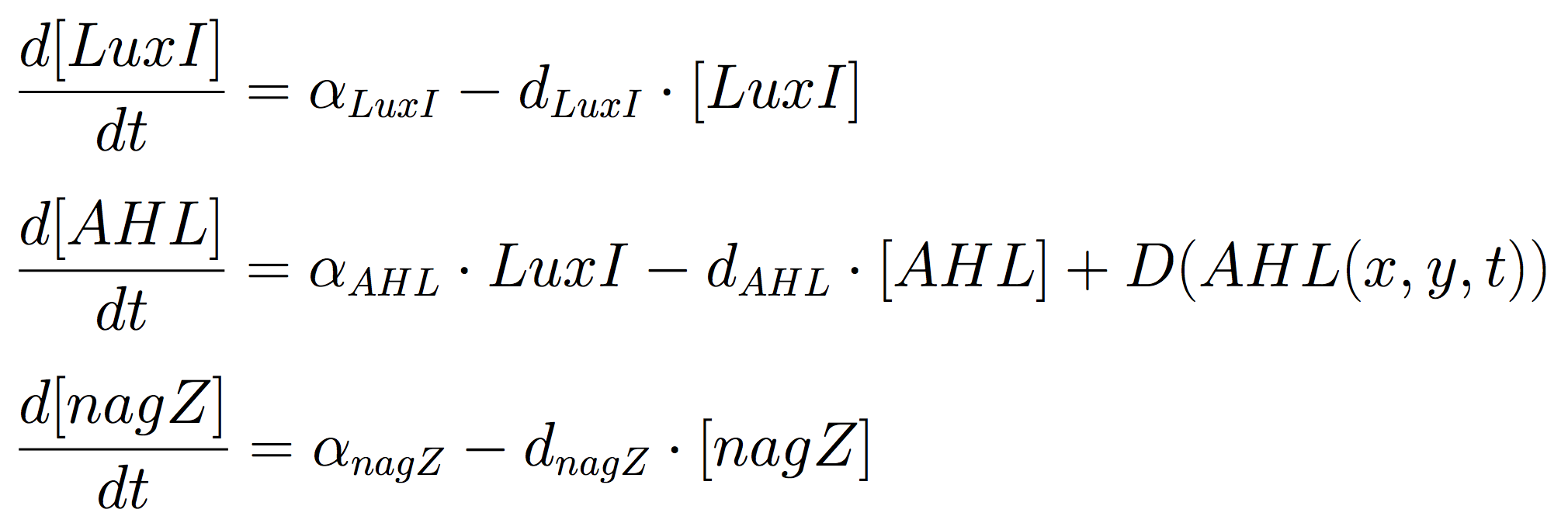Team:ETH Zurich/Modeling
From 2013.igem.org
(Difference between revisions)
| Line 24: | Line 24: | ||
=''Mine Cells''= | =''Mine Cells''= | ||
| + | [[File:eqnSender.png|500px|thumb|Equation system 1: ODE for Mine Cells.]] | ||
<p>The Mine Cells lead to the synthesis the signalling molecule, by constitutive expression of ''luxI'' gene. To reveal the nature of the cells, a coloured-substrate reaction is triggered upon addition of ''5-Bromo-4-chloro-3-indoxyl-N-acetyl-beta-D-glucosaminide''; given that the glycoside hydrolase ''NagZ'' is expressed constitutively. <br><br> The ODEs for the states involved in the sender module are given below: | <p>The Mine Cells lead to the synthesis the signalling molecule, by constitutive expression of ''luxI'' gene. To reveal the nature of the cells, a coloured-substrate reaction is triggered upon addition of ''5-Bromo-4-chloro-3-indoxyl-N-acetyl-beta-D-glucosaminide''; given that the glycoside hydrolase ''NagZ'' is expressed constitutively. <br><br> The ODEs for the states involved in the sender module are given below: | ||
</p> | </p> | ||
| - | + | ||
= ''Biosensors'' = | = ''Biosensors'' = | ||
hg | hg | ||
Revision as of 15:50, 17 August 2013
The change of the species concentrations in time is given by non-linear ordinary differential equations (ODEs), most of which follow Hill kinetics. The parameters we used in the model are derived from literature.
Mine Cells
The Mine Cells lead to the synthesis the signalling molecule, by constitutive expression of luxI gene. To reveal the nature of the cells, a coloured-substrate reaction is triggered upon addition of 5-Bromo-4-chloro-3-indoxyl-N-acetyl-beta-D-glucosaminide; given that the glycoside hydrolase NagZ is expressed constitutively.
The ODEs for the states involved in the sender module are given below:
 "
"