Team:TU-Munich/Results/GM-Moss
From 2013.igem.org
(→1. Generation of expression constructs) |
|||
| Line 5: | Line 5: | ||
<div id="wikicontent"> | <div id="wikicontent"> | ||
<!-- Start of content --> | <!-- Start of content --> | ||
| - | + | <!-- | |
== Creation of transgenic ''Physcomitrella patens'' plants== | == Creation of transgenic ''Physcomitrella patens'' plants== | ||
| Line 212: | Line 212: | ||
==4. Regeneration and Selection of transgenic plants== | ==4. Regeneration and Selection of transgenic plants== | ||
| - | After Transformation, the regenerating protoplasts were incubated in regeneration medium in 6-well-plates sealed with parafilm for 10 days and then cultivated on a layer of autoclaved cellophane (seperated with Whatman paper during autoclavation, so they dont stick together), on top of Knop medium agar plates for three days under standard conditions. Then we transferred the cellophane layers with the regenerated moss onto Knop medium agar plates containing 25 µg/ml G418 antibiotic for two weeks. The official protocol schedules a two week release period followed by another selection period to ensure stabile transormation, but we didn´t have that much time and went with a single round of selection. We plated only half of our transformed protoplasts. The other half was left in the 6-well-plates where 2 ml of selection medium were added to the 2 ml of regeneration medium from the transormation, with G418 diluted 1:8000. | + | After Transformation, the regenerating protoplasts were incubated in regeneration medium in 6-well-plates sealed with parafilm for 10 days and then cultivated on a layer of autoclaved cellophane (seperated with Whatman paper during autoclavation, so they dont stick together), on top of Knop medium agar plates for three days under standard conditions. Then we transferred the cellophane layers with the regenerated moss onto Knop medium agar plates containing 25 µg/ml G418 antibiotic for two weeks. The official protocol schedules a two week release period followed by another selection period to ensure stabile transormation, but we didn´t have that much time and went with a single round of selection. We plated only half of our transformed protoplasts. The other half was left in the 6-well-plates where 2 ml of selection medium were added to the 2 ml of regeneration medium from the transormation, with G418 diluted 1:8000.--> |
==5. Characterization of transgenic plants== | ==5. Characterization of transgenic plants== | ||
Revision as of 01:20, 28 October 2013
5. Characterization of transgenic plants
5.1 Testing our modular moss receptor
Idea - GFP fluorescence
Already during the planning phase of the trans membrane receptor we thought about how to track the localization of the receptor – or in other words how to proof the correct folding into the plasma membrane. As a very simple tool we fused genetically a GFP molecule C-terminal to the trans membrane domain of the receptor. A green fluorescence localized at the plasma membrane would hence indicate a correct folding of our receptor into the plasma membrane.
The animated Z-stack of transgenic moss cells expressing the trans membrane receptor presented in Figure 5 shows a bright green fluorescence as is characteristic for GFP. This fluorescence is mainly located at the plasma membrane which would indicate that our receptor proteins have the correct localization. In comparison to the Z-stack taken from transgenic moss expressing GFP in the cytoplasm which is shown in figure six the fluorescence of the moss carrying the trans membrane receptor seems to be more associated to the plasma membrane. However voluminous vacuoles can compress the cytoplasm to a thin layer so that the green fluorescence of cytoplasmic proteins would appear as membrane associated. Hence the fluorescence shown in figure 5 is a first indication that our receptor works properly but cannot serve as a reliable evidence. Therefore we decided to execute a second method for the proof of localization of our trans membrane receptor. Furthermore it is planned for the future to do confocal laser microscopy using fluorescein-biotin conjugate and quantum dots coupled to streptavidin. Therewith the localization of the trans membrane receptor carrying FluA as an effector domain at the N-terminus should be proofed doubtless through bright fluorescence of the used fluorophores on the outside of the plasma membrane and green fluorescence of the receptor GFP on the inner side of the plasma membrane.
2. Idea - Protease cleavage of the effector protein by TEV protease
We want to investigate the localisation of our synthetic receptor. For this purpose we used a transgenic moss strain which contains NanoLuc luciferase at the extracellular domain of the receptor. Between the luciferase and the transmembrane region we have added a linker consisting of a Streptag II and a TEV cleavage site. By adding the recombinant TEV protease, the TEV protease should be able to pass the cell wall and enter the appoplast, cleave the TEV cleavage site and liberate the NanoLuc luciferase. Afterwards the luciferase can be analyzed by its luminescense, the Streptag or by mass spectrometry.
To prove this process, we cut off our Nanoluciferase-Streptag-Construct by using the TEV-Protease. As we hypothetically know, our membrane construct is localized in the membrane and the Nanoluciferase-Streptag-Construct orientated to the Periplasma, protected by the cell-wall of our moss. In conclusion to that, the TEV protease needs to pass the cell-wall to cut off the Nanoluciferase-Streptag-Construct. This process is possible in liquid medium, because the TEV protease just has a molecular weight between 25 kDa and 27 kDa, so it can pass the cell-wall smoothly. After the TEV-Protease has entered the Periplasma of our moss cells, it cuts off the Nanoluciferase-Streptag-Construct at the TEV cleavage site. Again, this construct has in sum a molecular weight of 20 kDa (Nanoluciferase - 19 kDa, Streptag - 1 kDa), so it can pass the cell-wall due to diffusion to get to the exterior of the moss cell.
After this experimental part, it is necessary to prove the theoretical deliberation. In the following we performed several experiments like SDS-Pages, Luciferase Assays and tried to bind our construct by utilizing different beads.
Experimental Setup for localization process of membrane bound Nanoluciferase
First Step
The reagents which were necessary to perform the first step of the experiment have been 1,863 ml TEV Protease Buffer, 137 µl TEV Protease (0.364 mg/ml) to get a total enzyme activity of 500 Units in each sample and as much genetically modified moss, containing the membrane bound Streptag-Luciferase-Construct, as possible. All reagents have been added to a 2 ml Tube, gently mixed and incubated over night at 4°C. A negative control containing wild type moss instead of the transgenic one was prepared, too. Before starting the incubation of the prepared samples over night, we extracted 100 µl of each one, to enable the comparison of the amount of the cut-off construct at the beginning of the incubation time and the incubation over night.
Second Step
In the second step both samples, incubated over night, were centrifuged at 10000 rpm for 5 minutes to make the moss cells form a pellet at the bottom of the tube. Afterwards the supernatants, where the Nanoluciferase-Streptag-Construct was expected to be found, were transferred to new 2 ml tubes.
Third step
Afterwards we measured the Nanoluciferase activity in the supernatants of the transgenic moss at the beginning of the incubation time and over night. The same procedure was performed for the Wild-type moss. Additionally we measured the activity of the TEV-Protease buffer in the Luciferase-Assay, to be sure that it does not falsify our results. All samples were prepared with 50 µl supernatant mixed to 50 µl Luciferase-Substrate-Buffer. We concluded that the Luciferase activity was significant in our membrane-construct supernatants, whereas the wild-type moss did not show any significant Luciferase activity.
| Supernatant transgenic moss - T0 | Supernatant transgenic moss - Over night | Supernatant wild-type moss - T0 | Supernatant wild-type moss - Over night | TEV Buffer |
|---|---|---|---|---|
| 137047 | 1852775 | 5 | 20 | 3 |
Conclusion
The Luciferase-Assay showed a significant activity rate, so there has to be a significant amount of cut-off Streptag-Nanoluciferase-Construct in the transgenic supernatant. Due to the significant results of the Luciferase-Assay of the transgenic moss supernatant, the prove of the existence and the desired orientation of our Membrane-Nanoluciferase-Construct is confirmed.
5.2 Comparison of Signal Peptides for Secretion of Effector Proteins
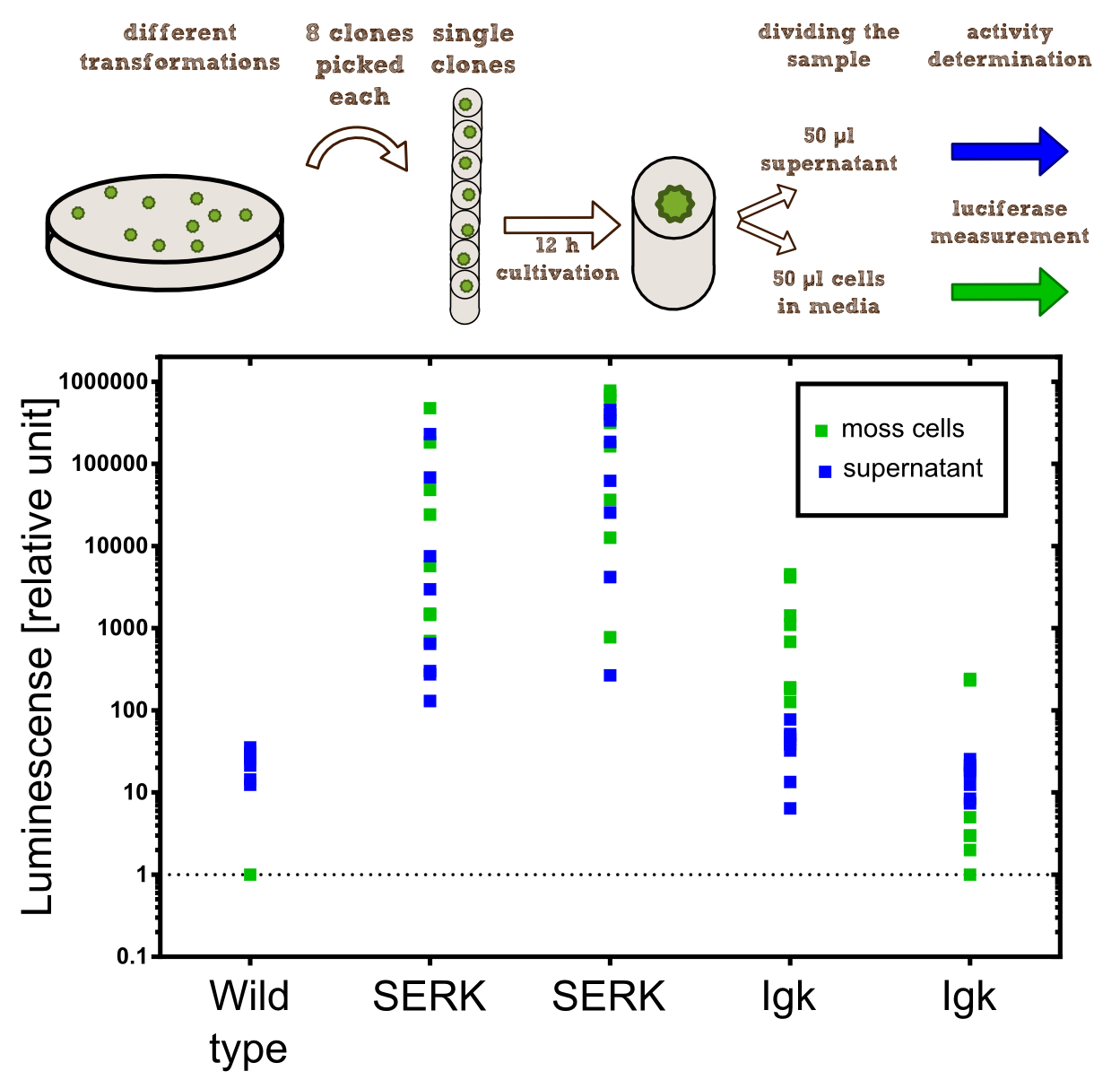
In our project the differential localisation of effector proteins was an important task which is also important for later iGEM teams who are interested in working with Physcomitrella patens. Through out the literature there are several different signal peptides which have been used to secrete proteins out of the moss cell. We decided to use two different signal peptides in our project. The first one is the Igkappa secretion signal from Mus musculus which has been recently shown to be functional as a signal peptide in Physco [http://www.ncbi.nlm.nih.gov/pubmed/19021876 Gitzinger et al., 2009]. Beside this proven signal peptide we also had the signal peptide which we derived from the SERK receptor. As it was interesting for us to know which of these signal peptides workes best we cloned both of them ahead of the NanoLuc luciferase in RFC[25]. As you can see above we cloned these genes of interest into our expression vector for stable integration. After the transfection we selected for four weeks stable transgenic moss lines using the antibiotic G418. In the end we picked 8 clones from the corresponding plates and performed an analysis of 8 single clones. This is necessary as the localisation of the transgene within the gene can have drastic effects on the expression strength of the transgene.
For the experiments 100 µl of Knop-media at pH 6.5 were transferred into 96-well plates. Subsequently single moss colonies were added to these wells and the the moss plants were incubated for 12 h in order to give the plants enough time to secrete the NanoLuc luciferase with the different signal peptides. The next day 50 µl of the supernatant in which the cells were cultivated was transfered to an empty well. The NanoGlow substrate (sponsored by Promega) was diluted in Water and 50 µl of the substrate was added rapidly to the 96-well plate. The luminescense was quantified for 1 sec in a BioTek II plate reader with a filter of 460 nm.
Results: The comparison of the two signal peptides gave a very clear result in our experiment. The Igkappa signal sequence only gives a luminescense signal which is slightly higher than the signal obtained for the wild type plants for which the autoluminescense of the substrate is responsible for the signal. In contrast the SERK signal peptide yielded luminescense signals in the supernatant that were about 1000-fold higher compared to the Igkappa signal sequence. This is also the proof that we successfully integrated a device encoding a secreted luciferase which is efficiently secreted by the moss into the supernatant. This is one of the major obstacles we had to solve in order to be able to secrete effector proteins by our PhyscoFilter.
5.3 The PhyscoFilter for Erythromycin
In order to check the activity of the erythromycine esterase B we pursued two different approaches: on the one hand we produced the enzyme recombinant in Escherichia coli and purified it. The purified protein was used for in vitro activity tests determining the depletion rates. The reaction was stopped at predetermined times by addition of three volumes acetonitrile and freezing in liquid nitrogen. On the other hand we added the substrate erythromycin at a concentration of 0.2 g L-1 to 3 mL of a polyclonal suspension of transgenic moss plants expressing the erythromycin esterase B in the cytoplasm. After 24 h of incubation at normal growth conditions 100 µL of the supernatant were used for further analysis.
Both samples were analyzed by liquid chromatography coupled to mass spectrometry (UPLC/MS/MS). The samples were measured at a Waters Xevo TQ-S using a Waters Acquity BEH C18 2,1 x 100mm column (1.7 µm, 130 Å pore size) and the following method: Solvent A: 0.1 % formic acid in water, B: 0.1 % formic acid in ACN and a linear gradient of 99 % A 0.5 min isocratic to 99 % B in 6 min. Column temperature 40 °C, flux 0.4 mL min-1, maximal pressure 703 bar. The tuning was executed with solutions consisting of the pure educt at concentrations of approximately 0.1 mgL-1.
In part A of figure seven the liquid chromatography chromatogram and the associated molecular mass for pure erythromycin as a standard is shown highlighted in red at a retention time of 3.65 min and a mass of ~734.3 g mol-1.
Part B of figure seven shows the corresponding LCMS spectra of purified erythromycin esterase B incubated for 0 min and 5 min with erythromycin. After 0 min of incubation the red highlighted peak with a retention time of 3.68 min and a mass of ~734.3 g mol-1 shows the initial presence of the substrate. However even merely mixing the substrate and enzyme was sufficient to degrade a small amount of substrate which is shown by the emerging peaks at 4.19 and 4.30 min retention time. The chromatogram of the sample taken after five minutes of incubation no longer shows the characteristic substrate peak at 3.68 min indicating that the whole amount of substrate was successfully degraded by the purified enzyme.
Part C of figure seven shows the corresponding LCMS spectra of the in vivo degradation experiment with transgenic moss expressing erythromycin esterase B in the cytoplasm. After 24 h of incubation at normal growth conditions the chromatogram of the sample containing the wild type moss as a negative control exclusively shows the characteristic peak for the substrate highlighted in red. The chromatogram of the sample in which the transgenic moss was utilized in contrast shows the clear degradation of the substrate because only the peaks at retention times of 4.19 and 4.30 min appear representing the degradation product as explained for part B of this figure.
To summarize this experiment demonstrates that our transgenic moss successfully degraded the environmental pollutant erythromycin!
5.4 The PhyscoFilter for Diclofenac and Ethinylestradiol
5.5 The PhyscoFilter for Microcstin
(receptor bound BioAccumulation)
5.6 The PhyscoFilter for Fluorescein
(receptor bound BioAccumulation using <partinfo>BBa_K1159003</partinfo>)
References:
- http://www.plant-biotech.net/paper/CurrGenet_2003_hohe.pdf Hohe, A., T. Egener, J. Lucht, H. Holtorf, C. Reinhard, G. Schween, R. Reski (2004): An improved and highly standardised transformation procedure allows efficient production of single and multiple targeted gene knockouts in a moss, Physcomitrella patens. Current Genet. 44, 339-347.
 "
"






AutoAnnotator:
Follow us:
Address:
iGEM Team TU-Munich
Emil-Erlenmeyer-Forum 5
85354 Freising, Germany
Email: igem@wzw.tum.de
Phone: +49 8161 71-4351