|
|
| (30 intermediate revisions not shown) |
| Line 22: |
Line 22: |
| | /*Add favicon*/ $("link[rel='shortcut icon']").remove(); $("head").append("<link rel='shortcut icon' type='image/png' href='https://static.igem.org/mediawiki/2013/6/6d/Uppsalas-cow-con.png'>"); }); | | /*Add favicon*/ $("link[rel='shortcut icon']").remove(); $("head").append("<link rel='shortcut icon' type='image/png' href='https://static.igem.org/mediawiki/2013/6/6d/Uppsalas-cow-con.png'>"); }); |
| | </script> | | </script> |
| | + | |
| | + | <link href="https://2013.igem.org/Team:Uppsala/lightbox-css-code.css?action=raw&ctype=text/css" type="text/css" rel="stylesheet"> |
| | + | |
| | + | <script src="https://2013.igem.org/Team:Uppsala/jquery-code.js?action=raw&ctype=text/javascript" type="text/javascript"></script> |
| | + | |
| | + | <script src="https://2013.igem.org/Team:Uppsala/lightbox-code.js?action=raw&ctype=text/javascript" type="text/javascript"></script> |
| | | | |
| | </head> | | </head> |
| Line 68: |
Line 74: |
| | <li><a href="https://2013.igem.org/Team:Uppsala/metabolic-engineering">Metabolic engineering</a> | | <li><a href="https://2013.igem.org/Team:Uppsala/metabolic-engineering">Metabolic engineering</a> |
| | <ul> | | <ul> |
| - | <li><a href="https://2013.igem.org/Team:Uppsala/p-coumaric-acid">P-coumaric acid</a></li> | + | <li><a href="https://2013.igem.org/Team:Uppsala/p-coumaric-acid">p-Coumaric acid</a></li> |
| | <li><a href="https://2013.igem.org/Team:Uppsala/resveratrol">Resveratrol</a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/resveratrol">Resveratrol</a></li> |
| | <li><a href="https://2013.igem.org/Team:Uppsala/lycopene">Lycopene</a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/lycopene">Lycopene</a></li> |
| Line 89: |
Line 95: |
| | <li><a href="https://2013.igem.org/Team:Uppsala/modeling" id="list_type1"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/6/63/Uppsala2013_Modeling.png"></a> | | <li><a href="https://2013.igem.org/Team:Uppsala/modeling" id="list_type1"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/6/63/Uppsala2013_Modeling.png"></a> |
| | <ul> | | <ul> |
| - | <li><a href="https://2013.igem.org/Team:Uppsala/P-Coumaric-acid-pathway">P-Coumaric acid</a></li> | + | <li><a href="https://2013.igem.org/Team:Uppsala/P-Coumaric-acid-pathway">Kinetic model</a></li> |
| | <li><a href="https://2013.igem.org/Team:Uppsala/modeling-tutorial">Modeling tutorial </a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/modeling-tutorial">Modeling tutorial </a></li> |
| | + | |
| | + | <li><a href="https://2013.igem.org/Team:Uppsala/toxicity-model">Toxicity model</a></li> |
| | </ul></li> | | </ul></li> |
| | <li><a href="https://2013.igem.org/Team:Uppsala/parts" id="list_type2"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/e/eb/Uppsala2013_parts.png"></a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/parts" id="list_type2"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/e/eb/Uppsala2013_parts.png"></a></li> |
| Line 109: |
Line 117: |
| | <li><a href="https://2013.igem.org/Team:Uppsala/public-opinion">Public opinion </a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/public-opinion">Public opinion </a></li> |
| | <li><a href="https://2013.igem.org/Team:Uppsala/Outreach">High school & media </a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/Outreach">High school & media </a></li> |
| - | | + | <li><a href="https://2013.igem.org/Team:Uppsala/bioart">BioArt</a></li> |
| | + | <li><a href="https://2013.igem.org/Team:Uppsala/LactonutritiousWorld">A LactoWorld</a></li> |
| | + | <li><a href="https://2013.igem.org/Team:Uppsala/killswitches">Killswitches</a></li> |
| | + | <li><a href="https://2013.igem.org/Team:Uppsala/realization">Patent</a></li> |
| | </ul></li> | | </ul></li> |
| | <li><a href="https://2013.igem.org/Team:Uppsala/attribution" id="list_type4"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/5/5d/Uppsala2013_Attributions.png"></a></li> | | <li><a href="https://2013.igem.org/Team:Uppsala/attribution" id="list_type4"><img class="nav-text" src="https://static.igem.org/mediawiki/2013/5/5d/Uppsala2013_Attributions.png"></a></li> |
| Line 129: |
Line 140: |
| | <h1>Building bridges between E. coli and Lactobacillus</h1> | | <h1>Building bridges between E. coli and Lactobacillus</h1> |
| | | | |
| - | <p>A shuttle vector is a plasmid that can be transferred between two different species and is able to replicate in both. Such a vector is key for bioengineering in Lactobacillus. The low transformation frequencies of ligations and longer generation times slows down the work pace tremendously if you have to assembly and test everything in Lactobacillus. A solution to this problem is if you instead build everything in E.coli and then transfer the finished construct to Lactobacillus. We have created two shuttle vectors that we have sent to the registry.<br><br> | + | <p>A shuttle vector is a plasmid that can be transferred between two different species and is able to replicate in both. Such a vector is key for bioengineering in Lactobacillus. The low transformation frequencies of ligations and longer generation times slows down the work pace tremendously if you have to assembly and test everything in Lactobacillus. A solution to this problem is if you instead build everything in E. coli and then transfer the finished construct to Lactobacillus. We have created two shuttle vectors that we have sent to the registry.</p><br> |
| | | | |
| - | Resveratrol belongs to a group of molecules known as Phyloalexin which is used by many plants to battle infections of all sorts.
| |
| - | Since the early 90’s there has been many studies done on resveratrol showing it has a wide range of beneficial properties ranging from skin cancer reduction to anti-inflamatory effects and antioxidant properties.<br><br>
| |
| | | | |
| | <h3>Shuttle vectors</h3> | | <h3>Shuttle vectors</h3> |
| Line 142: |
Line 151: |
| | </div> | | </div> |
| | <div class="text"> | | <div class="text"> |
| - | <a id="b1"></a><h1>Construction of shuttle vectorns</h1> | + | <a id="b1"></a><h1>Construction of shuttle vectors</h1> |
| | <p> | | <p> |
| - | To create the shuttle vector we replaced the replicon in a biobrick compatible plasmid BBa_K864003?) with a replicon from a plasmid known to work in both E.coli and Lactobacillus pSH71. pSH71 originates from a plasmid, pJP059, from Lactococcus lactis but it is well known to replicate in both the Lactobacillus genus and in E.coli through rolling circle replication.<sup><a href="#ref_point">[1]</a></sup><br><br> | + | To create the shuttle vector we replaced the replicon in a biobrick compatible plasmid BBa_K864003 with a replicon, pSH71, from a plasmid known to work in both E. coli and Lactobacillus. pSH71 originates from a plasmid, pJP059, from Lactococcus lactis but it is well known to replicate in both the Lactobacillus genus and in E. coli through rolling circle replication.<sup><a href="#ref_point">[1]</a></sup><br><br> |
| | | | |
| - | Many species in the Lactobacillus genus have inherent antibiotic resistances.<sup><a href="#ref_point">[2]</a></sup>, because of that we have been limited in our selection of antibiotic resistance. Most commonly used are chloramphenicol and erythromycin and we made versions of both for our shuttle vector. BBa_K1033207 contains an erythromycin cassette from pLUL631 and BBa_K1033206 contains the resistance cassette from pSB1C3 but has had its promoter replaced with our cp29 promoter.<sup><a href="#ref_point">[3]</a></sup><br><br> </p> | + | Many species in the Lactobacillus genus have inherent antibiotic resistances.<sup><a href="#ref_point2">[2]</a></sup>, because of that we have been limited in our selection of antibiotic resistance. Most commonly used are chloramphenicol and erythromycin and we made versions of both for our shuttle vector. BBa_K1033207 contains an erythromycin cassette from pLUL631 and BBa_K1033206 contains the resistance cassette from pSB1C3 but has had its promoter replaced with our cp29 promoter.<sup><a href="#ref_point3">[3]</a></sup><br><br> </p> |
| | </div> | | </div> |
| - | <img class="vector-plasmid" src="https://static.igem.org/mediawiki/2013/9/92/Shuttle-vector_pSBLbC.png"> | + | <a href="https://static.igem.org/mediawiki/2013/9/92/Shuttle-vector_pSBLbC.png" data-lightbox="roadtrip"><img class="vector-plasmid" src="https://static.igem.org/mediawiki/2013/9/92/Shuttle-vector_pSBLbC.png"></a> |
| | | | |
| | <h1>Results</h1> | | <h1>Results</h1> |
| | | | |
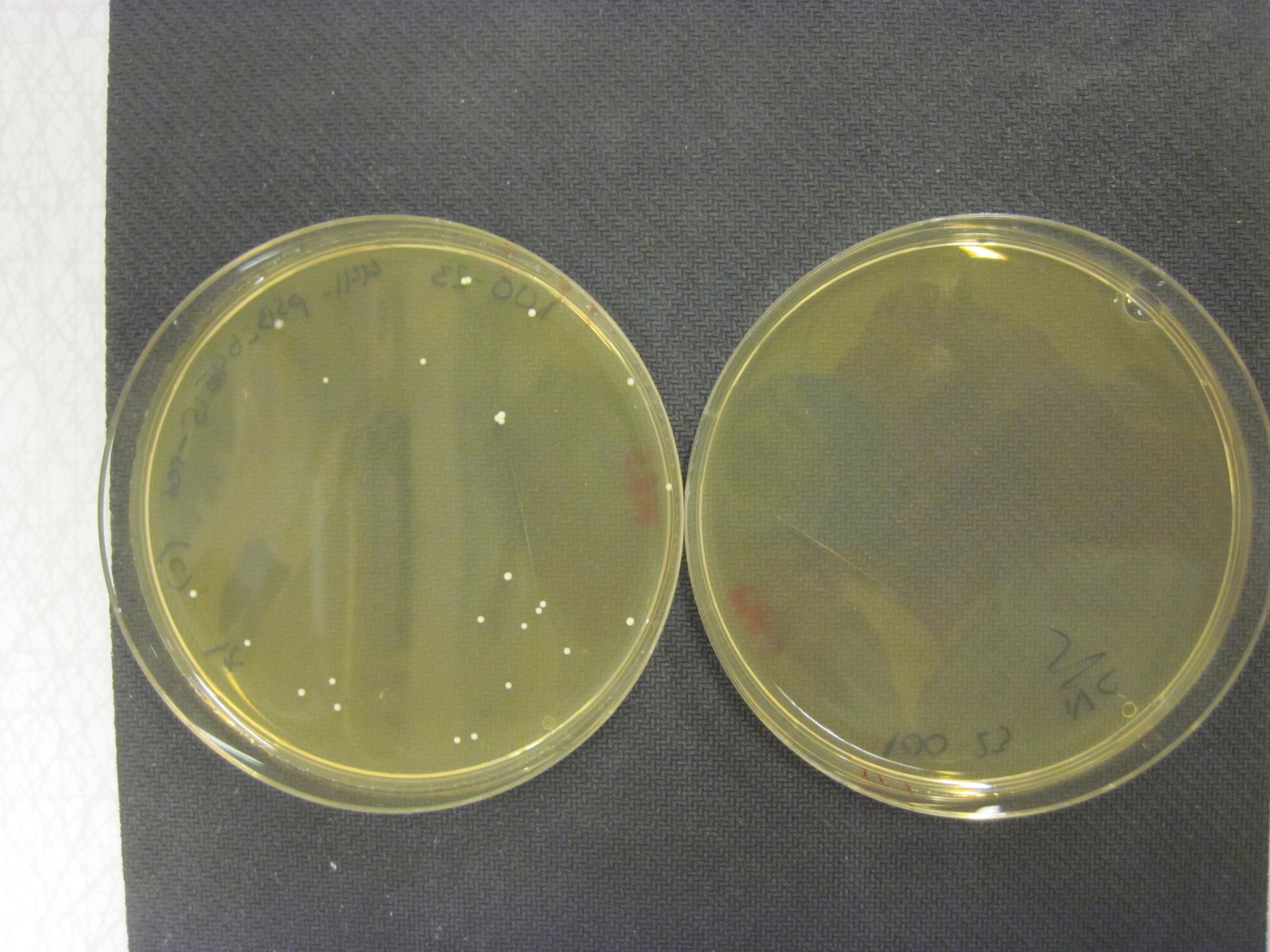
| - | <p>We have managed to transform both shuttle vectors to E. coli and verified them by sequencing. We have also managed to subclone with them and replaced the red insert with chromoproteins expressed by our CP promoters that works both in E. coli and Lactobacillus, clearly yielding coloured colonies (see picture on the right below).</p> | + | <p>We have managed to transform both shuttle vectors to E. coli and verified them by sequencing. We have also managed to subclone with them and replaced the red insert with chromoproteins expressed by our CP promoters that works both in E. coli and Lactobacillus, clearly yielding blue colonies.</p> |
| | | | |
| - | <img class="pic_type2" src="https://static.igem.org/mediawiki/2013/e/e1/IMG_6644.JPG"> | + | <div id="pic_type2"> |
| | + | <a href="https://static.igem.org/mediawiki/2013/e/e1/IMG_6644.JPG" data-lightbox="roadtrip" title="BBa_K1033207 transformed to Lactobacillus reuteri 100-23"><img class="pic_type2" src="https://static.igem.org/mediawiki/2013/e/e1/IMG_6644.JPG"></a> |
| | | | |
| - | <img class="pic_type3" src="https://static.igem.org/mediawiki/2013/2/28/Uppsala2013_chromo.JPG"> | + | <i>To the left, BBa_K1033207 transformed to Lactobacillus reuteri 100-23, to the right, negative control without plasmid</i> |
| | + | </div> |
| | | | |
| - | <img class="shuttle_vec" src="https://static.igem.org/mediawiki/2013/1/18/Uppsala2013_Shuttle_Vector_pSBLBC_cp29_amilCP1.png"> | + | <div id="pic_type3"> |
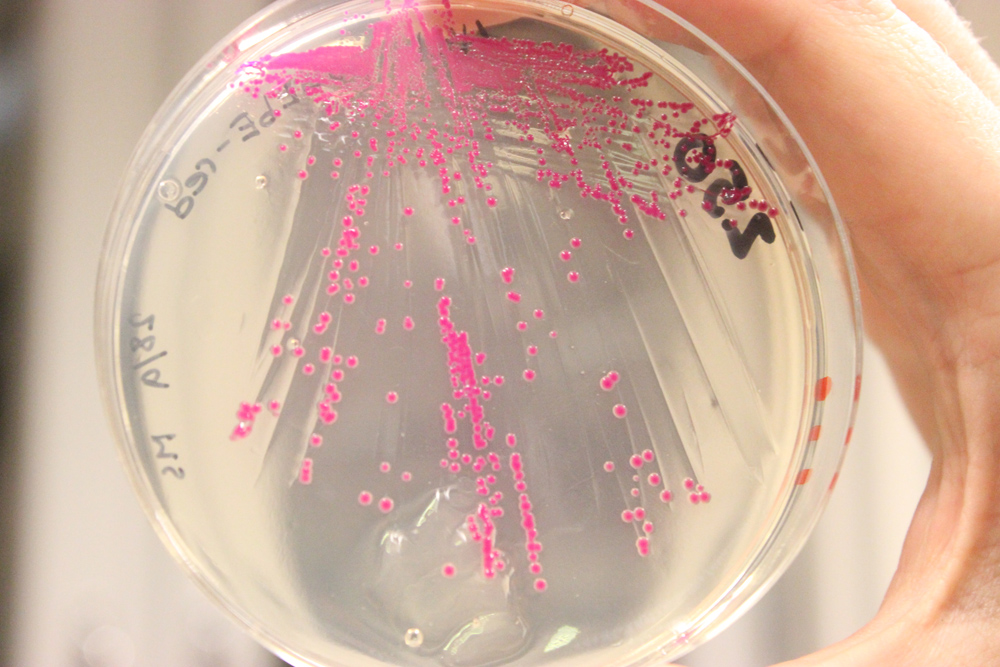
| | + | <a href="https://static.igem.org/mediawiki/2013/2/28/Uppsala2013_chromo.JPG" data-lightbox="roadtrip" title="BBa_K1033207 and RFP, BBa_J04450 insert in E. coli"><img class="pic_type3" src="https://static.igem.org/mediawiki/2013/2/28/Uppsala2013_chromo.JPG"></a> |
| | + | |
| | + | <i class="middle-text">BBa_K1033207 and RFP, BBa_J04450 |
| | + | <br> |
| | + | <i class="middle-text">insert in E. coli</i> |
| | + | </div> |
| | + | |
| | + | <div id="pic_type4"> |
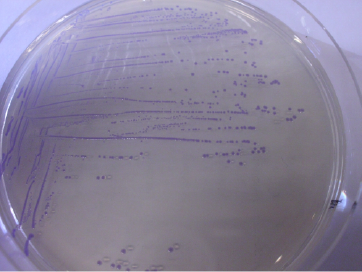
| | + | <a href="https://static.igem.org/mediawiki/2013/b/ba/Uppsala2013_Shuttle_Vector_pSBLBC_cp29_amilCP.JPG" data-lightbox="roadtrip" title="BBa_K1033206 with a subcloned chromoprotein BBa_K1033282 in E. coli"><img class="shuttle_vec" src="https://static.igem.org/mediawiki/2013/1/18/Uppsala2013_Shuttle_Vector_pSBLBC_cp29_amilCP1.png"></a> |
| | + | |
| | + | <i>BBa_K1033206 with a subcloned chromoprotein BBa_K1033282 in E. coli</i> |
| | + | </div> |
| | | | |
| | <div id="clear"></div> | | <div id="clear"></div> |
| | | | |
| - | <p>Finally we have also gained positive results when transforming BBa_K1033207 to Lactobacillus reuteri and Lactobacillus plantarum. See the picture above on the left. The plate on the left contains Lactobacillus reuteri 100-23 transformed with BBa_K1033207. The plate on the right is a negative control of 100-23 without the plasmid. <p> | + | <p>We have also gained positive results when transforming BBa_K1033207 to Lactobacillus reuteri and Lactobacillus plantarum. <p> |
| | | | |
| | <div id="reference"> | | <div id="reference"> |
| Line 168: |
Line 191: |
| | <p class="reference"> | | <p class="reference"> |
| | | | |
| - | <a>[1]</a> <a href:"http://www.ncbi.nlm.nih.gov/pubmed/11591136> "Construction of compatible wide-host-range shuttle v... </a> [Plasmid. 2001] - PubMed - NCBI." National Center for Biotechnology Information. N.p., n.d. Web. 1 Oct. 2013. | + | <a id="ref_point">[1]</a> <a href="http://www.ncbi.nlm.nih.gov/pubmed/11591136"> Construction of compatible wide-host-range shuttle v... </a> [Plasmid. 2001] - PubMed - NCBI." National Center for Biotechnology Information. N.p., n.d. Web. 1 Oct. 2013. |
| | | | |
| | <br><br> | | <br><br> |
| | | | |
| - | <a>[2]</a> <a href:"http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3378904> "High efficiency recombineering in lactic acid bacteria." </a> National Center for Biotechnology Information. N.p., n.d. Web. 1 Oct. 2013. | + | <a id="ref_point2">[2]</a> <a href="http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3378904"> "High efficiency recombineering in lactic acid bacteria. </a> National Center for Biotechnology Information. N.p., n.d. Web. 1 Oct. 2013. |
| | <br><br> | | <br><br> |
| | | | |
| - | <a>[3]</a> <a href="http://www.researchgate.net/publication/226939823_Transformation_ofLactobacillus_reuteri_with_electroporation_Studies_on_the_erythromycin_resistance_plasmid_pLUL631?ev=pub_cit%22,%22http://www.researchgate.net/publication/226939823_Transformation_ofLactobacillus_reuteri_with_electroporation_Studies_on_the_erythromycin_resistance_plasmid_pLUL631?ev=pub_cit%22) | + | <a id="ref_point3">[3]</a> <a href="http://www.researchgate.net/publication/226939823_Transformation_ofLactobacillus_reuteri_with_electroporation_Studies_on_the_erythromycin_resistance_plasmid_pLUL631?ev=pub_cit%22,%22http://www.researchgate.net/publication/226939823_Transformation_ofLactobacillus_reuteri_with_electroporation_Studies_on_the_erythromycin_resistance_plasmid_pLUL631?ev=pub_cit%22) |
| | "> Transformation of Lactobacillus reuteri with electroporation: Studies on the erythromycin resistance plasmid pLUL631 </a> , Siv Ahrné, Göran Molin, Lars Axelsson, Januari 1992 | | "> Transformation of Lactobacillus reuteri with electroporation: Studies on the erythromycin resistance plasmid pLUL631 </a> , Siv Ahrné, Göran Molin, Lars Axelsson, Januari 1992 |
| | | | |
 "
"