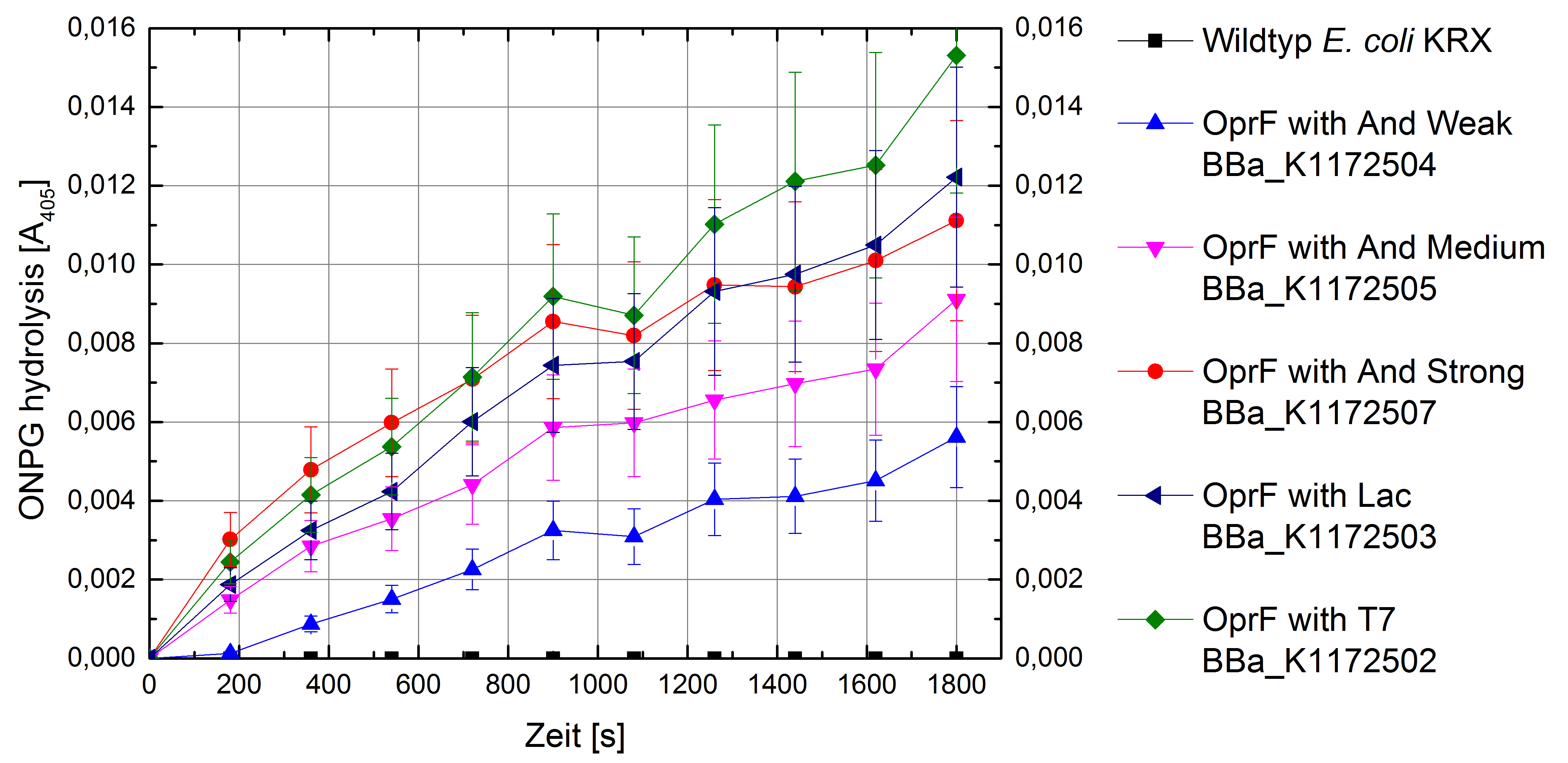
Figure 5: Results of the ONPG-uptake-assay. Comparison of ONPG hydrolysis between Escherichia coli KRX wild type and Escherichia coli KRX with <bbpart>BBa_K1172502</bbpart>, <bbpart>BBa_K1172503</bbpart>, <bbpart>BBa_K1172504</bbpart>, <bbpart>BBa_K1172505</bbpart> and <bbpart>BBa_K1172507</bbpart>. Absorbance at 405 nm wavelength with standard deviation is shown.
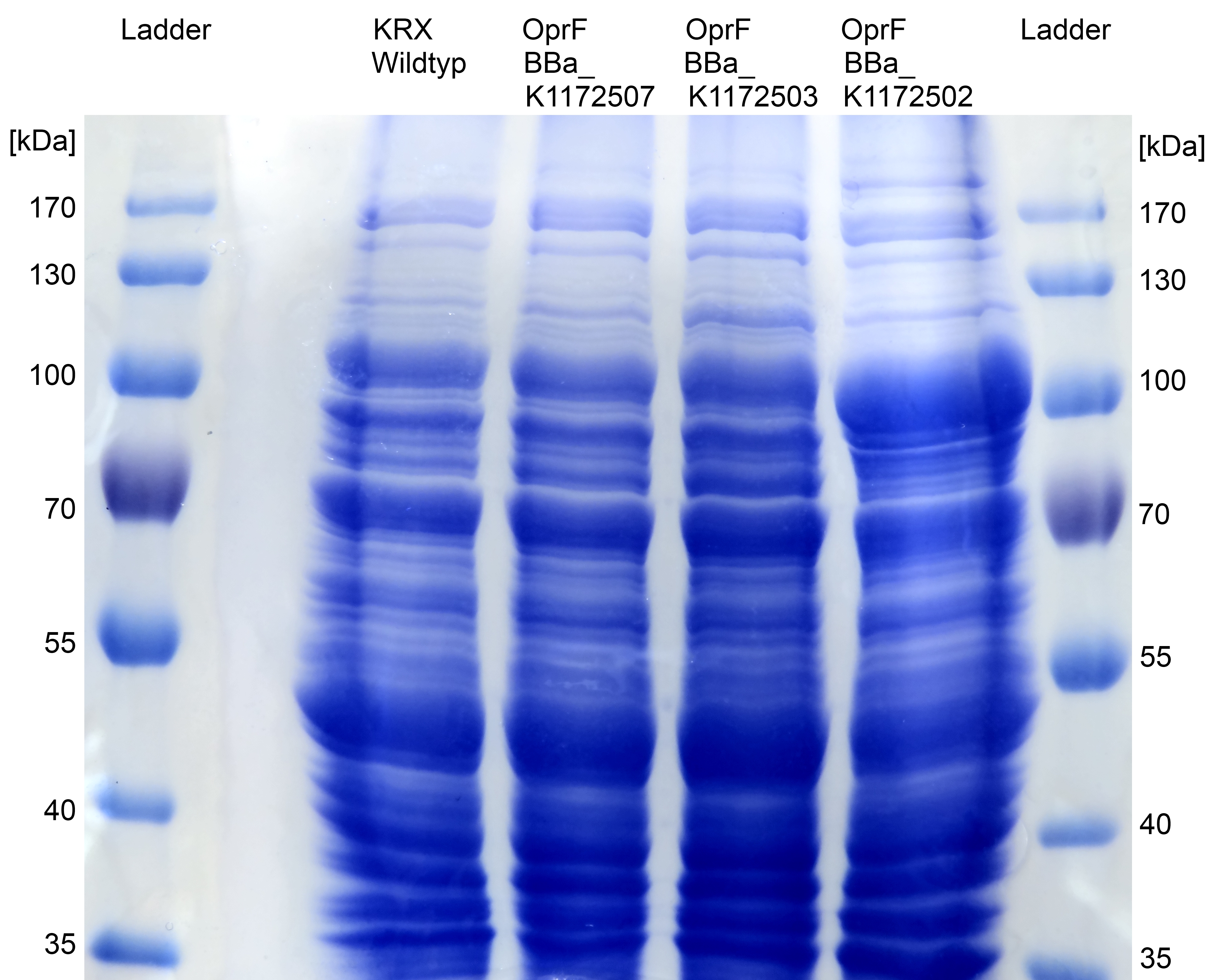
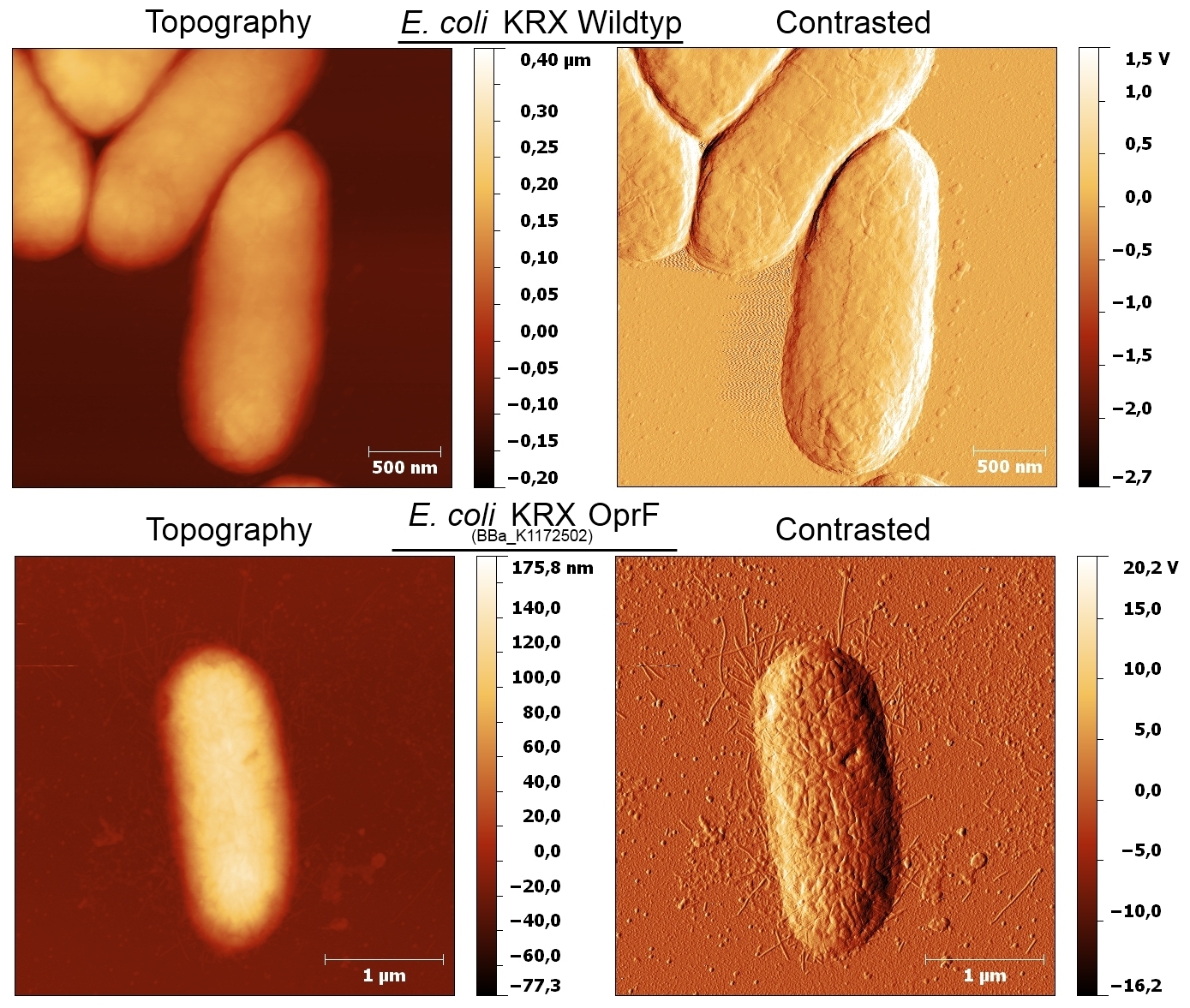
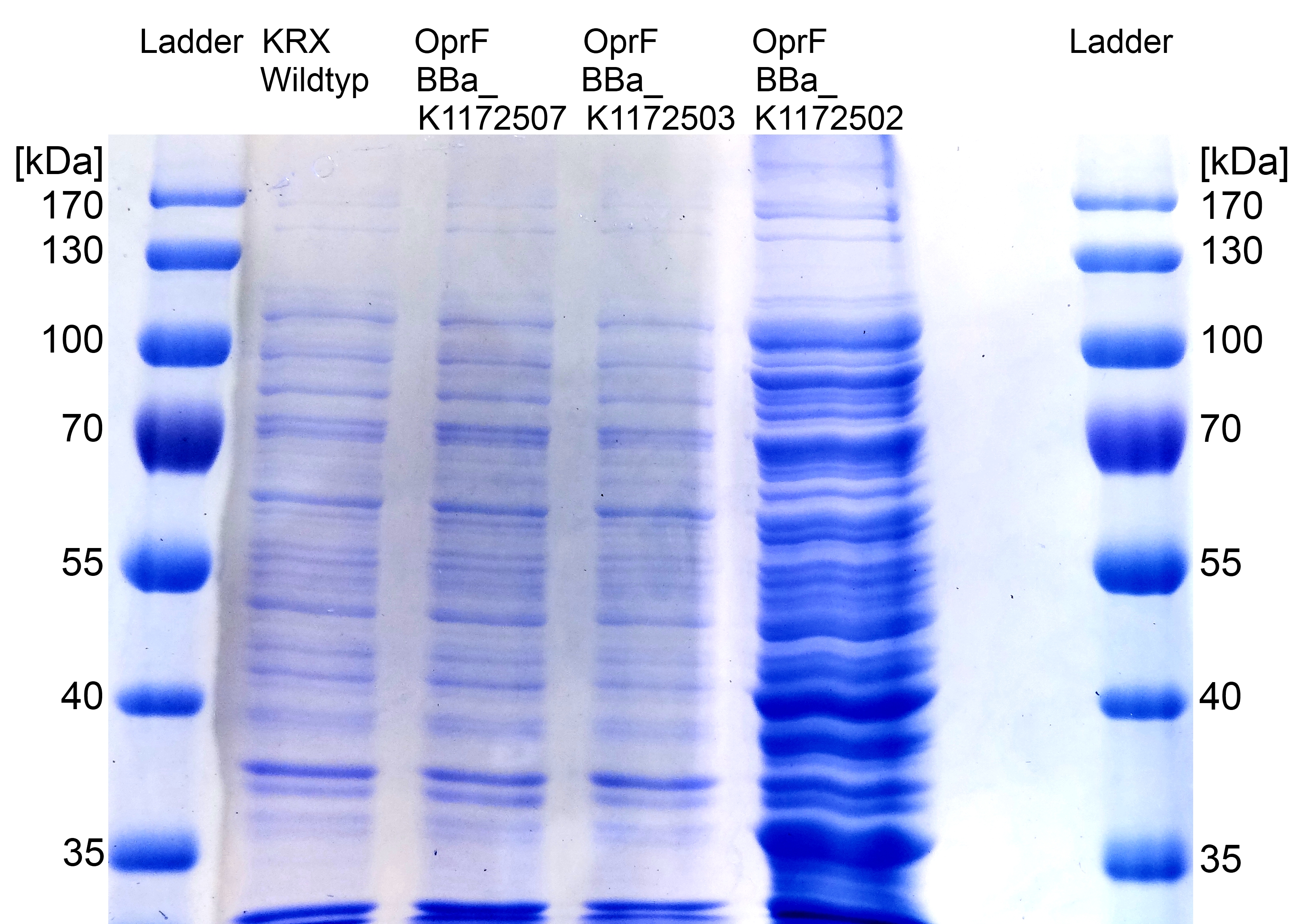
- Beside testing OprF Biobrick (<bbpart>BBa_K1172501</bbpart>) with different assays and SDS-PAGE, the membrane should be visually displayed. The technique of choice is atomic force microscopy.
- The glass slides were coated with Escherichia coli KRX wild type and Escherichia coli KRX with <bbpart>BBa_K1172502</bbpart> in order to compare the membrane differences. The AFM itself was performed with the help of the working group of Prof. Dr. Dario Anselmetti, with special help from Dr. Volker Walhorn.
- AFM was carried out using the MultiMode® 8 AFM from Bruker. The measurements were performed on air with ‘Tapping Mode’ and in water with ‘Peak Force Mode’.
- According to AFM images, Escherichia coli KRX with OprF and T7 promotor (<bbpart>BBa_K1172502</bbpart>) shows a slightly rougher cell surface morphology in contrast to Escherichia coli KRX Wildtyp.
Week 19
Organization
- Our WDR TV contribution could be seen in the ‘OWL local time’ and in the WDR library.
- We have successfully performed the day of synthetic biology in the city of Bielefeld. We were able to explain many people our project with the help of our poster and the kids had a lot of fun with performing different experiments for introducing them into scientific work. DNA isolation from fruit and vegetables, pipetting of bright colors, chromatography with markers, a potato battery and microscopy were on the program. All in all a very successful day.
MFC
Mediators
- Riboflavin
- Due to false bands we were forced to extract Fragment 3 from a gel. We were able to generate a concentration of 2,1 ng/µl and 3,0 ng/µl respectively. So evaporation of the probes in a SpeedVac was necessary. Afterwards, we had practical concentrations of 40 ng/µl and 30 ng/µl.
- For fragments 1 and 2, as well as X and Y, a simple DpnI digest was sufficient.
- 45 µl + 5 µl FD-Buffer + 1 µl FD DpnI -> 80min/37C°; 20min/80C°
- We had now succesfully generated multiple fragments that would not carry any illegal restriction sites.
- Subsequently, we could now start the decisive Gibson assemblies:
- We transformed the eletrocompetent e.coli KRX with 0,8 µl of the assembly mix. Each mix was transformed three times and then transferred to 450 µl SOC-Media. After one hour in a 37 C° water bath, we plated our cells on LB-plates supplemented with chloramphenicol.
- The next day, we found that a staggering amount of cells had been growing on our plates. We were a bit skeptic, but picked eight to sixteen colonies per plate anyway. We diluted the colonies in 20 µl of H20 and plated them on ¼ Cm-Plates.
- Twenty-four hours later we did several miniPreps and executed a test-digestion with XbaI and PstI. The test rendered very good results: BIOBRICKS! <bbpart>BBa_K1172303</bbpart> and <bbpart>BBa_K1172301</bbpart>.
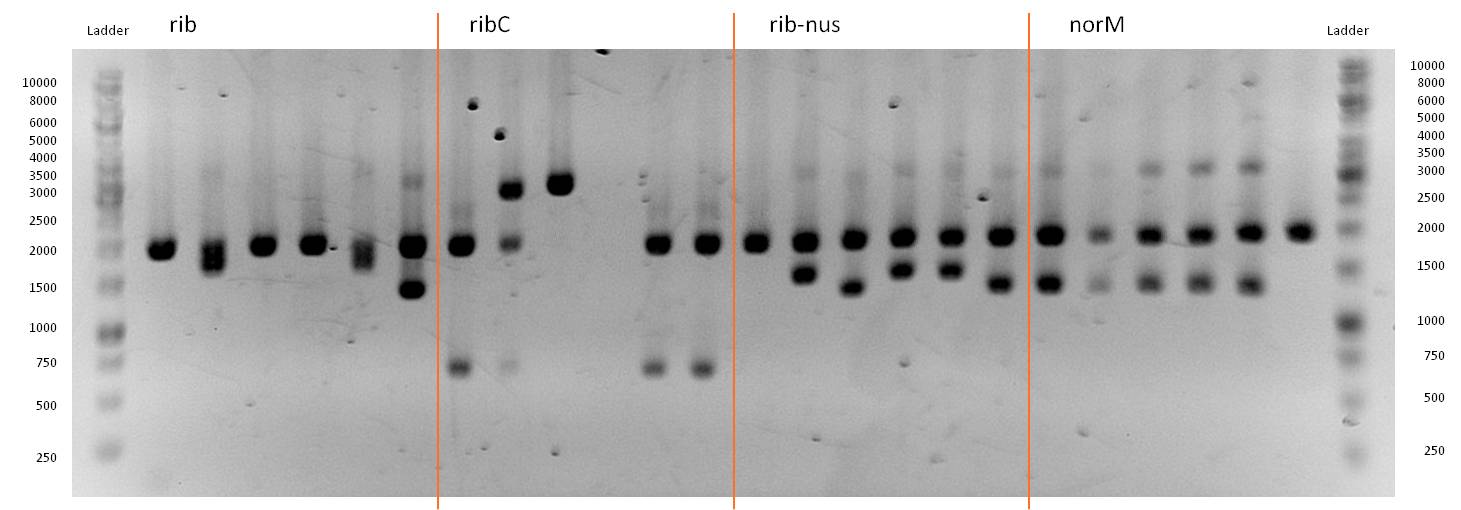
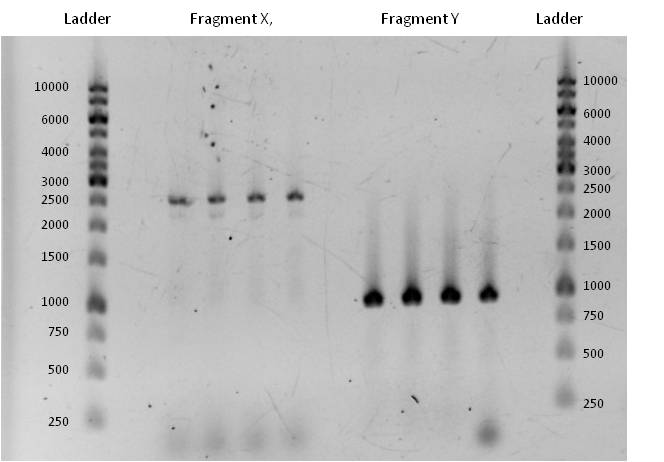
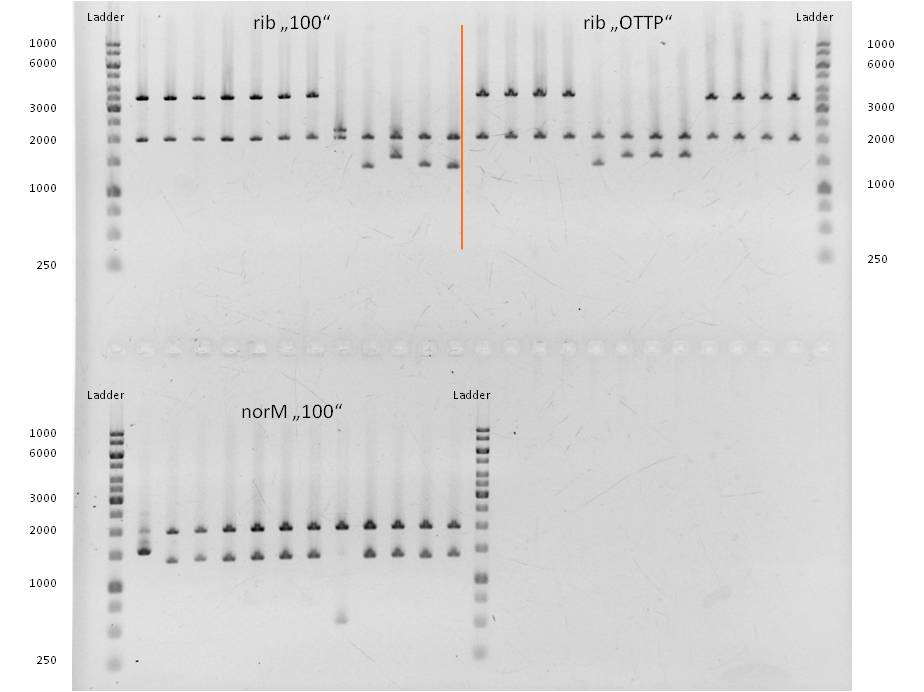
- As you can see above, in figure X, both Gibson aaproaches gave viable results. In the upper part of the gel we can see bands at ca. 2000 bp that belong to linear pSB1C3. The rib-gene-cluster has a length of 3597 bp. The negative bands occurred only when a flash could be seen during electroporation. In the lower part of the gel, we see bands at approx. 1300 bp in every lane but two. 1330bp is the length of norm.
- We were able to eliminate the illegal restriction sites from norm and most importantly the rib-cluster and had them already available in pSB1C3.
- We followed up our positive digestion results with a precautionary sequencing, which proofed the correct form of the BioBricks norM (<bbpart>BBa_K1172301</bbpart>), rib (<bbpart>BBa_K1172303</bbpart>)and although ribC (<bbpart>BBa_K1172302</bbpart>).
- Once again we abandoned parts on our way to riboflavin overexpression in e. coli. The first was norM. The natrium antiporter has enough homologues in e. coli and might be completely unserviceable after a fusion of the rib-cluster and the OprF-gen (Link). This second part left behind was ribC. We resigned on equipping it with promoters. When ribC was taken in by microbial ancestors through horizontal gene transfer, it overtook the job of SO_3468 (which is located in the rib-cluster). Nevertheless, SO_3468 and ribC still share the same function, encode for the same protein and, as SO_3468 lies in the middle of an active operon, we assumed that we might be able to reenact its functionality.
- The most promising BioBrick <bbpart>BBa_K1172303</bbpart> (rib) was then provided with a set of three different Promoters:
- <bbpart>BBa_K608002</bbpart> (strong Andersen promoter and strong RBS) aka And77
- <bbpart>BBa_K608006</bbpart> (medium Anderson promoter and weak RBS) aka And33
- <bbpart>BBa_K525998</bbpart> (T7 Promoter and strong RBS)
- These promoters can be found in the Distribution Kit and are already located in pSB1C3. Therefore they are perfectly suitable for Suffix Insertion. Isolation of Anderson Promoters
- The rib BioBrick (<bbpart>BBa_K1172303</bbpart>) was digested with XbaI and PstI for 1,5 h at 37 C° followed up by an inactivation step at 80 C° for 20 min. Naturally, we than had to extract the insert from a gel and to clean it up.
- The most promising BioBrick <bbpart>BBa_K1172303</bbpart> (rib) was then provided with a set of three different Promoters:
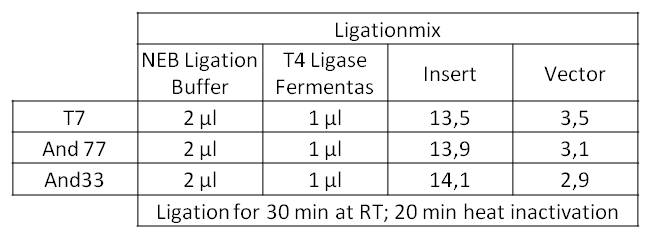
- The Ligationmix looked like the following:
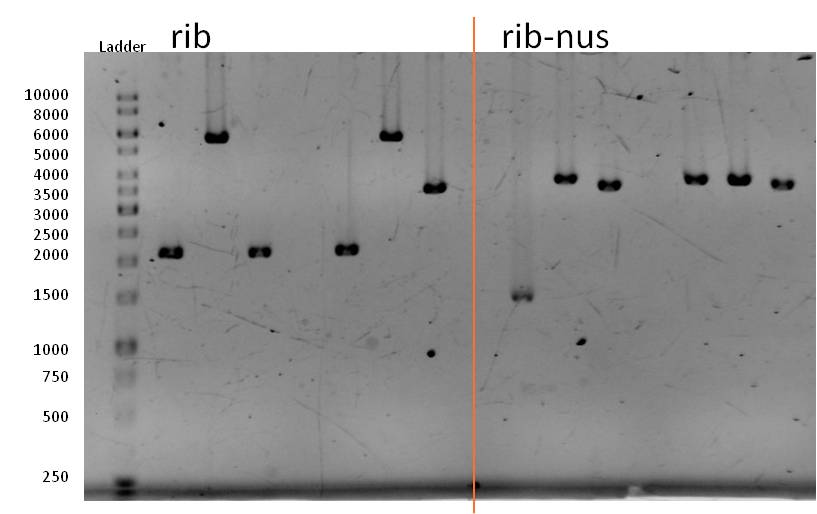
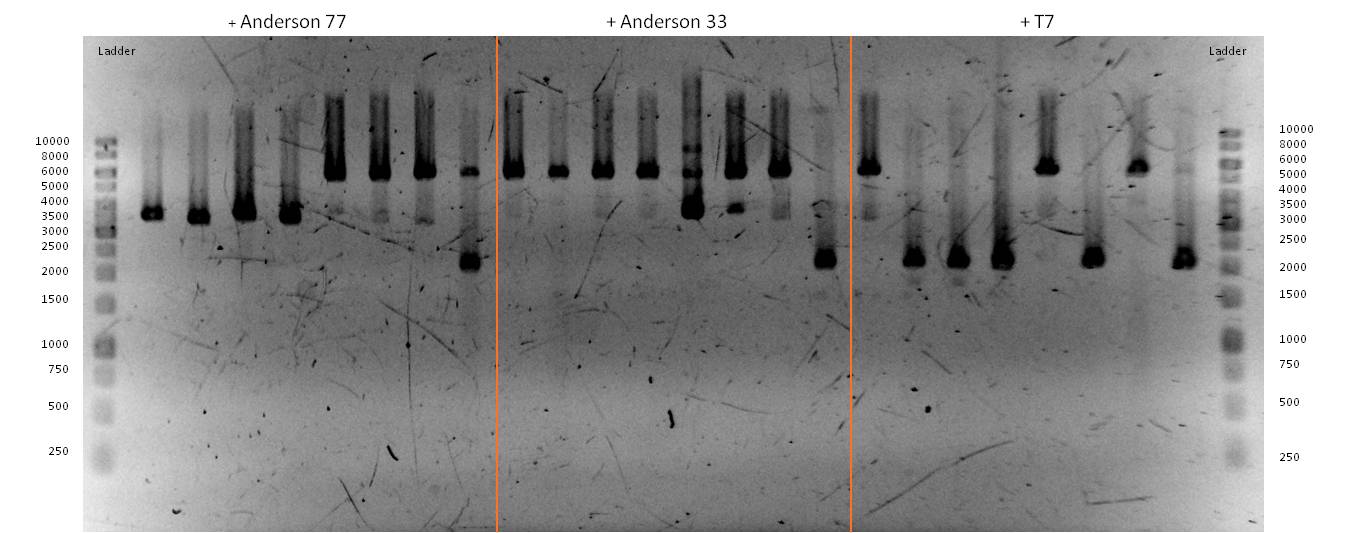
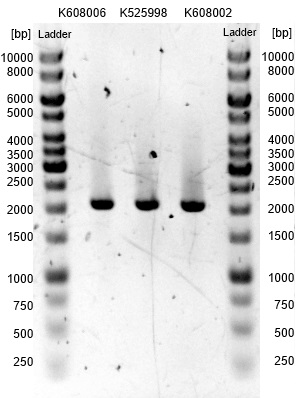
- We transformed the mix three times according to our standard protocol. Two days later we could execute a plasmidisolation and a test-digestion with PstI. Figure X confirms the creation of three new devices:
- <bbpart>BBa_K608002</bbpart> + <bbpart>BBa_K1172303</bbpart> = <bbpart>BBa_K1172306</bbpart>
- <bbpart>BBa_K608006</bbpart> + <bbpart>BBa_K1172303</bbpart> = <bbpart>BBa_K1172305</bbpart>
- <bbpart>BBa_K525998</bbpart> + <bbpart>BBa_K1172303</bbpart> = <bbpart>BBa_K1172304</bbpart>
Cytochromes
- Attempted another Gibson assembly with psB1C3 and the three mtrCAB fragments. The inserts were added equimolar and in a 3-fold excess to the vector.
| psB1C3 | 4-0609-301 | 110ng/ul | 0.6ul | 66ng |
| Fragment1 | 4-2608-302 | 48.8ng/ul | 3.1ul | 151ng |
| Fragment2 | 4-2008-304 | 68.3ng/ul | 0.4ul | 27ng |
| Fragment3 | 4-0509-301 | 285ng/ul | 0.9ul | 256ng |
- Various cells of the transformation were transfered to new plates and incubated again.
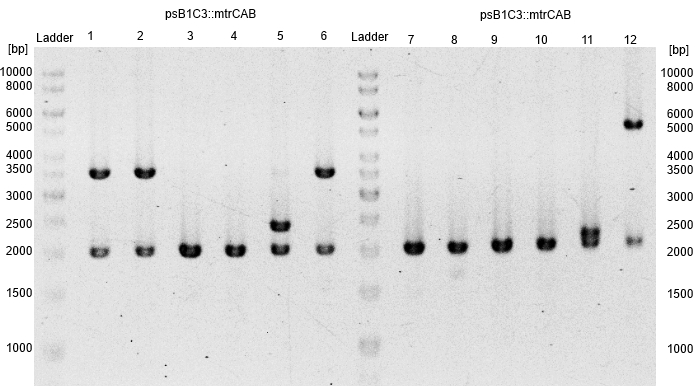
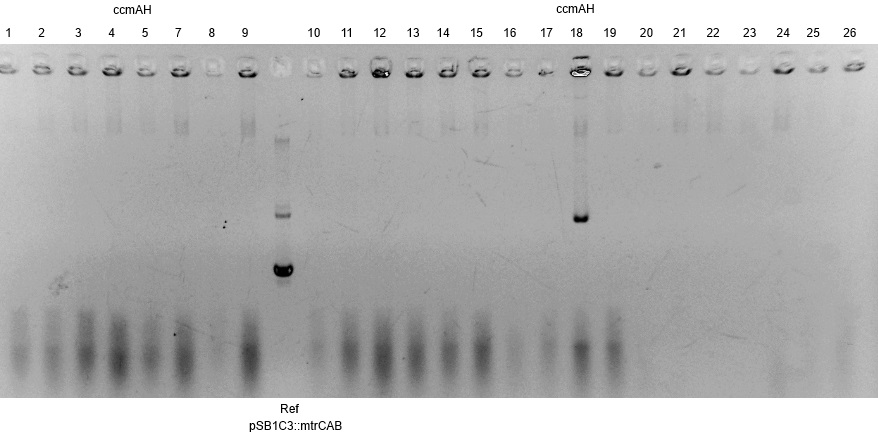
- Subsequently plasmids were isolated from this cultures and a restriction analysis with the enzymes EcoRI and PstI and a total amount of 500ng isolated plasmid DNA was performed.
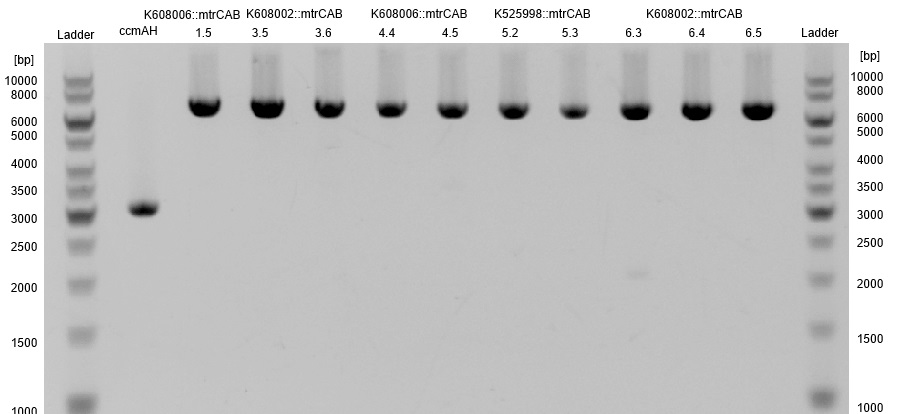
- The resulting fragment should be the respective insert mtrCAB with a size of approx. 5.1kb. The Results are shown in Figure 1 below. The correct sample is located in lane 12.
- An aliquot of the plasmid DNA was used for sequencing.
- The corresponding results verify the correct insertion of the mtrCAB cluster into the psB1C3 backbone.
- Another Gibson assembly was attempted with the ccmAH cluster. The insert was added in a 3-old excess to the vector.
| 4-0609-303 | 4-0509-301 | 75.7ng/ul | 0.7ul | 52.5ng |
| ccmAH | 4-0509-301 | 157.0ng/ul | 3.1ul | 486.7ng |
| Part | Vector | Insert | Volume |
|---|---|---|---|
| K608006::mtrCAB | 50ng | 374ng | 20ul |
| K608002::mtrCAB | 50ng | 374ng | 20ul |
| K525998::mtrCAB | 50ng | 374ng | 20ul |
| K608006::mtrCAB | 10ng | 150ng | 10ul |
| K608002::mtrCAB | 10ng | 150ng | 10ul |
| K525998::mtrCAB | 10ng | 150ng | 10ul |
- Notes: Interestingly the ligations with only 10ng vector and 6-fold excess of the insert seemed to be more effecient, since they yield significantly more colonies.
- The succesfull ligation was confirmed via restriction analysis. The isolated plasmids from the aforementioned transformations were digested with the enzyme EcoRI, which should linearised them and generate a 7.2kb fragment. The Results are shown in Figure3 and verify the correct ligation, leading to the following succesfully constructed devices:
- <bbpart>BBa_K1172403</bbpart>: mtrCAB cluster with Anderson 0.33 and weak rbs
- <bbpart>BBa_K1172404</bbpart>: mtrCAB cluster with Anderson 0.77 and strong rbs
- <bbpart>BBa_K1172405</bbpart>: mtrCAB cluster with T7 and strong rbs
- Subsequently glycerol stocks were made from the ligations and the further characterisation was started.
Biosafety
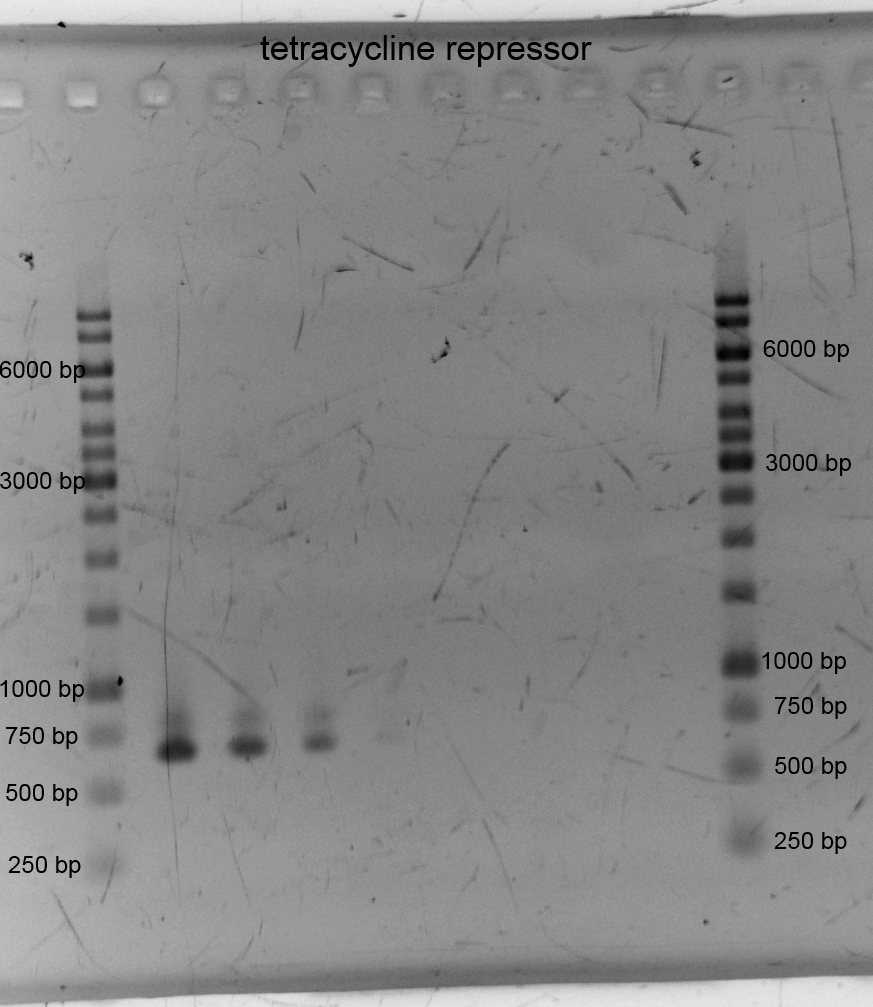
- We transformed the tetracyclin repressor into KRX via electroporation for plasmid isolation. After this we isolated the plasmids and other plasmids which are listend below.
- BBa_C0040_3 250,2 ng/µL (9-108-451)
- BBa_C0040_4 124,5 ng/µL (9-108-452)
- Ptac_alr1 115,8 ng/µL (9-108-453)
- Ptac_alr5 68,1 ng/µL (9-108-454)
- Ptac_alr6 163,6 ng/µL (9-108-455)
- DNase Ba3 86,6 ng/µL (9-108-456)
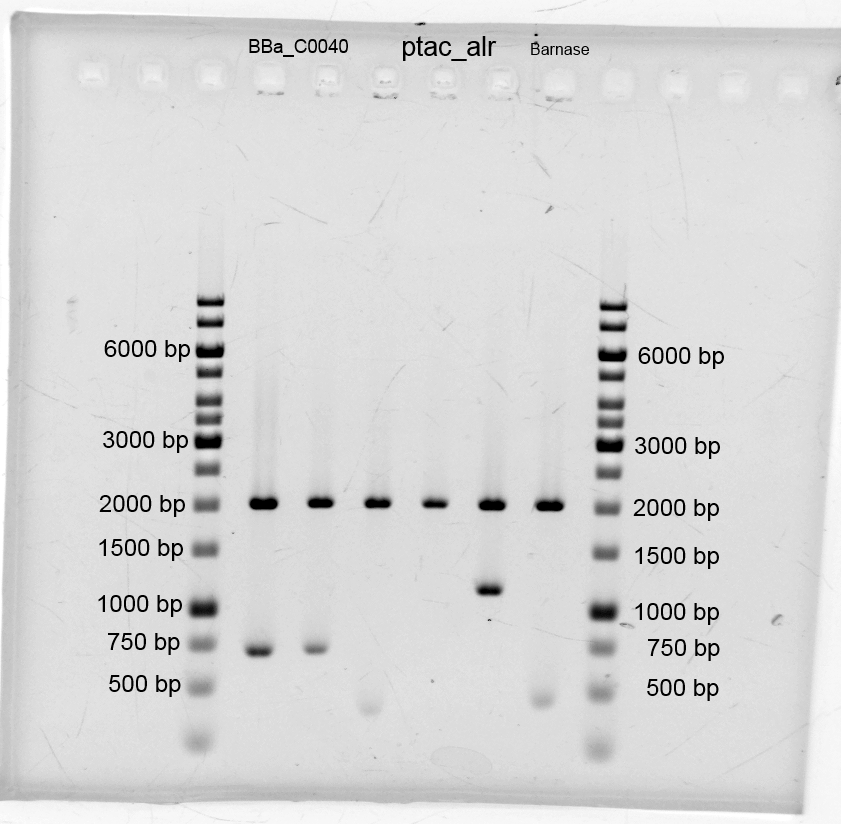
- We did purification of the restriction and measured the dna concentration via nanodrop:
- BBa_C0040 TetO: 35,0 ng/µL
- B0015 Terminator 22,1 ng/µL
- GFP 4,4 ng/µL
- Alr 8,7 ng/µL
- After we isolated the different plasmids we assembled the alanine racemase with the terminator (<bbpart>BBa_K1172003</bbpart>), TetO with GFP (<bbpart>BBa_K1172014</bbpart>) lacI+terminator. We transformed them into competent cells and did plasmid isolation.
- LacI (C0012) 86,5 ng/µL (9-129-451)
- Terminator (B0015) 95,4 ng/µL (9-129-452)
- Terminator (B0015) 46,1 ng/µL (9-129-453)
- Alr_Terminator 84,6 ng/µL (9-129-454)
- Alr_Terminator 65,5 ng/µL (9-129-455)
- Alr_Terminator 33,4 ng/µL (9-129-456)
- Alr_Terminator 98,2 ng/µL (9-129-457)
- TetO_GFP 102,6 ng/µL (9-129-458)
- TetO_GFP 77,1 ng/µL (9-129-459)
- TetO_GFP 91,8 ng/µL (9-129-460)
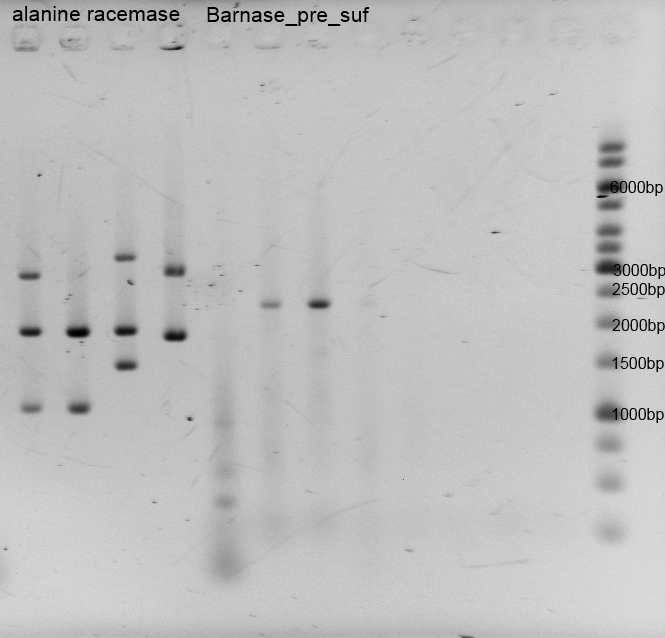
- After the plasmid isolation we checked with a restriction analysis with EcoR1 and Pst1 if the correct inserts are in the vectors. The results are schown below.
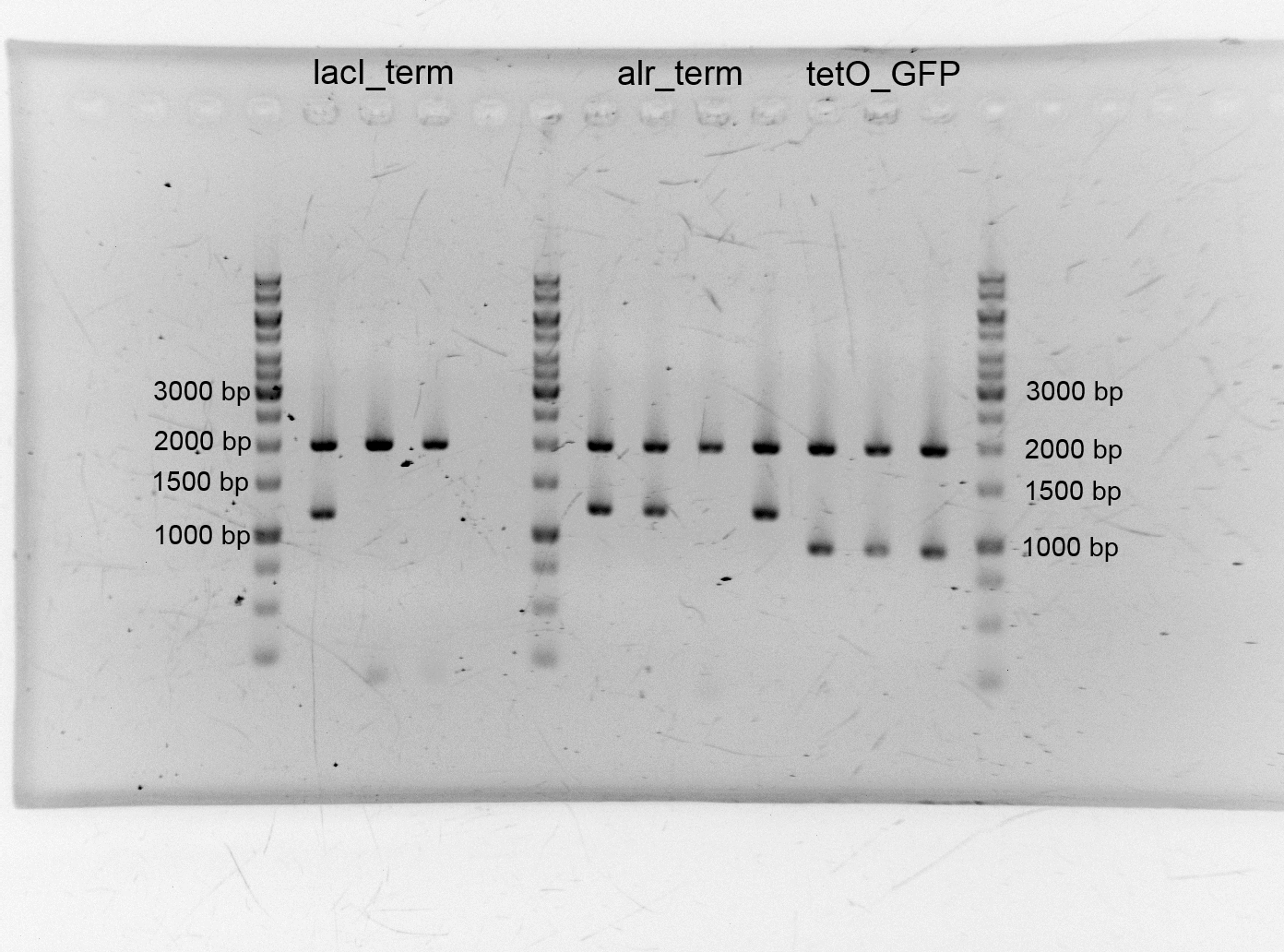
- We send them for sequence analysis to the sequencing service. The results confirmed our constructs.
- We did a PCR to get araC. It was done by executing the phusion protocoll and recipe. The results are listened below:
- After cheking the PCR result we purified the samples with the PCR clean up Kit and measured the DNA concentration.:
- araC 12a 58,8 ng/µL (9-129-463)
- araC 12b 84,1 ng/µL (9-129-464)
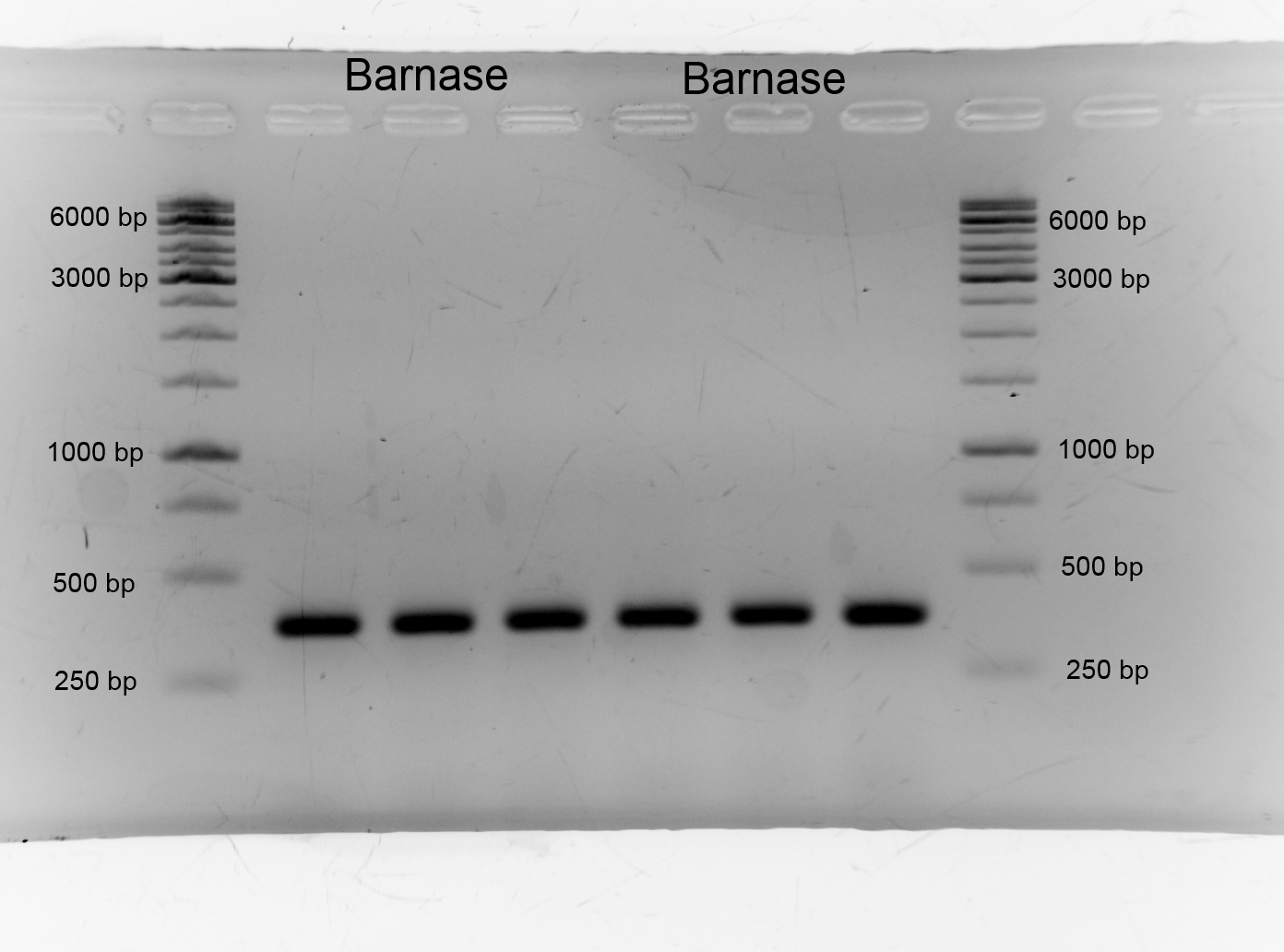
- We did PCR with Tetracylin repressor purificated the samples and did gelelectrophoresis:
- We assembled the three front constructs of our biosafety system via Gibson Assembly:
- BSVaraC
- 1,5 µL pRha (BBa_K914003)
- 2,3 µL alr_terminator (BBa_K1172003)
- 1,5 µL araC
- BSVTetR
- 1,5 µL pRha (BBa_K914003)
- 2,3 µL alr_terminator (BBa_K1172003)
- 1,5 µL TetR (BBa_C0040)
- BSVlacI
- 1,0 µL pRha (BBa_K914003)
- 7,0 µL lacI (BBa_C0012)
- 0,7 µL alr_terminator
- We had assembled TetO with GFP before. pBAD with GFP and plac with GFP were already in the parts registry.
- Back constructs:
- BSVaraC
- pBAD+GFP
- BSVaraC
- BSVTetR
- TetO+GFP
- BSVTetR
- BSVlacI
- plac+GFP
- BSVlacI
Porines
Week 21
Organization
MFC
Mediators
- Glycerol dehydrogenase
- Glycerol dependent NADH-Assay with Escherichia coli KRX Wildtyp strain and E. coli KRX with pSB1C3 and <bbpart>BBa_K1172203</bbpart> by testing different glycerol concentrations of M9-medium.

Table 6: Experimental design of the glycerol dependent NADH-Assay. Different concentrations of glycerol in M9-medium should show the effect of glycerol on NADH overproduction.
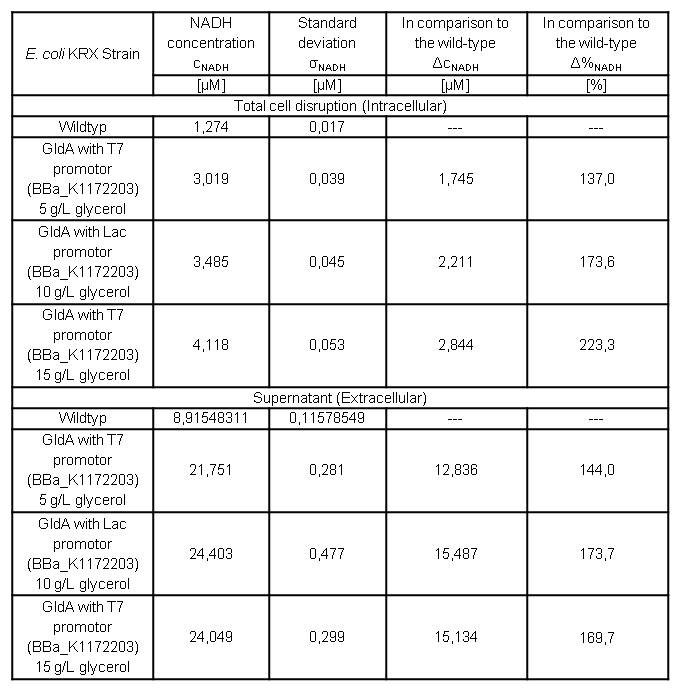
Table 7: Results of the NADH-Assay. Comparison between Escherichia coli KRX Wildtyp and Escherichia coli KRX with <bbpart>BBa_K1172203</bbpart> grown in M9-medium with different concentrations of glycerol. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown for cell disruption and supernatant fraction.
- NADH-Assay shows an increasing intra- and extracellular NADH concentration for <bbpart>BBa_K1172203</bbpart> with increasing glycerol concentration in M9-medium.
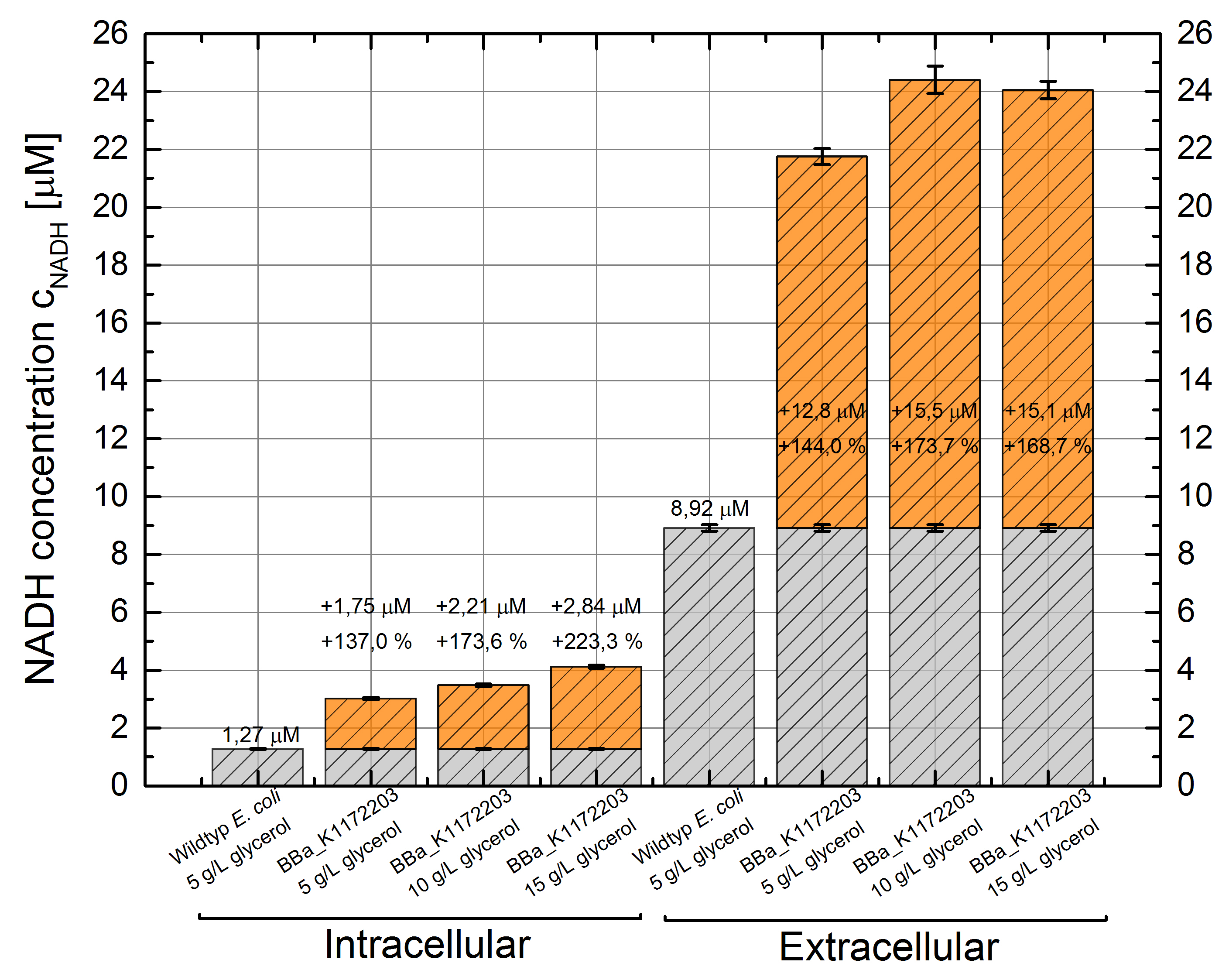
Figure 9: Column Chart with results of the NADH-Assay. Comparison between Escherichia coli KRX Wildtyp and Escherichia coli KRX with <bbpart>BBa_K1172203</bbpart> grown in M9-medium with different concentrations of glycerol. NADH concentration with standard deviation and concentration in comparison to the wild-type is shown for cell disruption (Intracellular) and supernatant (Extracellular) fraction.
Cytochromes
- For characterisation the mtrCAB cluster in with the three different promoters <bbpart>BBa_K1172403</bbpart>, <bbpart>BBa_K1172404</bbpart> and <bbpart>BBa_K1172405</bbpart>,the latter induced and uninduced were cultivated anaerob to ensure the the expression of the ccmAH cluster.
- Subsequently the periplasmic and membrane protein fraction from E. coli was released by cold osmotic shock
- Samples from each fraction were analyzed by SDS-PAGE.
- The resulting gel did not show a significant difference between the samples and the control
- Furthermore a cytochrome type c redox activity assay for the different fractionations was carried out.
- The absorbance was measured at wavelength between 350 and 650nm.
- The oxidized cytochromes were expected to have peaks at 410nm and 530nm.
- The 410nm peak could be measured around 400nm
- The reduced ones should show a shift to 420nm and 552 nm, respectively.
- No significant peak could be measured in this area.
Biosafety
- We cultivated the different biosafety systems for measuring how the specific product building rate looks like in M9 mediumwith different carbon sources. As mentioned before, we used GFP instead of Barnase for characterisation. We sampled in regulary intervals to measure the absorbance and the fluorescence.
Porines
 "
"