Team:Bielefeld-Germany/Project/Riboflavine
From 2013.igem.org
Riboflavin
Overview

Tubes from left to right:
- Tube 1: A 72-hour culture of E. coli KRX [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1172306 BBa_K1172306]
- Tube 2: A 72-hour culture of E. coli KRX [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1172306 BBa_K1172306], supernatant
- Tube 3: A 72-hour culture of E. coli KRX [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1172306 BBa_K1172306], cell lysate
- Tube 4: A 72-hour culture of a wild type E. coli KRX, supernatant
- Tube 5: A 72-hour culture of a wild type E. coli KRX, cell lysate
Riboflavin, or Vitamin B2 is a redox-active substance that plays an essential role in living cells. As precursor of flavin mononucleotide (FMN) and flavin adenine dinucleotide (FAD) it is crucial for diverse energy supplying metabolic processes, e.g. beta-oxidation or oxidative phosphorylation. Secreted into the medium, it can be
effectively used by some bacteria for electron transfer. Presence of riboflavin in anaerobic cultures leads to higher current flow in a Microbial Fuel Cell, which made riboflavin overproduction a suitable target for optimisation of our MFC.
We have shown that cloning of the riboflavin cluster from the metal-reducing bacterium Shewanella oneidensis MR-1 in E. coli is sufficient to achieve significant riboflavin overproduction detectable both in supernatant and in cells.
Theory
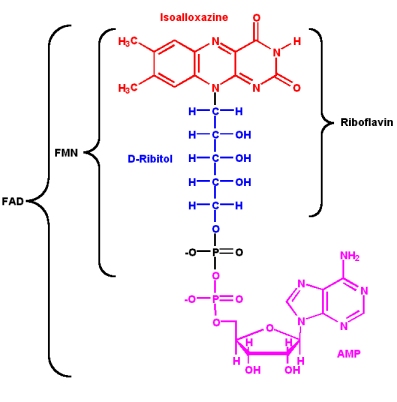
Since its discovery in 1879 and its first structural characterization in the 1930’s, a lot of properties of riboflavin were elucidated. This substance is a precursor of FMN (flavin mononucleotide) and FAD (flavin adenine dinucleotide), which play an essential role as cofactors in many oxidative processes.
The modern name riboflavin, also named lactoflavin, is composed of two parts: «ribo» indicating the presence of the sugar alcohol ribitol, and «flavin» meaning «yellow»; to accentuate the yellow coloring of the oxidized molecule. Chemically this substance consists of two functional subunits, an already mentioned short-chain ribitol and a tricyclic heterosubstituted isoalloxazine ring.
The latter, also known as a riboflavin ring, exists in three redox states and is responsible for the diverse chemical activity of riboflavin. A fully oxidized quinone, a one-electron semiquinone and a fully reduced hydroquinone are the three states of riboflavin oxidation. In an aqueous solution, the quinone (fully oxidized) form of riboflavin has a typical yellow coloring. It becomes red in a semi-reduced anionic or blue in a neutral form and is colorless when fully reduced.
All these forms are present in different proportions in a living cell, making previous oxidation a necessary step if riboflavin analysis is to be conducted. Flavins have a typical yellow-green fluorescence in the UV light. The peaks of absorbance occur at 223, 266, 373 and 445 nm. The maximum fluorescence emission of the neutral solution is at 535 nm [Charles A. Abbas et al., [http://mmbr.asm.org/content/75/2/321.full#ref-292 2011]]. These fluorimetric properties are widely used in the analysis of riboflavin.
Due to its structure, which allows a transfer of two electrons from hydrogen and hydrid ions, riboflavin can be imagined as a potential electron shuttle. It was previously known, that the electron transfer from the outer membrane-associated proteins to an inorganic electron acceptor is the main limiting growth factor for Fe(III)-reducing prokaryotes, so a few mechanisms that enhance this process have been discovered. One of them was a secretion of water-soluble redox mediators. It was proven, that secretion of riboflavin and FMN enhances the rate of insoluble mineral oxides reduction. Indeed, Shewanella oneidensis, a facultative Fe-III respiring bacterium uses secreted riboflavin as its electron transmitter [Harald von Canstein et al., [http://aem.asm.org/content/74/3/615.full 2008]]. Therefore, we decided to overproduce riboflavin in E. coli to improve its efficiency in our MFC.
First of all, we searched for a suitable microorganism, which has an active riboflavin cluster with known coding sequences. We have already used the remarkable versatility of Shewanella in order to clone the anaerobic respiratory chain (mtrCAB cluster), so now we were able to skip some initial steps, like the whole genome DNA isolation and advanced rapidly to more specific steps.
Before planning the cloning strategy, we checked [http://parts.igem.org/Main_Page the Parts registry] for any parts we could use or enhance. A [http://parts.igem.org/Part:BBa_K769203 ribC part] was listed, but it was neither submitted nor available.
We also had a close look on the [http://www.chem.qmul.ac.uk/iubmb/enzyme/reaction/misc/riboflavin2.html riboflavin biosynthesis pathway]. This process is well studied and a lot of appropriate literature is available. There are three types of riboflavin overproducers used in industry: yeast (Candida famate), fungi (Ashbya gossypii), and bacteria (Bacillus subtilis) [Overview, Seong Han Lim et al., [http://www.bbe.or.kr/storage/journal/BBE/6_2/6657/articlefile/article.pdf 2001]], but a prevailing majority of microorganisms also synthesize riboflavins in low concentrations. In E. coli, for instance, genes are scattered through the whole genome and riboflavin is produced constitutively.
The biosynthesis of riboflavin starts with ribulose-5-phosphate and GTP, converted to formate and DHBP (L-3,4-dihydroxybutan-2-one 4-phosphate). The final stage involves forming of the third isoalloxazine ring by an exchange of a 4-carbon part, catalysed by riboflavin synthetase (EC 2.5.1.9). We assumed, that the operon in Shewanella could be similar to a well-studied [http://www.ncbi.nlm.nih.gov/pubmed/8159171 rib-operon] of Bacillus subtilis. The operon is transcribed as a one polycistronic RNA, making a single promoter sufficient. Introduction of multiple copies of a ribA gene (coding for GTP cyclohydrolase II (EC 3.5.4.25)), comprised in the rib-operon, results in riboflavin overproduction in B. subtilis [Hohmann H.P.,Stahmann K.P. 2010. Biotechnology of riboflavin production, p. 115–139.], so we predicted a notable riboflavin synthesis gain following a successful introduction of a multiple-copy plasmid harbouring the rib-operon under an active promoter.
Genetic Approach
Riboflavin Cluster

Below we shortly describe each functional member of this cluster.
- Gene: ribD http://www.ncbi.nlm.nih.gov/gene/1171145 Sequence, 1145 bp
- Protein: bifunctional diaminohydroxyphosphoribosylaminopyrimidine deaminase/5-amino-6-(5-phosphoribosylamino) uracil reductase RibD
- Enzyme: (EC: 3.5.4.26)
- Molecular weight: 41,257 kDa
- Gene: SO_3468 http://www.ncbi.nlm.nih.gov/gene/1171144 Sequence, 656 bp
- Protein: Riboflavin synthase alpha subunit RibC-like protein
- Enzyme: EC 2.5.1.9
- Molecular weight: 23,483 kDa
- Gene: ribBA → ribA & ribB http://www.ncbi.nlm.nih.gov/gene/1171143 Sequence, 1103 bp
- ribA
- Protein: GTP cyclohydrolase-2
- Enzyme: EC 3.5.4.25
- Molekular weight: 22,852 kDa
- ribB
- Protein: 3,4-dihydroxy-2-butanone-4-phosphate synthase
- Enzyme: EC 3.5.4.25
- Molecular weight: 22,956 kDa
- Gene: ribE http://www.ncbi.nlm.nih.gov/gene/1171142 Sequence, 476 bp
- Protein: 6,7-dimethyl-8-ribityllumazine synthase (aka: riboflavin synthase beta subunit RibE)
- Enzyme: EC 2.5.1.78
- Molecular Weight: 16,689 kDa
Figure 5 presents the initial plasmid we established: The rib-gene cluster from the genome of Shewanella oneidensis cloned into pSB1C3.

The second step is shown in Figure 6: We disrupted the plasmid at the illegal restriction-sites. Thus enabling the deletion of the sites by using special designed primers for the following Gibson-Assembly.
Finally, the assemblage of the fragments lead to a functioning BioBrick shown in Figure 7.
Supplementary Genes
We also separately amplified some other genes that theoretically could be important for riboflavin synthesis or regulation.
- Gene: ribC http://www.ncbi.nlm.nih.gov/gene/1170021 Sequence, 621 bp
- Protein: 6,7-dimethyl-8-ribityllumazine synthase alpha subunit RibC, analogous to EC 2.5.1.9
- Molecular Weight: 22,110 Da
- Gene: nusB (full name: N utilization substance protein B homolog), http://www.ncbi.nlm.nih.gov/gene/1171141 Sequence, 403 bp
- Protein: transcription antitermination factor
- Molecular weight: 14,740 Da
- Gene: norM, http://www.ncbi.nlm.nih.gov/gene/1170020 Sequence, 1380 bp
- Protein: probable multidrug eflux protein NorM
- Molecular weight: 50,490 Da

Results
Confirming overexpression of the rib-gene cluster
The overexpression of <bbpart>BBa_K1172303</bbpart> and its derived devices <bbpart>BBa_K1172306</bbpart>,<bbpart>BBa_K1172305</bbpart>, <bbpart>BBa_K1172304</bbpart> is assured by verifying the protein Riboflavin synthase beta subunit RibE The protein RibE is part of the riboflavin synthesis pathway of Shewanella oneidensis. The corresponding gene is ribE. RibE belongs to the rib-gene cluster, which we managed to isolate, removing all the illegal restriction sites and subsequently cloned into pSB1C3.
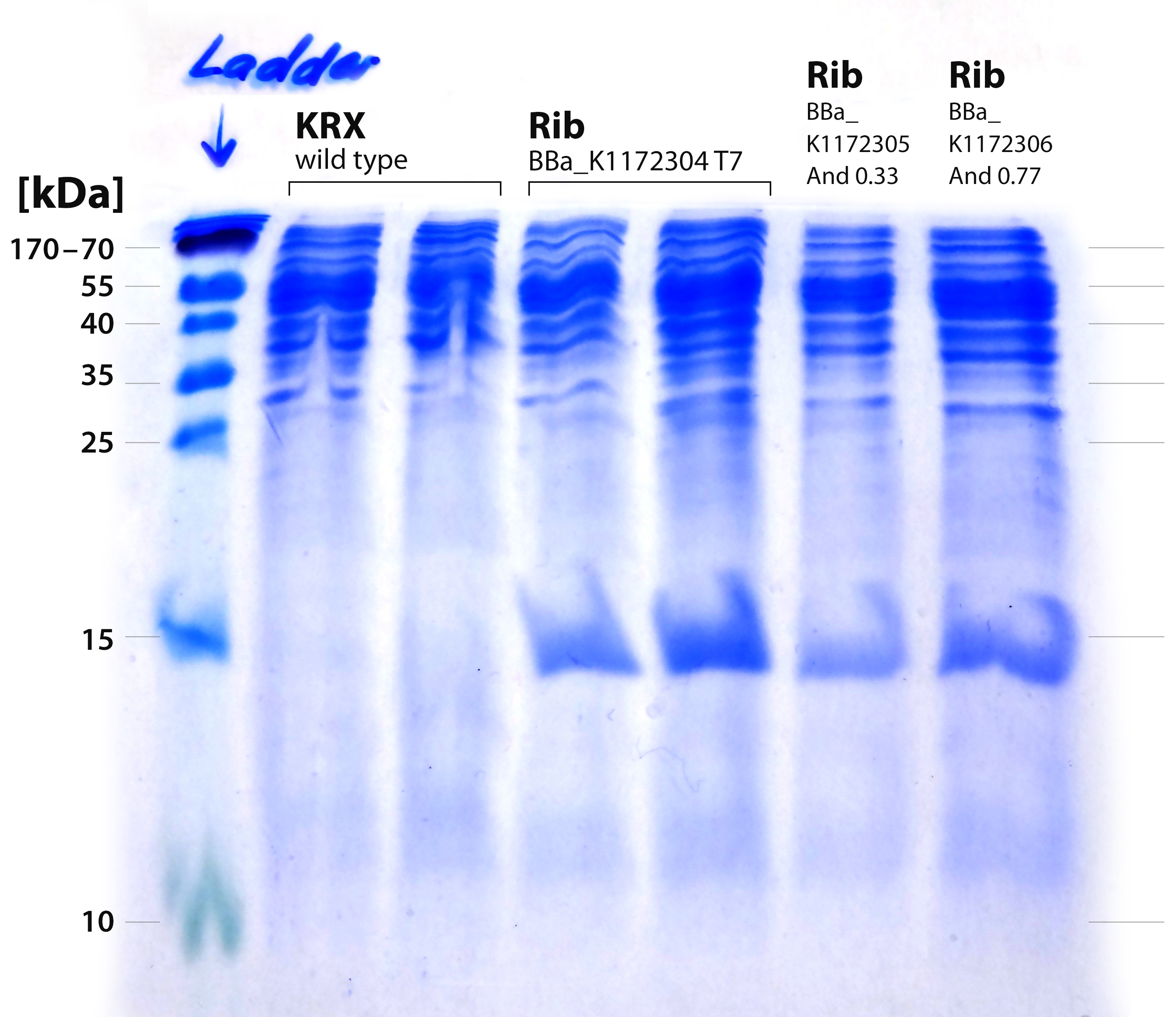
SDS-PAGE
The performed SDS-PAGE shows a distinct band at ~15 kDa. The exact size of the riboflavin synthase beta subunit RibE is 16.7 kDa. The band was cut out and analyzed by MALDI-TOF.
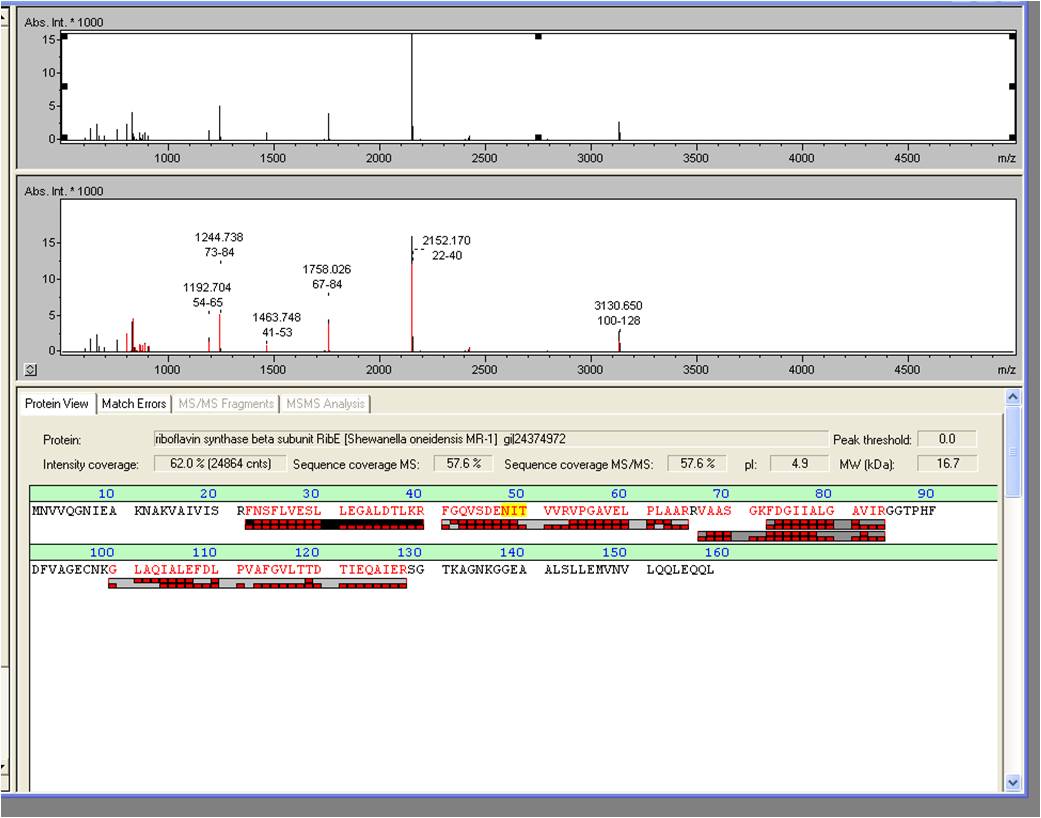
MALDI-TOF
The spot, described above, was picked and digested with trypsine. Afterwards the sample was spotted on the target and analyzed by MALDI-TOF MS/MS. RibE was identified with a Mascot Score of 906 against the NCBI database concerning bacterial organisms.
Analysis of riboflavin in supernatants
Absorbance measurement
Riboflavin has an absorption peak at 446 nm. The absorbance was measured in a TECAN infinite plate reader. The samples consisted of supernatant derived from E. coli KRX with <bbpart>BBa_K1172306</bbpart> and KRX as the "wild type" (both strains were cultivated over 72 hours). Further intracellular measurements of both strains were obtained. Therefore, the cells were disrupted via mechanical disruption, the debrisremoved by centrifugation and the supernatant was analyzed.
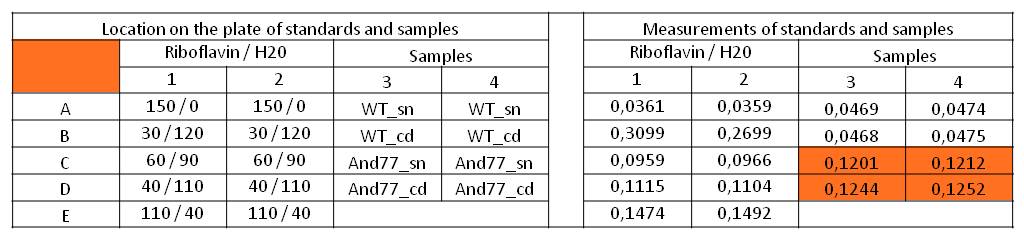
Table 1: Pipetting scheme and measurement results of riboflavin standards and cell samples for absorbance measurement at 446 nm in the [http://www.tecan.com/platform/apps/product/index.asp?MenuID=1812&ID=1916&Menu=1&Item=21.2.10.1 Tecan Infinite® M200 platereader]. WT = wild type, And77 = E. coli carrying <bbpart>BBa_K1172306</bbpart>, sn = supernatant, cd = cell disruption.
Riboflavin in known concentrations (5.31 * 10^-5 M) and dilutions was measured to generate a calibration curve. The subsequently computed riboflavin concentrations were 5773.3 µg / L for the supernatant of E. coli KRX with <bbpart>BBa_K1172306</bbpart> and 6112.63 µg /L for the cell disruption samples of E. coli KRX with <bbpart>BBa_K1172306</bbpart>. The concentration of riboflavin in the wild type strain was below the detection limit.
- Absorbance measurement is the least sensitive method used for riboflavin detection. Therefore, the slightly higher yields should be taken with a grain of salt.
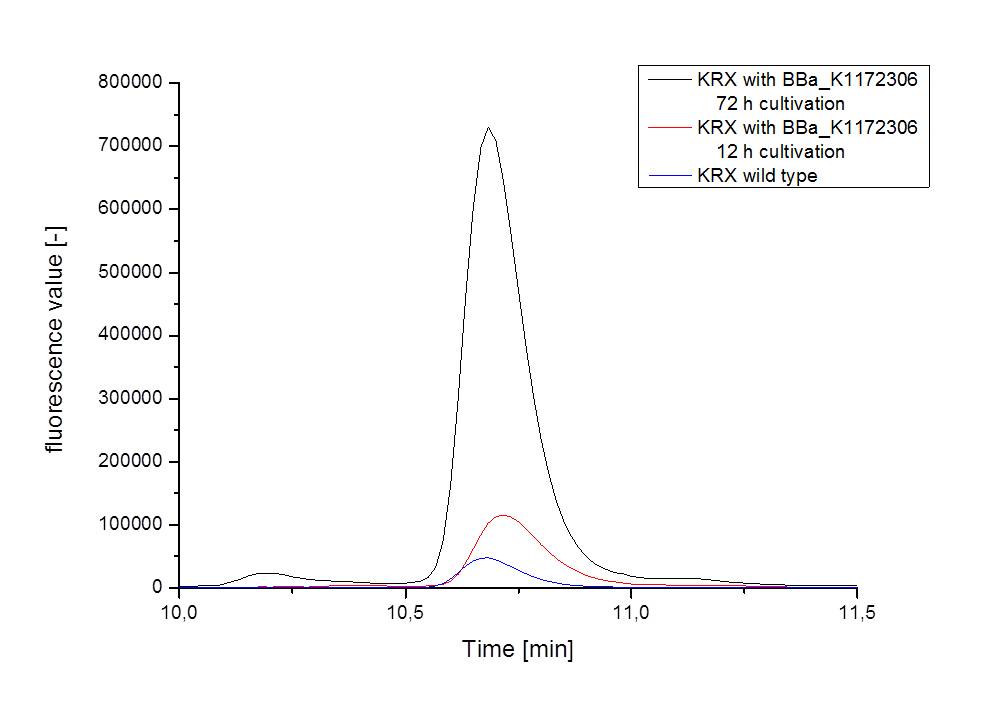
Fluorescence measurement
Riboflavin absorbs light at 440 nm with a corresponding emission at 535 nm. The fluorescence was measured in a TECAN infinite plate reader. The samples consisted of supernatant samples from E. coli KRX with <bbpart>BBa_K1172306</bbpart> (grown for 72 hours) , E. coli KRX with <bbpart>BBa_K1172306</bbpart> (grown for 12 hours) and E. coli KRX wild type bacteria (grown for 72 hours)
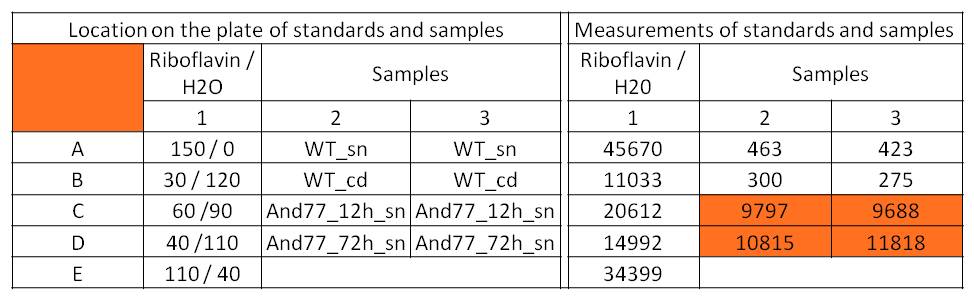
Table 2: Pipetting scheme and measurement results of riboflavin standards and cell samples for fluorescence measurement, emission at 535 nm. Measured in the [http://www.tecan.com/platform/apps/product/index.asp?MenuID=1812&ID=1916&Menu=1&Item=21.2.10.1 Tecan Infinite® M200 platereader]. WT = wild type, And77 = E. coli carrying <bbpart>BBa_K1172306</bbpart>, sn = supernatant, cd = cell disruption.
Riboflavin in known concentrations and dilutions was measured to generate a calibration curve. The subsequently calculated riboflavin concentrations were 308.1 µg / L for the supernatant sample after 12 hours and 3821.5 µg /L for the supernatant sample after 72 hours. The concentration of riboflavin in the wild type strain was below the detection limit.
HPLC measurement
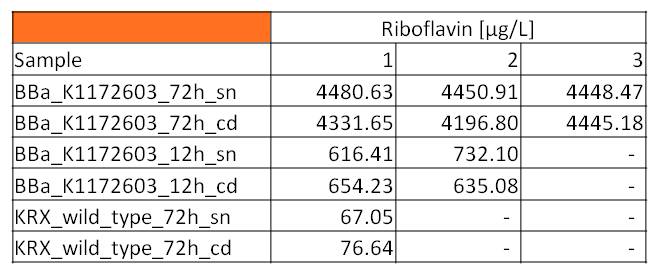
Supernatant and cell disruption samples of E. coli KRX with BBa_K1172306 (grown for 72 hours) , E. coli KRX with <bbpart>BBa_K1172306</bbpart> (grown for 12 hours) and E. coli KRX wild type bacteria (grown for 72 hours) were measured in a HPLC detector.
In summary, the obtained data showed, that after 72 hours of cultivation the concentration of riboflavin in the supernatant and cell disruption samples of E. coli KRX carrying <bbpart>BBa_K1172306</bbpart> was 60fold higher than in the E. coli KRX wild type: Approx. 4400 µg/L, yielded with BioBrick carrying KRX compared to 67.05 µg/L for the KRX wild type. Even after 12 hours, the riboflavin producing strain had already generated ten times as much riboflavin as the wild type: Approx. 700 µg/L compared to 76.64 µg/L.
- HPLC yielded the only results for riboflavin concentration in KRX wild type supernatants due to its sensitivity.
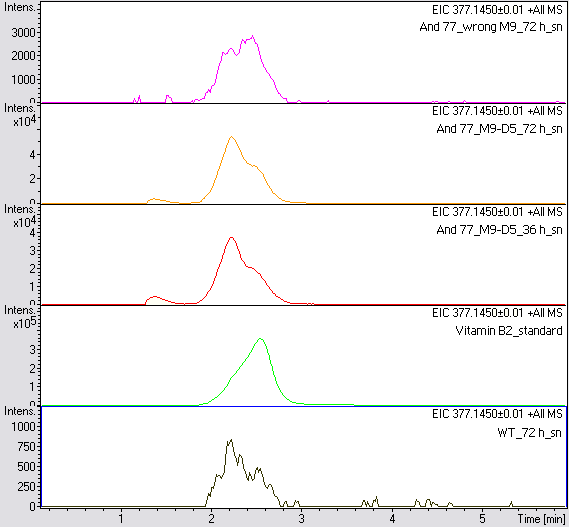
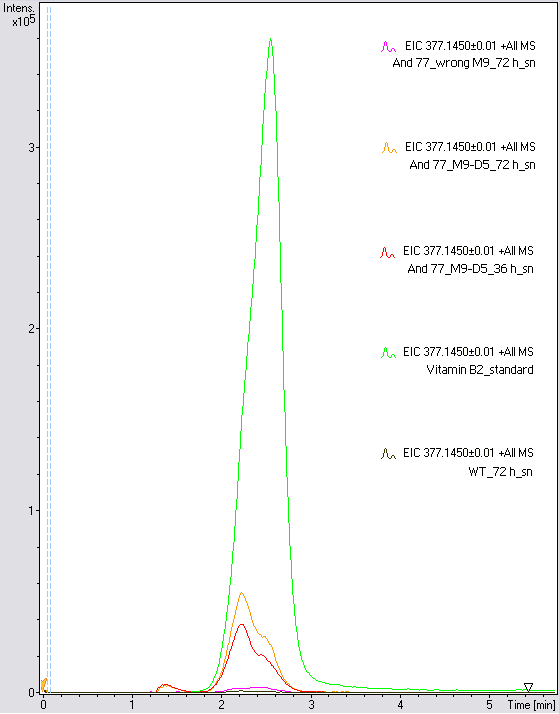
LC/MS measurement
Supernatant samples of a wild type E. coli KRX (grown for 72 hours) and E. coli KRX with BBa_K1172306 (grown for 72 hours and 36 hours) were measured. One sample of E. coli KRX with BBa_K1172306, which had been grown in a M9 medium for 72 hours was measured additionally.
The <bbpart>BBa_K1172306</bbpart> carrying strains produced a much higher amount of riboflavin compared to the wild type E. coli KRX strain. The LC/MS results do not allow for a statement on how much more riboflavin E. coli KRX carrying <bbpart>BBa_K1172306</bbpart> produces. Nevertheless, it is obvious that riboflavin was overproduced in a remarkable quantity.
Evaluation of the measurements
The quantitative data obtained using absorbance, fluorescence and HPLC measurements shows a distinct trend. All samples generated from E. coli KRX carrying <bbpart>BBa_K1172306</bbpart> (grown for 72 h) showed similar values of approx. 4000 µg/ml. This is a considerable increase in riboflavin production compared to the wild type KRX strains.
Conclusion
Riboflavin possesses the ability to be a potent redoxmediator. By turning the rib-gene cluster from Shewanella oneidensis into a BioBrick and subsequently cloning it into the desired chassis Escherichia coli, the iGEM Team Bielefeld was able to raise the amount of riboflavin produced by E. coli significantly. The results indicate that the transformation of E. coli with <bbpart>BBa_K1172303</bbpart>, respectively <bbpart>BBa_K1172306</bbpart> represents a viable option when considering genetically optimization of microorganisms intended for usage in microbial fuel cells (MFC).
References
C. A. Abbas and A. S. Sibirny. (2011) Genetic control of biosynthesis and transport of riboflavin and flavin nucleotides and construction of robust biotechnological producers. [http://mmbr.asm.org/content/75/2/321.full#ref-292| Microbiology and Molecular Biology Reviews 75(2): 321-360].
Hohmann H. P., Stahmann K. P. (2010). Biotechnology of riboflavin production, p. 115–139. In Mander L., Liu H. W. (ed.), Comprehensive natural products. II. Chemistry and biology, vol. 7. Cofactors. Elsevier, Philadelphia, PA.
von Canstein H., Ogawa J., Shimizu S., Lloyd J. R. (2008). Secretion of flavins by Shewanella species and their role in extracellular electron transfer. [http://aem.asm.org/content/74/3/615.full Appl. Environ. Microbiol. 74:615–623].
Bacher A., et al. (2001). Biosynthesis of riboflavin. [http://www.ncbi.nlm.nih.gov/pubmed/11153262 Vitam. Horm. 61:1–49.]
Tesliar G. E., Shavlovskii G. M. (1983). Localization of the genes coding for GTP cyclohydrolase II and riboflavin synthase on the chromosome of Escherichia coli K-12. Tsitol. Genet. 17:54–56. (In Russian.)
Seong Han Lim, Jong Soo Choi and Enoch Y. (2001). Park Microbial Production of Riboflavin Using Riboflavin Overproducers, Ashbya gossypii, Bacillus subtilis, and Candida famate: An Overview.[http://www.bbe.or.kr/storage/journal/BBE/6_2/6657/articlefile/article.pdf Biotechnol. Bioprocess Eng., 6: 75-88]
 "
"