Team:Duke/Project/Basics
From 2013.igem.org
Contents |
Background Information
Synthetic Gene Circuits and Bistable Switches
A key feature of synthetic biology is the “bottom-up” design of biological systems using existing biological parts or through the development of novel parts. Of particular interest in the field of synthetic biology is the understanding and development of genetic networks and engineering them to elicit a specific response. Given the dynamic nature of gene-protein interactions that can be explained by chemical kinetics and mathematical modeling, synthetic biologists can utilize these tools to design novel gene circuits guided by quantitative predictions. In 2000, three articles detailed the rational design of a genetic toggle switch, an oscillatory circuit called the repressilator, and an autoregulatory feedback circuit in Escherichia coli. Since then, the regulatory motifs have been expanded and combined to generate new responses such as logical computing or to improve the existing modules, such as robust oscillators.
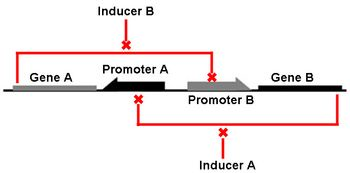
We focus our attention on the genetic toggle switch developed by Gardner, Cantor, and Collins in 2000, as this has guided the design of our project. The genetic toggle switch consists of two repressor proteins λ cI-ts, a temperature sensitive protein, and lacI, the lac repressor, linked in mutual repression in E. coli. The toggling comes from the presence of certain stimuli that turns on or off the production of one of the repressors. If the temperature is high (42 degrees Celsius), then λ cI-ts would be turned on and repress the production of lacI. This could be switched by changing the stimulus to IPTG, a chemical inducer that mimics allolactose to turn on the lacI gene and shut off the λ cI-ts gene. The abil
or the chemical inducer IPTG one repressor protein would be produced
Characteristics of Genetic Toggle Switches
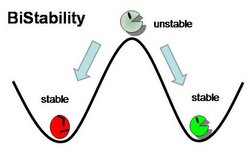
Genetic Toggle Switches have several characteristics which make them unique from other Gene Regulatory networks. For example, Genetic Toggle Switches must be capable of bi-stable behavior, meaning that they should only exist in one of two stable states instead of in a variety of intermediate states. Bi-stability also suggests that a certain threshold must be passed before the toggle switch can switch from one state to another. In addition, toggle switches should have low transcriptional noise to prevent random switching without induction. Finally, a reporter or marker structural gene, such as a fluorescent protein, should be present in order to characterize the effectiveness of the toggle switch. <From: iGEM 2011 Team Duke>
The graph below shows nullclines for the level of two mutually repressive proteins in a bistable toggle switch. Each curve on the graph represents concentrations of lacI and cI that makes its change in concentrations respect to time (ie. d[lacI]/dt = 0, d[cI]/dt=0). These curves are called nullclines, and the points at which the two curves intersect are steady-state solutions of the system because the concentrations of the two repressors do not change with time. However, mathematical analysis shows that the intersection in the middle is an unstable steady state, where minute perturbations can quickly shift the equilibrium point to either direction. On the other hand, the two points labeled “State 1” and “State 2” are stable stead-state solutions. At these two points, the concentration of only one repressor is high while the concentration of the other repressor is minimal. Therefore, with these two stable steady-state solutions, the system is called bistable. Only when the majority repressor is degraded or the minority repressor is over-expressed to “go-over” the separatrix line, the system will flip to the opposite steady state (hence the name bistable toggle switch).
TAL Effector
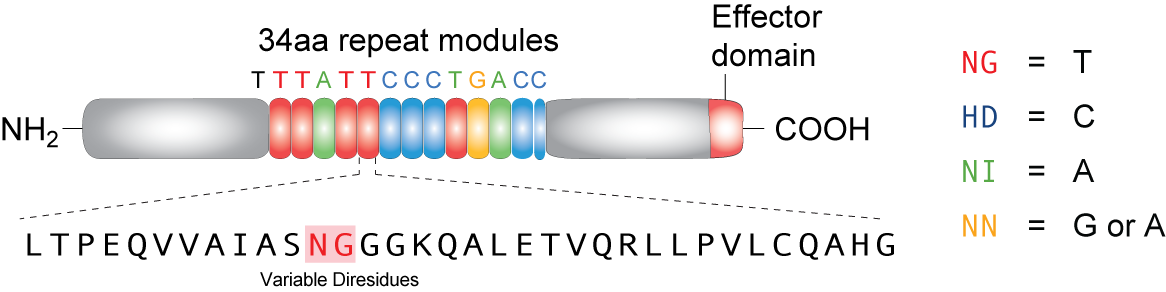
TALEs (Transcription Activator-Like effectors or TAL effectors) are DNA binding proteins derived from a plant pathogen Xanthomonas sp. which uses these proteins to regulate the host plant’s genome and aid in bacterial infection. Biochemical analysis of the TALEs revealed a highly repetitive primary structure with two residues at positions 12 and 13 out of a 33-34 amino acid protein that varied based on the target nucleotide. These two amino acids, referred to as repeat variable di-residues (RVDs) can be selected to target any DNA sequence and linked together to bind sequences up to 30 nucleotides long. The four RVDs that we utilize are NG targeting T, HD targeting C, NI targeting A, and NN targeting G, although other RVDs exist that target various nucleotides with varying affinity. Using Golden Gate cloning, we are able to rapidly synthesize TALEs for uses in regulating gene expression by fusing activator or repressor domains to the TALEs. In addition, TALEs have been fused to nuclease domains (TALENs) for applications in genetic engineering and genome editing in plant and mammal cells by editing specific mutations, knocking out genes, or adding genes. We utilize a TALE fused to the fluorescent protein mCherry to act as a transcriptional roadblock rather than a repressor protein with the hopes of achieving the same result.
CRISPR
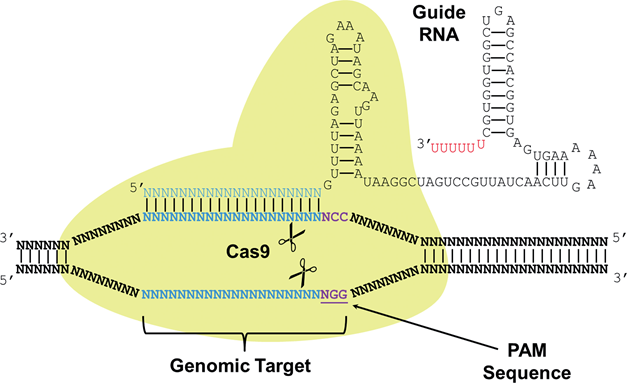
CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) are a part of a system joined with Cas (CRISPR associated) proteins that function as a immune system found in around 90% of archaea and 40% of bacteria species. CRISPR-Cas systems recognize foreign DNA sequences and silence them through interactions with Cas proteins that have nuclease or helicase activity. A small RNA called sgRNA (small-guide RNA) recognizes a specific DNA sequence and contains a region of significant secondary structure called the Cas9 handle that Cas proteins bind to either cut the DNA sequence or silence it. The use of the CRISPR-Cas9 system as a tool for genetic engineering is recent and rapidly growing because of the ability to multiplex the genome editing process by designing multiple sgRNAs targeting various regions in the genome to be edited simultaneously in conjunction with the nuclease activity of Cas9. Researchers have also discovered two nonsynonymous amino acid substitutions that deactivate the nuclease activity of Cas9 while preserving the binding activity of Cas9 to sgRNAs. This catalytically dead Cas9 is referred to as dCas9 and can be utilized in a similar fashion to TALEs where sgRNAs target promoters and the dCas9 protein acts as a roadblock to transcriptional activation. Recently, activating and repressing domains were fused to dCas9 to develop a gene regulation CRISPR/dCas9 system. We utilize the dCas9 protein in order to cause a transcriptional roadblock for our toggle switch, very much like a repressor protein.
References
DiCarlo et al., 2013
- Gardener, T. et al. Construction of a genetic toggle switch in Escherichia coli. Nature. 403, 339-342 (2000).
- TAL Effectors Resources. Introduction to TAL Effectors. Accessd on 9/3/13 <http://www.genome-engineering.org/taleffectors/>.
- DiCarlo, J. et al. Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems. Nucleic Acids Research. 41(7), 4336-4343 (2013).
 "
"