Team:ETH Zurich/Experiments 3
From 2013.igem.org
| Line 123: | Line 123: | ||
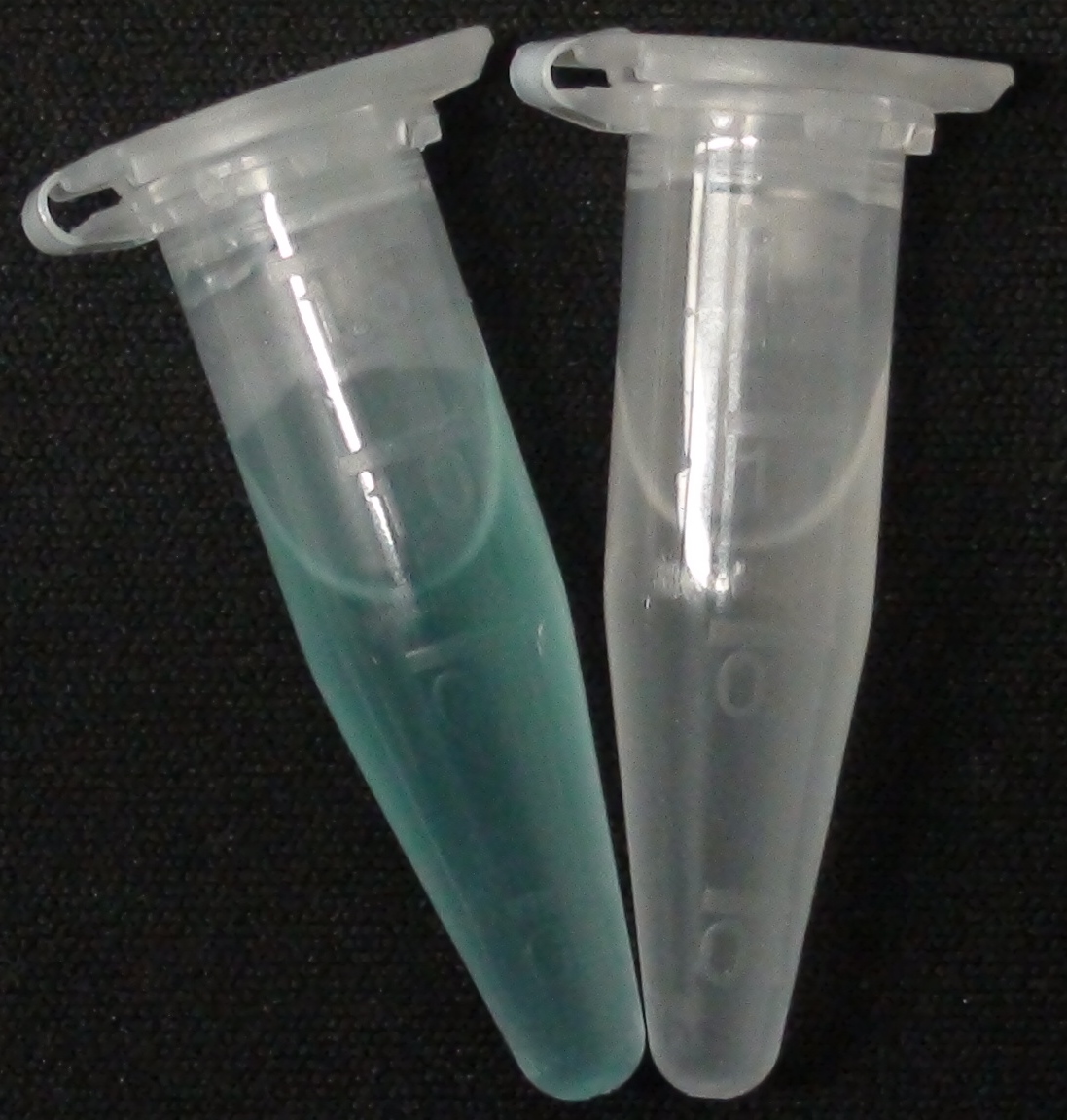
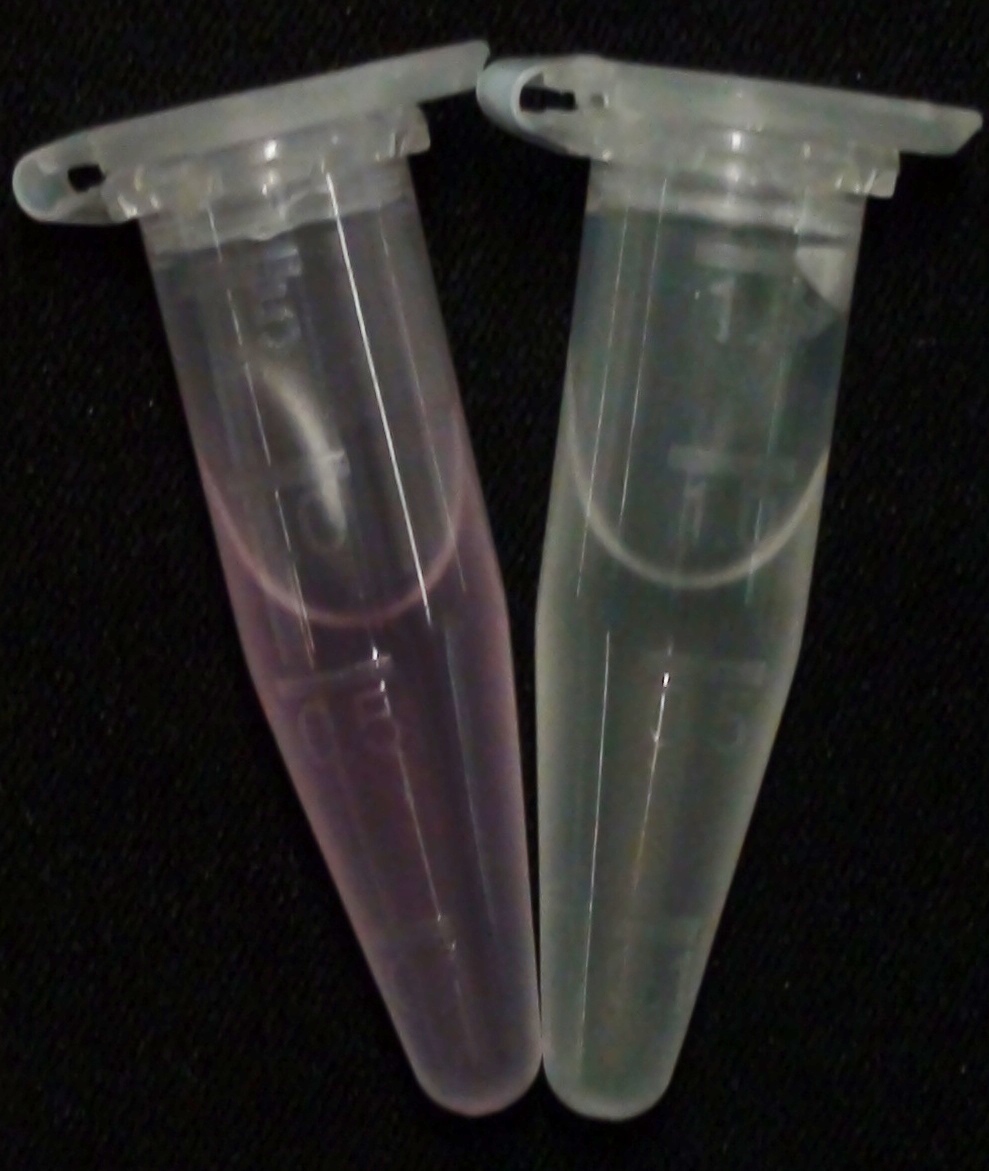
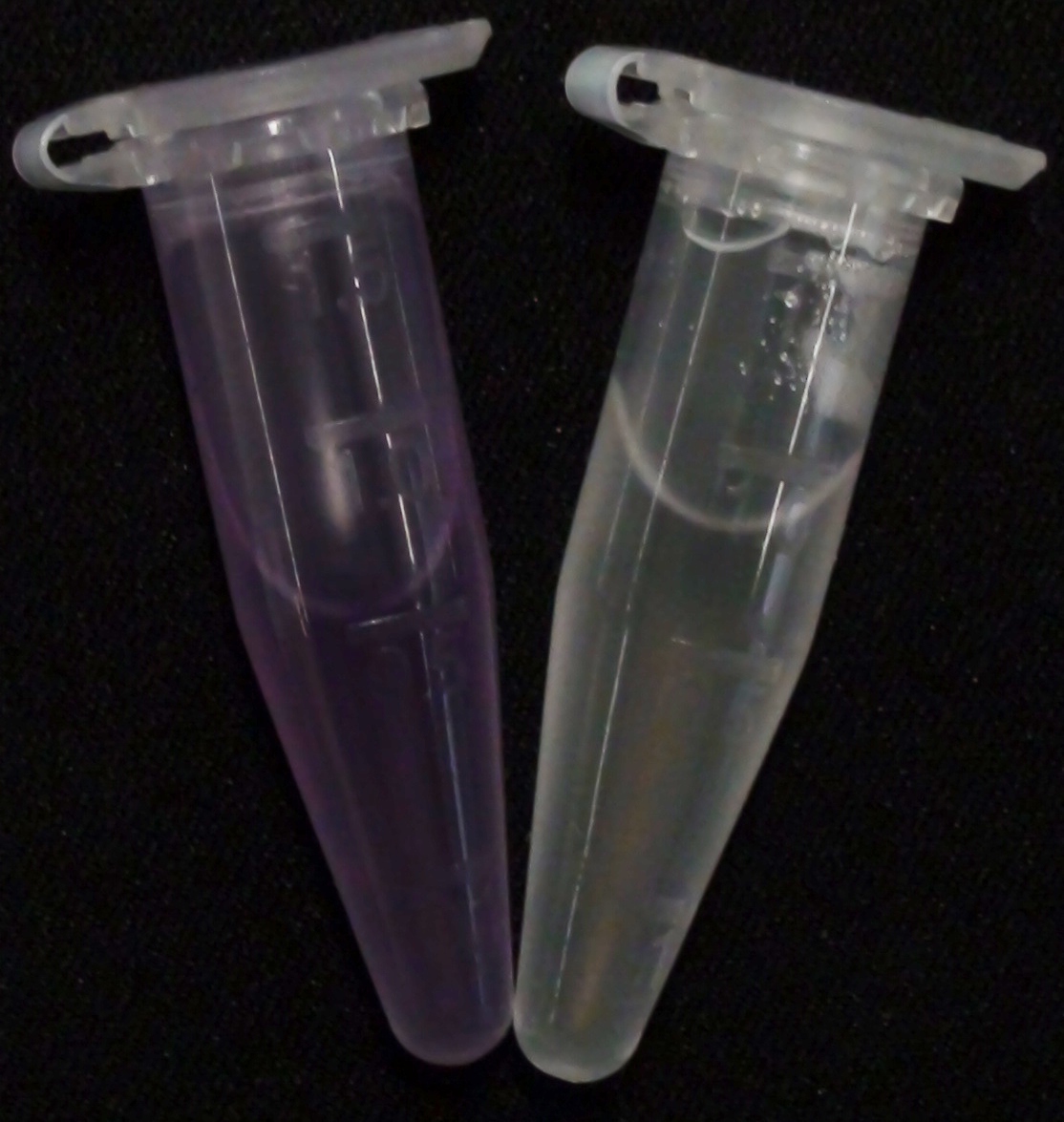
[[File:Aes MagentaButyrate.JPG|thumb|right|200px| <b>Figure 2.</b> Liquid culture of the triple knockout <i>Escherichia coli</i> strain left: overexpressing Aes and right: without overexpressing Aes; after addition of magenta-butyrate.]] | [[File:Aes MagentaButyrate.JPG|thumb|right|200px| <b>Figure 2.</b> Liquid culture of the triple knockout <i>Escherichia coli</i> strain left: overexpressing Aes and right: without overexpressing Aes; after addition of magenta-butyrate.]] | ||
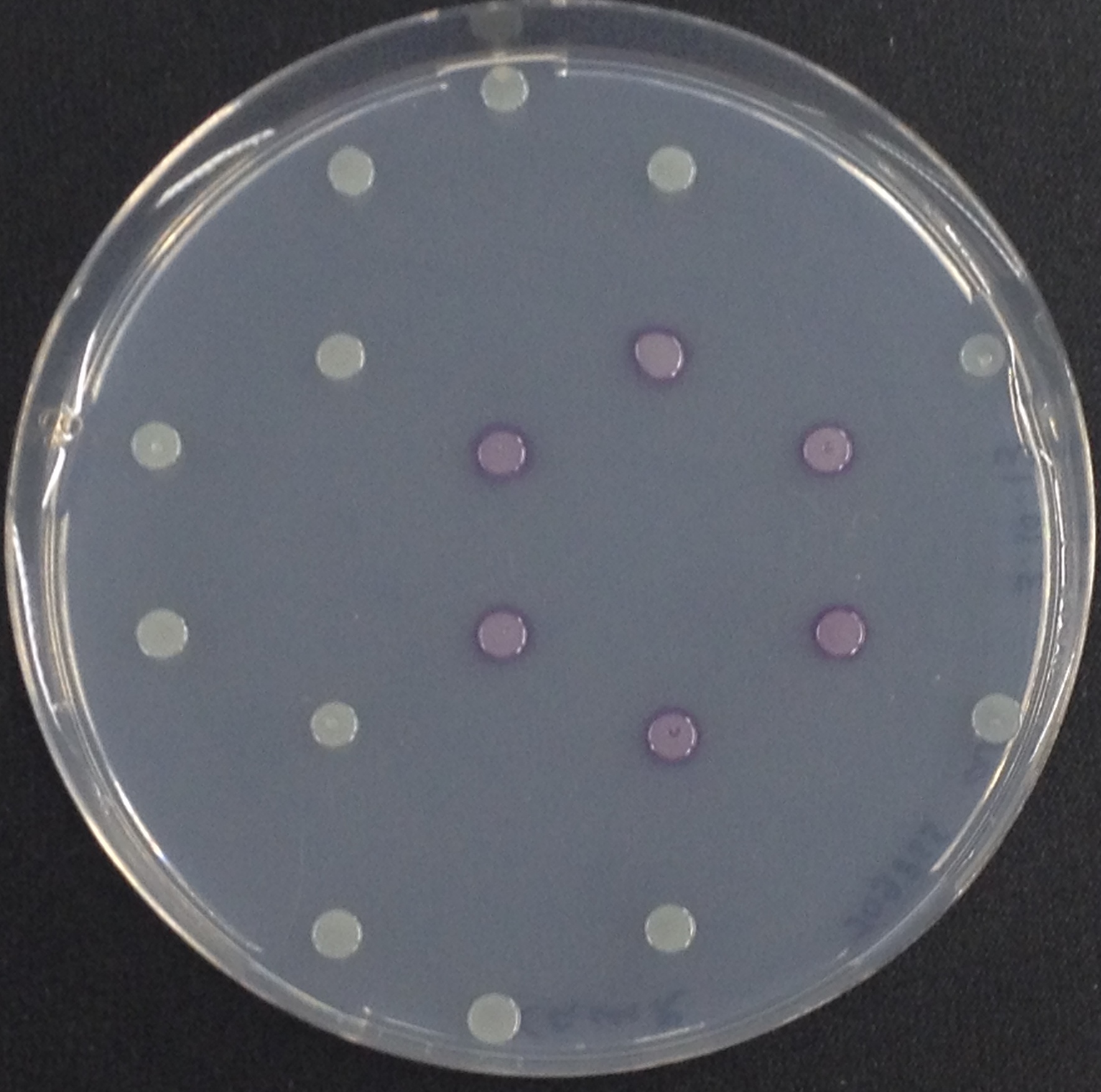
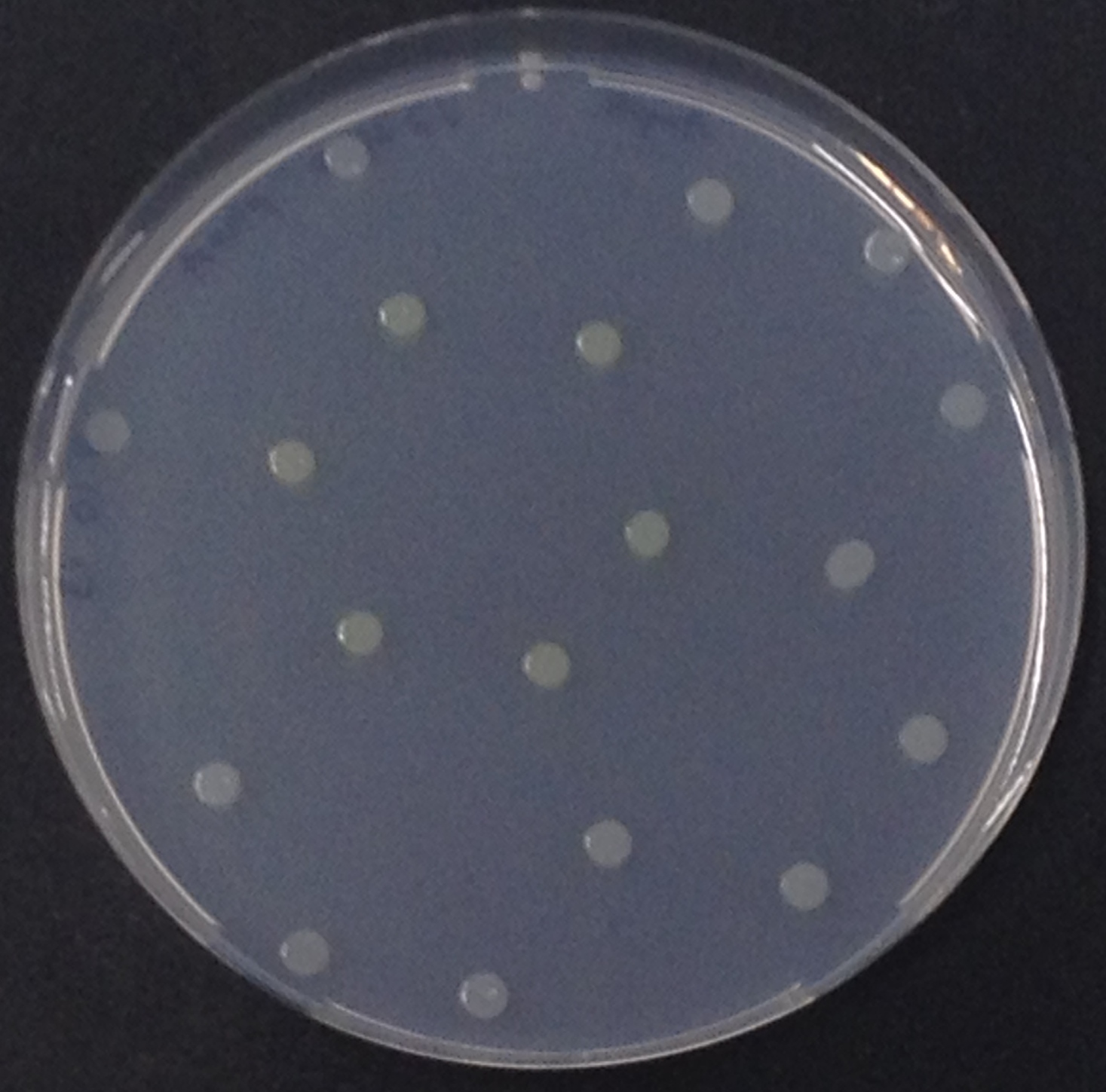
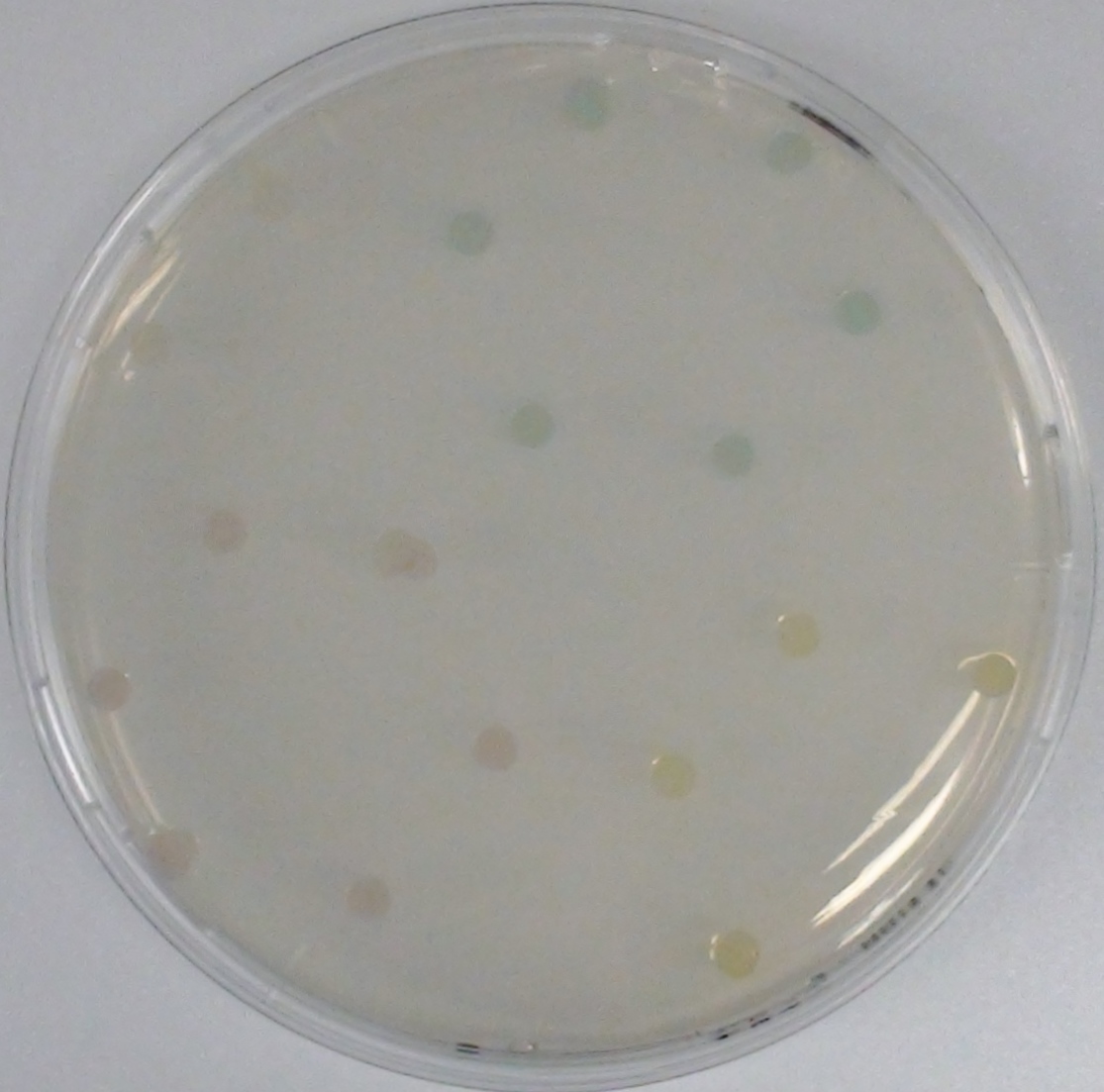
[[File:Aes and Magenta Butyrate.jpeg|thumb|right|200px| <b>Figure 3.1.</b> Colonies of ''Escherichia coli'' overexpressing the Aes. Colonies produce magenta color after addition of magenta butyrate.]] | [[File:Aes and Magenta Butyrate.jpeg|thumb|right|200px| <b>Figure 3.1.</b> Colonies of ''Escherichia coli'' overexpressing the Aes. Colonies produce magenta color after addition of magenta butyrate.]] | ||
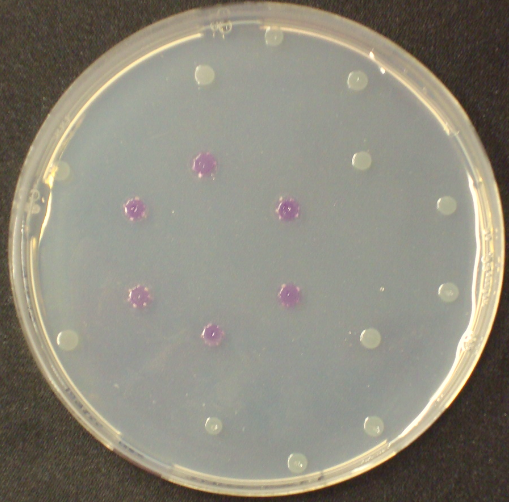
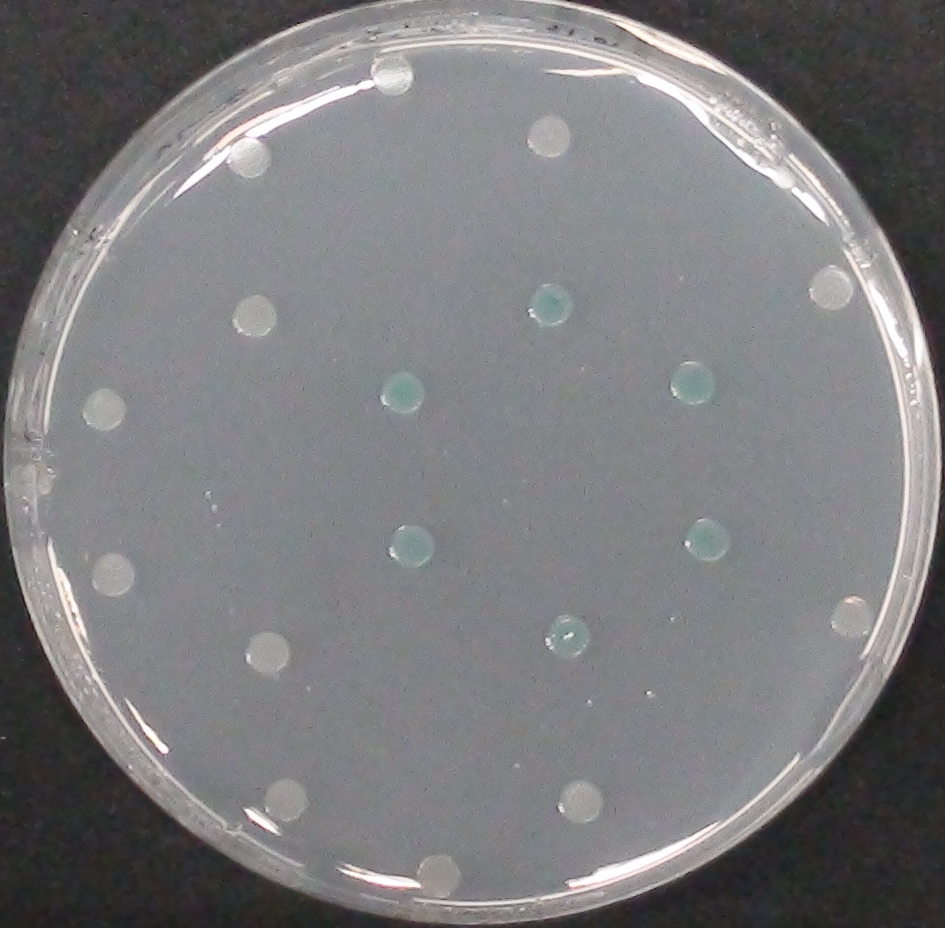
| - | [[File:MagentaCaprylate.png|thumb|right|200px| <b> Figure 3.2.</b> Colonies of ''Escherichia coli'' overexpressing the hydrolase Aes. Upon addition of magenta caprylate, a magenta color is produced after approximately 30 minutes incubation at 37°C.]] | + | [[File:MagentaCaprylate.png|thumb|right|200px| <b> Figure 3.2.</b> Colonies of the triple knockout ''Escherichia coli'' strain overexpressing the hydrolase Aes. Upon addition of magenta caprylate, a magenta color is produced after approximately 30 minutes incubation at 37°C.]] |
<p align="justify"><b>Chromogenic assay</b><br>To assess color development after reaction of the enzyme with the chromogenic substrate, a liquid culture of our triple knockout <i>Escherichia coli</i> strain overexpressing Aes was grown until an OD<sub>600</sub> of 0.4 - 0.6 was reached before addition of 5-Bromo-6-Chloro-3-indolyl butyrate (magenta butyrate, dissolved in acetone) to a final concentration of 100 µM (Figure 2). To study the color development in the actual Colisweeper game setup, colonies were plated by pipetting 1.5 µl of the triple knockout <i>Escherichia coli</i> liquid culture (OD<sub>600</sub> of 0.4 - 0.6) on an M9 agar plate and incubated overnight. Addition of 1.5 µl of 20 mM magenta butyrate onto each colony results in color generation visible after two to three minutes at room temperature (Figure 3.1).<br> | <p align="justify"><b>Chromogenic assay</b><br>To assess color development after reaction of the enzyme with the chromogenic substrate, a liquid culture of our triple knockout <i>Escherichia coli</i> strain overexpressing Aes was grown until an OD<sub>600</sub> of 0.4 - 0.6 was reached before addition of 5-Bromo-6-Chloro-3-indolyl butyrate (magenta butyrate, dissolved in acetone) to a final concentration of 100 µM (Figure 2). To study the color development in the actual Colisweeper game setup, colonies were plated by pipetting 1.5 µl of the triple knockout <i>Escherichia coli</i> liquid culture (OD<sub>600</sub> of 0.4 - 0.6) on an M9 agar plate and incubated overnight. Addition of 1.5 µl of 20 mM magenta butyrate onto each colony results in color generation visible after two to three minutes at room temperature (Figure 3.1).<br> | ||
Unfortunately, substrate tests in our triple knockout strain without Aes overexpression have shown that when using the butyrate as a substrate, there is background activity observable, meaning that enzymes other than the Aes can catalyze hydrolysis of the butyrate substrate. Therefore, 5-Bromo-6-Chloro-3-indolyl caprylate (magenta caprylate) was chosen as an alternative substrate, which has shown to be more specifically cleaved by the Aes in our knockout strain. However, when using magenta caprylate as the substrate, color on colonies developed only after at least half an hour of incubation at 37°C, or overnight at room temperature (Figure 3.2).<br> | Unfortunately, substrate tests in our triple knockout strain without Aes overexpression have shown that when using the butyrate as a substrate, there is background activity observable, meaning that enzymes other than the Aes can catalyze hydrolysis of the butyrate substrate. Therefore, 5-Bromo-6-Chloro-3-indolyl caprylate (magenta caprylate) was chosen as an alternative substrate, which has shown to be more specifically cleaved by the Aes in our knockout strain. However, when using magenta caprylate as the substrate, color on colonies developed only after at least half an hour of incubation at 37°C, or overnight at room temperature (Figure 3.2).<br> | ||
| - | Because the player of Colisweeper is not expected to incubate plates after each move, we made an assay comparing time differences of the color development in colonies overexpressing and not overexpressing Aes, using different concentrations of the substrates as well. Results have shown that with the butyrate substrate (20 mM in DMSO) and at room temperature (RT), colonies overexpressing Aes already developed color within the first few minutes, whereas colonies not overexpressing Aes did not show color until six hours after addition of the substrate. Even at twelve hours after adding the substrate, only a faint color was observed in the colonies not overexpressing Aes. Therefore, we continued using magenta butyrate for further tests and finally for the game, due to speed and intensity of color development compared to magenta caprylate. | + | Because the player of Colisweeper is not expected to incubate plates after each move, we made an assay comparing time differences of the color development in colonies of the triple knockout ''Escherichia coli'' strain overexpressing and not overexpressing Aes, using different concentrations of the substrates as well. Results have shown that with the butyrate substrate (20 mM in DMSO) and at room temperature (RT), colonies overexpressing Aes already developed color within the first few minutes, whereas colonies not overexpressing Aes did not show color until six hours after addition of the substrate. Even at twelve hours after adding the substrate, only a faint color was observed in the colonies not overexpressing Aes. Therefore, we continued using magenta butyrate for further tests and finally for the game, due to speed and intensity of color development compared to magenta caprylate. |
</p><br><br> | </p><br><br> | ||
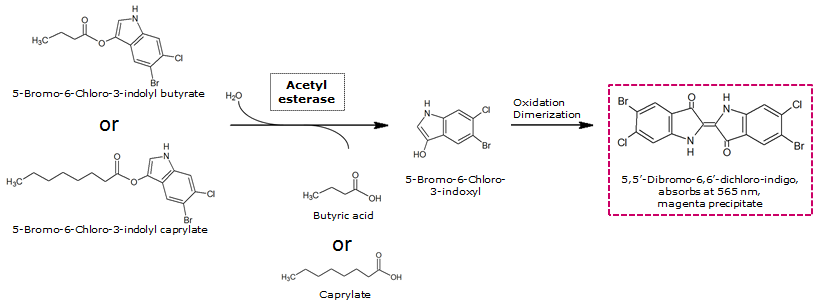
[[File:AesSubstrates.png|thumb|left|743px| <b>Figure 4.</b> Aes catalyzed hydrolysis reaction of magenta butyrate or magenta caprylate.]] | [[File:AesSubstrates.png|thumb|left|743px| <b>Figure 4.</b> Aes catalyzed hydrolysis reaction of magenta butyrate or magenta caprylate.]] | ||
Revision as of 18:53, 28 October 2013
Contents[hide] |
Enzyme-substrate reactions
To generate visible output by adding substrates to colonies in Colisweeper, we made use of orthogonal enzyme-substrate reactions. A set of chromogenic substrates was chosen to produce different colors depending on the abundant hydrolases and thereby to uncover the identity of each colony to the player.
The set of enzyme-substrate pairs chosen for the Colisweeper project, and characterizations of their reactions, are described below. Information on other possible substrates that can be used for the enzymes of the Colisweeper reporter system can be found in the reporter system section.
To characterize the hydrolases used in Colisweeper, we conducted substrate tests in liquid cultures as well as on colonies, enzyme kinetics and detection of HIS-tagged protein by Western Blot. Additionally, crosstalk assays and color overlay tests (multiple enzyme-substrate reactions together) have been performed for this project. A listing of chromogenic substrates used can be found in the Materials section.
General characterisitcs overview
| Hydrolase | Substrate | Color, Absorption λmax |
Stock | Liquid culture | Colonies | Response time on E. coli colonies, at RT |
|---|---|---|---|---|---|---|
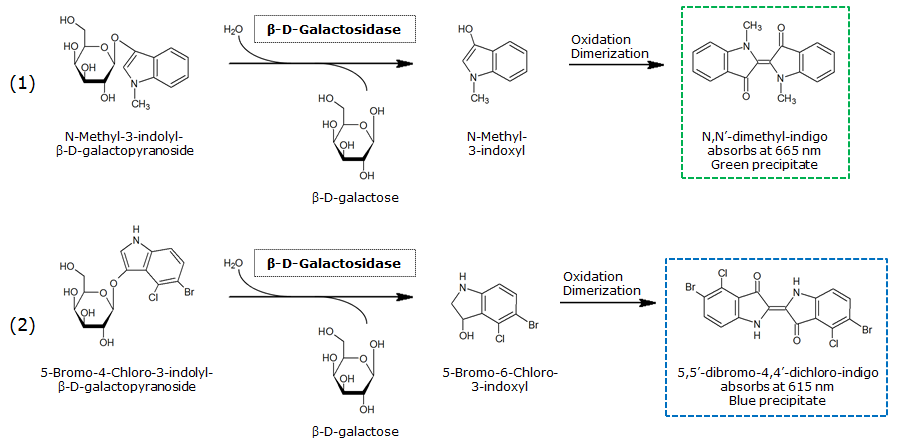
| LacZ | 5-Bromo-4-Chloro-3-indoxyl-β-D-galactopyranoside (X-Gal) | Blue, 615 nm |
0.5 M in DMSO | 1 mM | 50 mM | ~ 10 minutes |
| N-Methyl-3-indolyl-β-D-galactopyranoside (Green-Gal) | Green, 665 nm |
0.1 M in DMSO | 1 mM | 50 mM | ~ 15 minutes | |
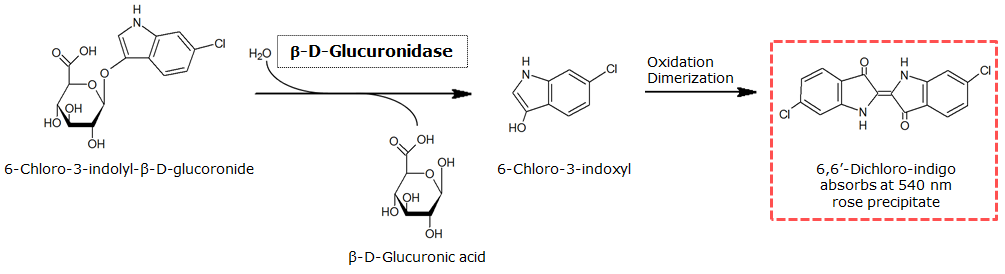
| GusA | 6-Chloro-3-indolyl-β-D-glucuronide (Salmon-Glc) | Salmon, 540 nm |
0.3 M in DMSO | 1.5 mM | 0.1 M | ~ 5 minutes |
| PhoA | 4-Nitrophenoyl-phosphate (pNPP) | Yellow, 405 nm |
0.5 M in DEA | 50 mM | 0.5 M | ~ 1 minutes |
| 5-Bromo-4-Chloro-3-indolyl phosphate (BCIP) | Blue, 615 nm |
0.1 M in H2O | 1 mM | 50 mM | ~ 30 minutes | |
| Aes | 5-Bromo-6-Chloro-3-indoxyl butyrate (Magenta butyrate) | Magenta, 565 nm |
0.5 M in Acetone | 0.1 mM | 20 mM | ~ 2 minutes |
| 5-Bromo-6-Chloro-3-indoxyl caprylate (Magenta caprylate) | Magenta, 565 nm |
0.2 M in Acetone | 1 mM | 0.2 M | overnight | |
| NagZ | 4-Nitrophenyl- N-acetyl-β-D-glucosaminide (pNP-GluNAc) | Yellow, 405 nm |
15 mM in H2O | 0.01 mM | 15 mM | ~ 1 minutes |
| 5-Bromo-6-Chloro-3-indolyl N-acetyl-β-D-glucosaminide (Magenta GluNAc) | Magenta, 565 nm |
0.1 M in DMSO | 1 mM, supplemented with BSA | 50 mM, with BSA added | ~ 15 minutes | |
| 5-Bromo-4-Chloro-3-indolyl N-acetyl-β-D-glucosaminide (X-GluNAc) | Blue, 615 nm |
0.1 M in DMSO | 1 mM, supplemented with BSA | 50 mM, with BSA added | ~ 15 minutes |
Expand the boxes to see the characterization of the hydrolases
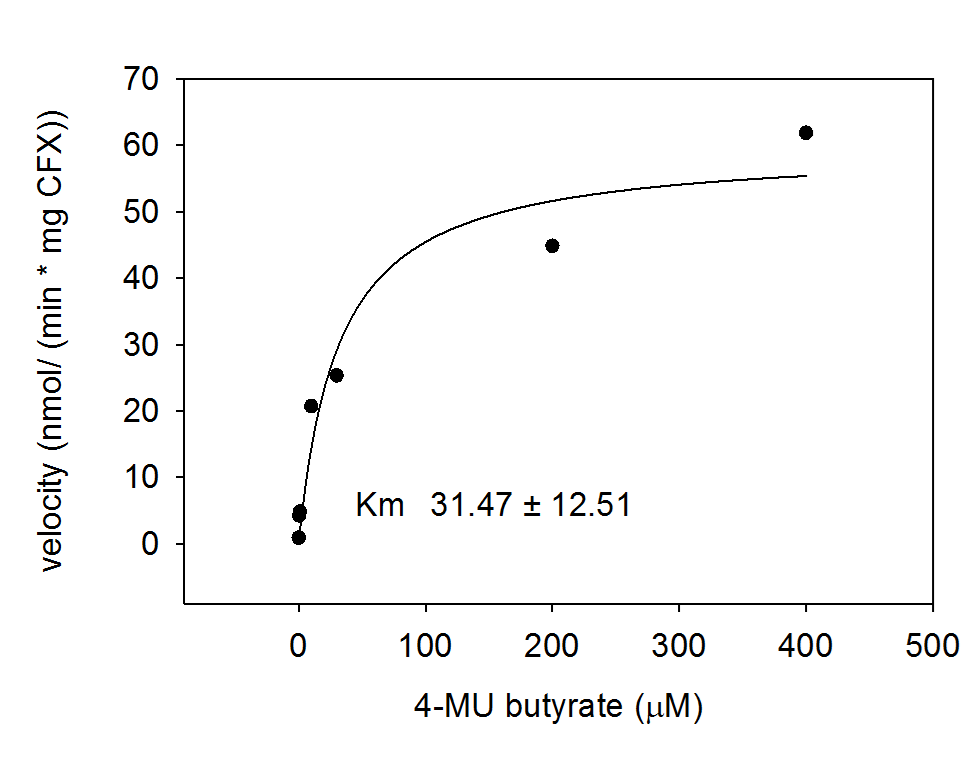
Acetyl esterase (Aes)
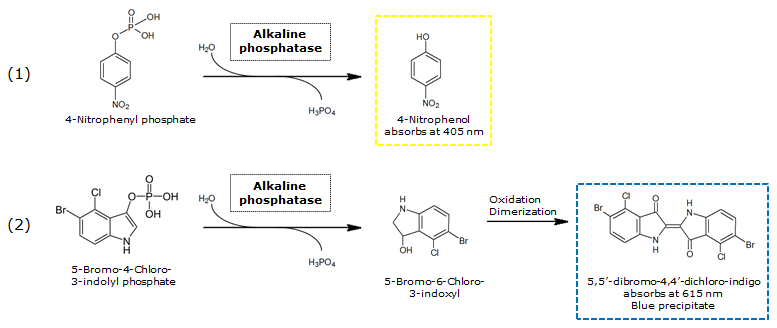
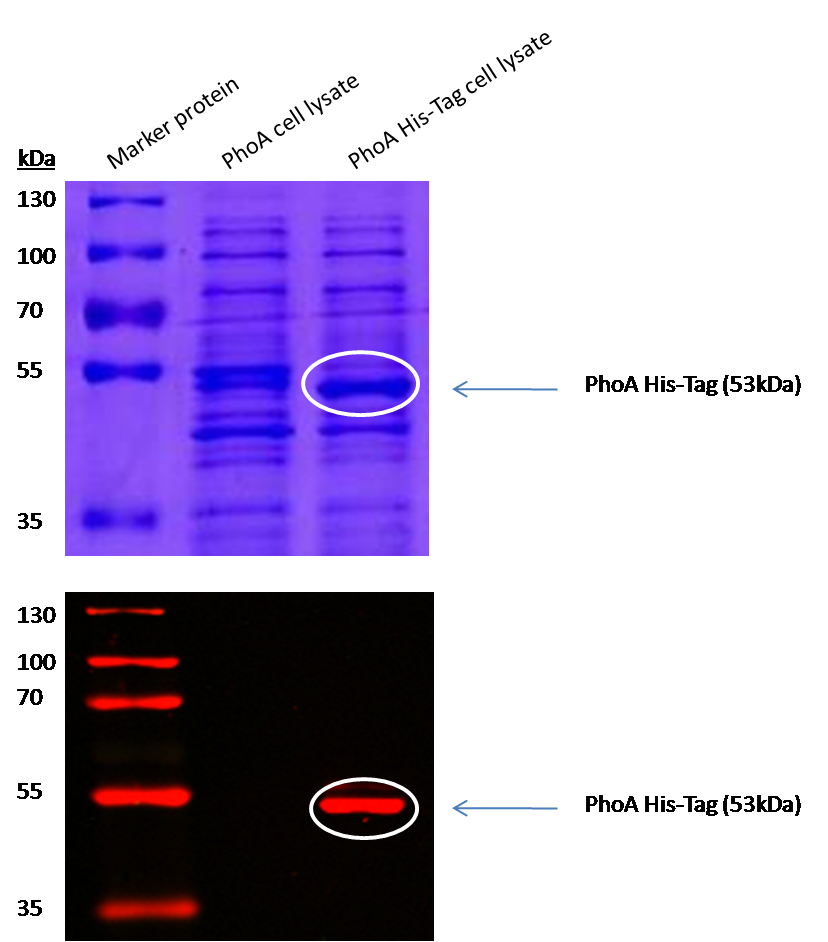
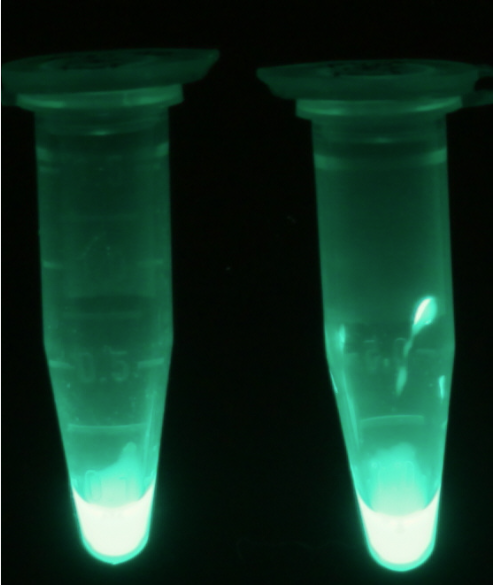
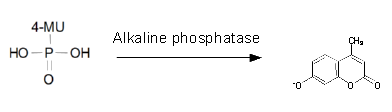
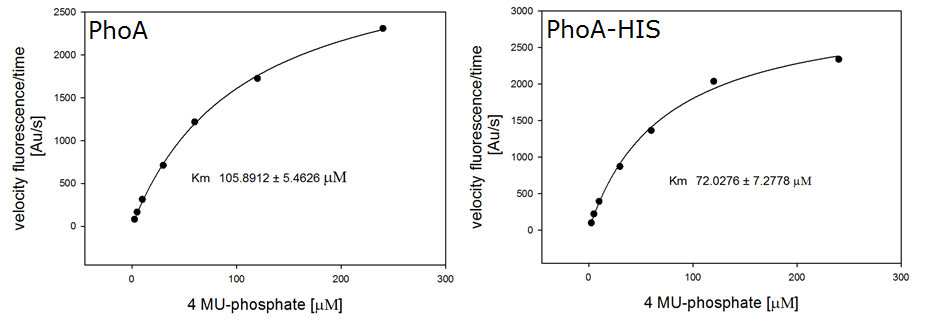
Alkaline phosphatase (PhoA)
β-Galactosidase (LacZ)
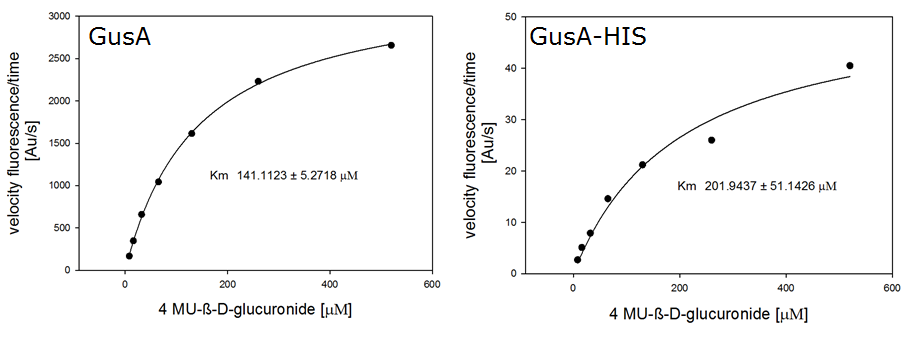
β-Glucuronidase (GusA)
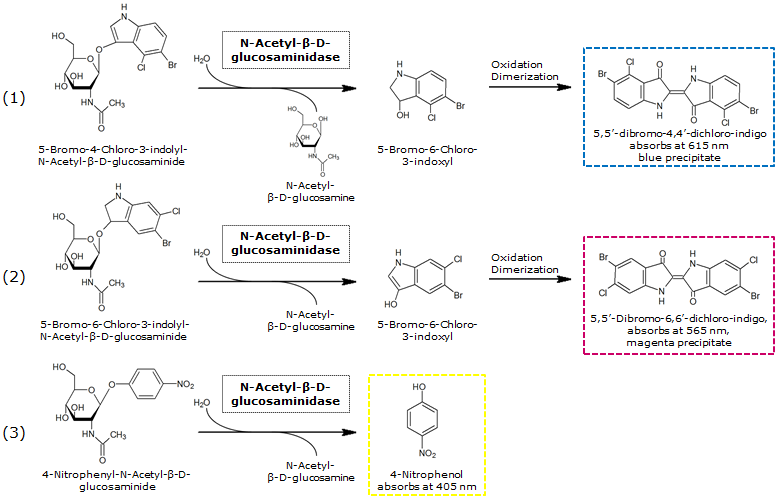
β-N-Acetylglucosaminidase (NagZ)
Crosstalk
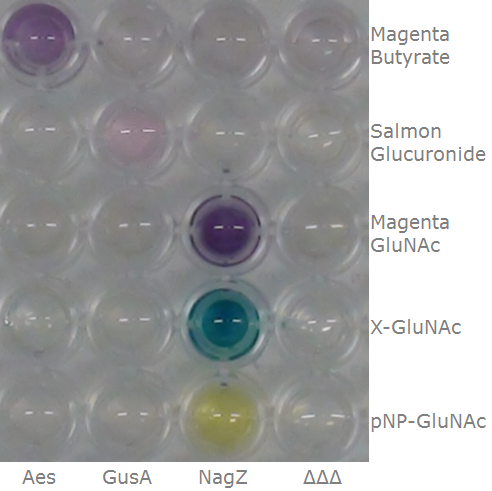
To ensure specificity of the enzyme-substrate pairs used in Colisweeper, a crosstalk test was done to make sure that all overexpressed enzymes specifically cleave their assigned substrate.
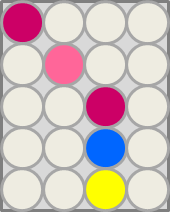
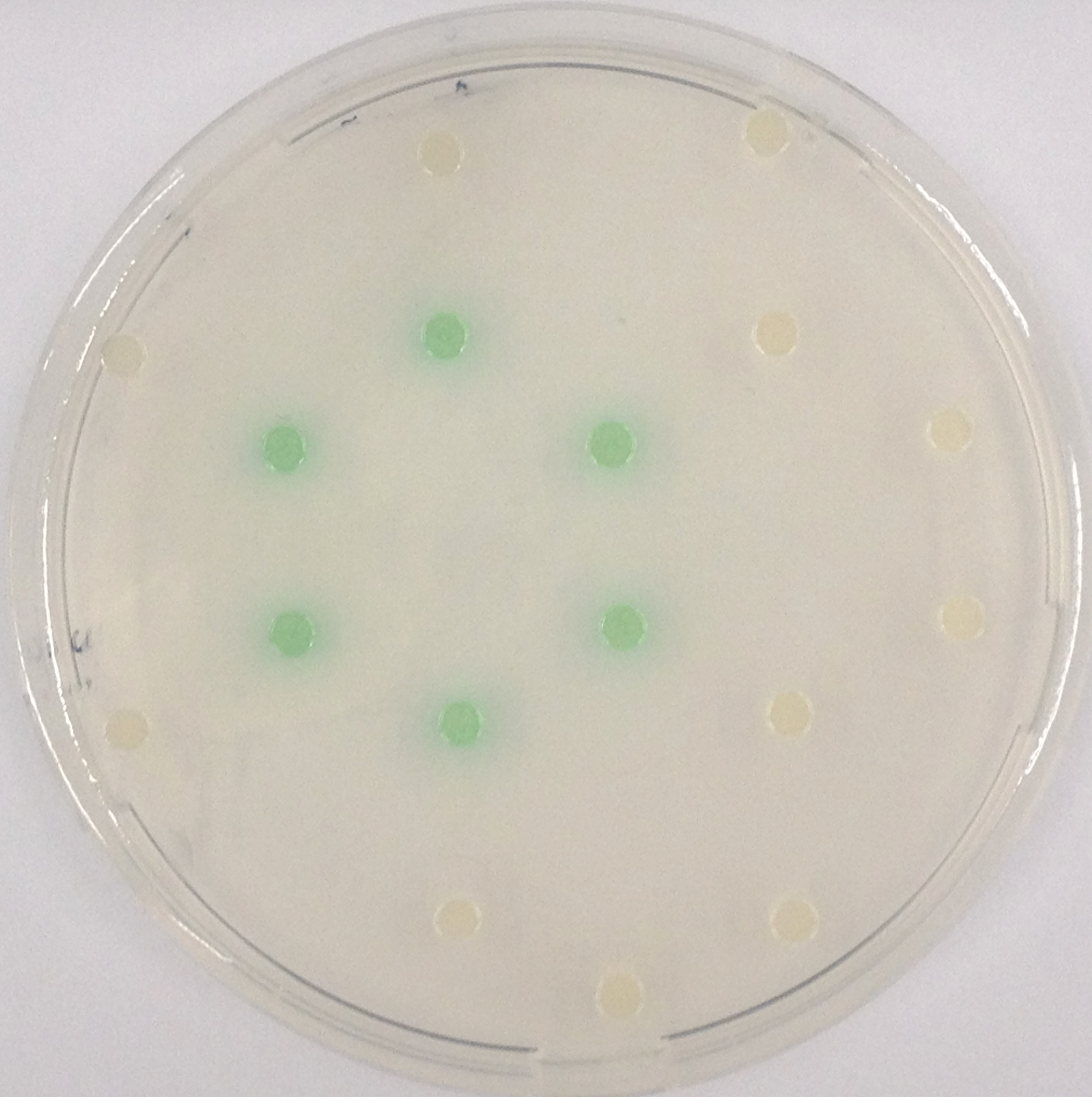
This crosstalk test was done in a 96-well plate, each well containing 200 μl from liquid cultures of our ΔaesΔgusAΔnagZ Escherichia coli strain overexpressing either Aes, GusA, NagZ or none, each distributed among the column-wells of the plate. Horizontally, the chromogenic substrates were pipetted to the liquid cultures in the same order as their corresponding hydrolase. If specificity of the chosen enzyme-substrates pairs were given, we would expect an output as shown in the figure below (Figure 26.1.). As Figure 26.2. shows, the overexpressed hydrolases cleave only the substrates they were expected to.
Color overlay

To play Colisweeper, colors for each colony identity have to be clearly distinguishable by eye. Because we are using high-pass filters to differentially process certain AHL levels, we had to make sure that a mix of cleaved product would show distinguishable colors.
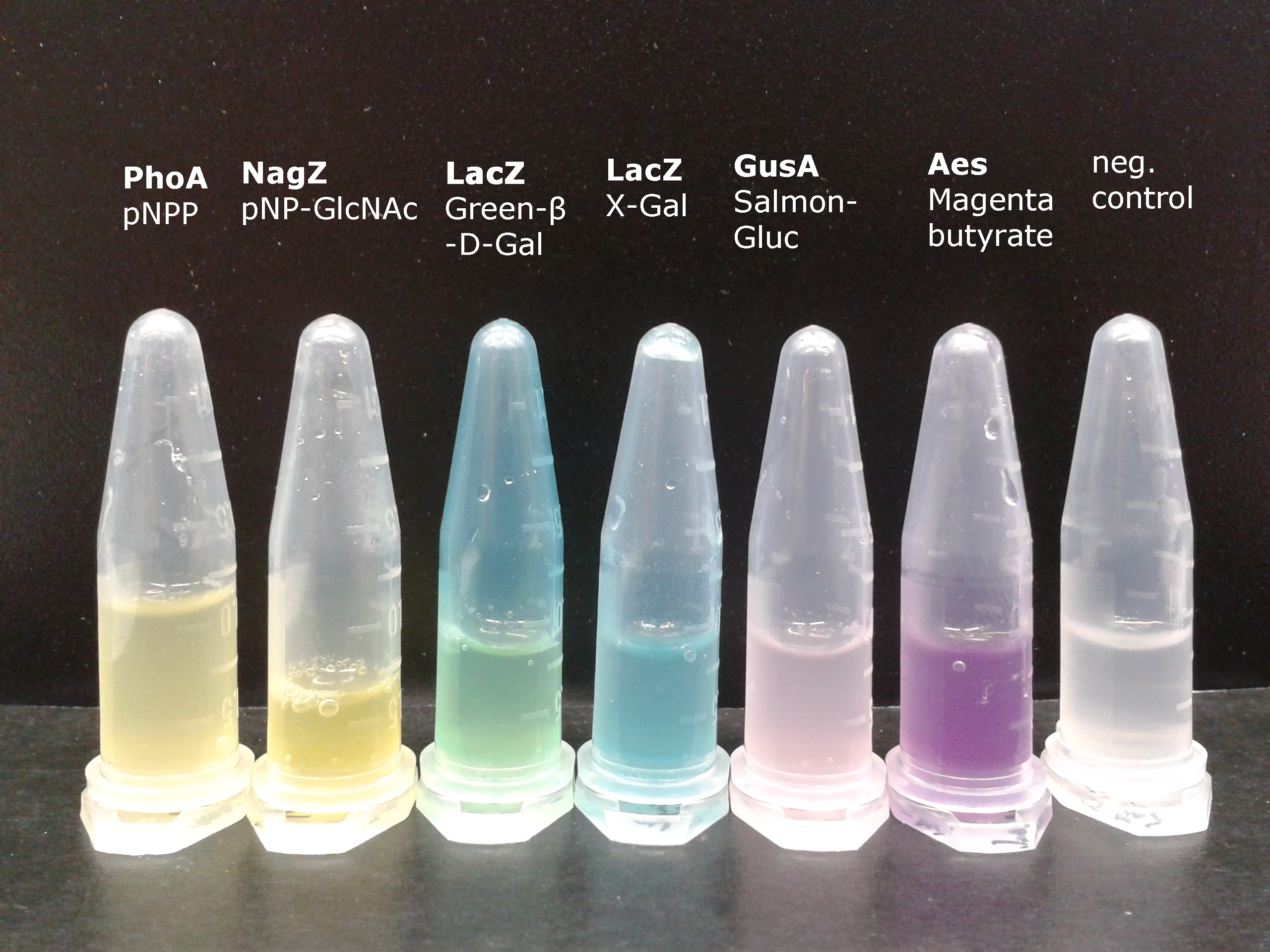
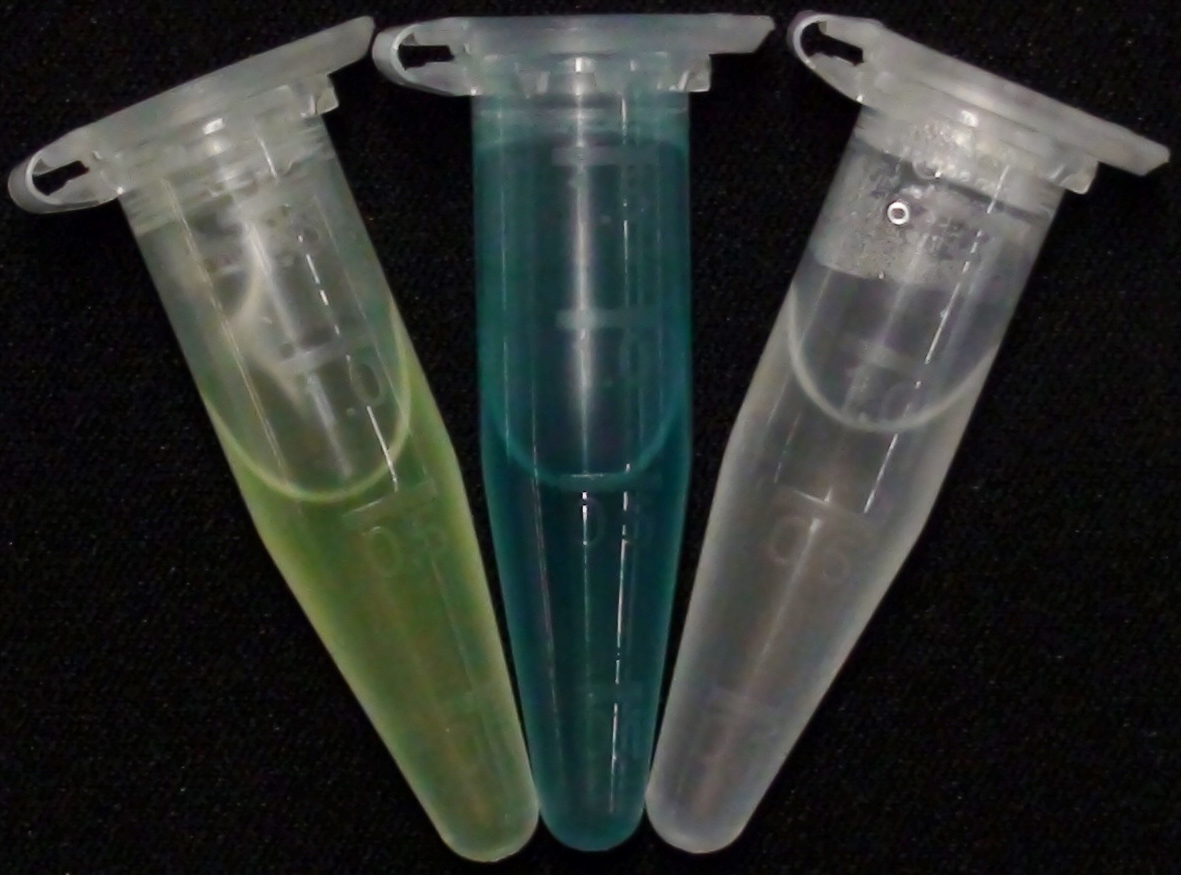
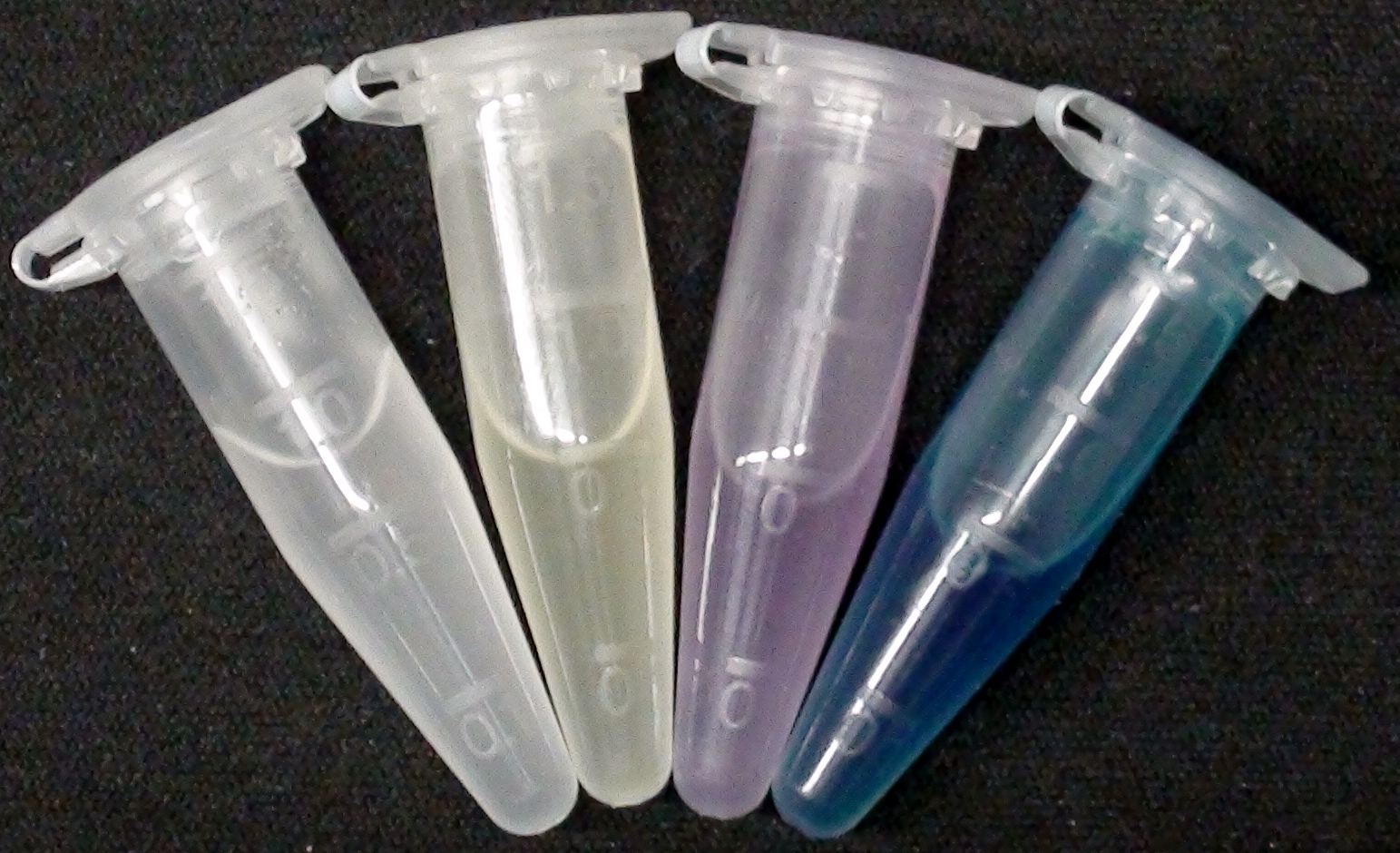
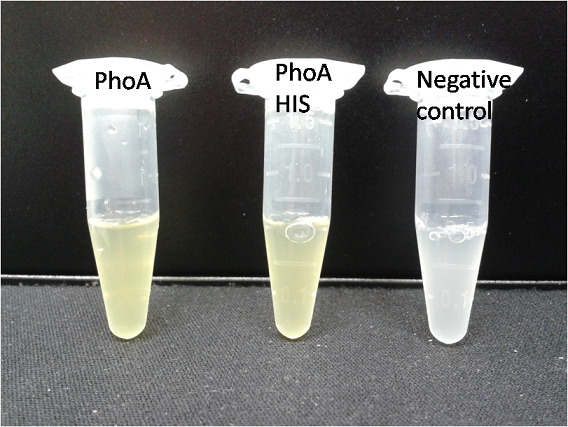
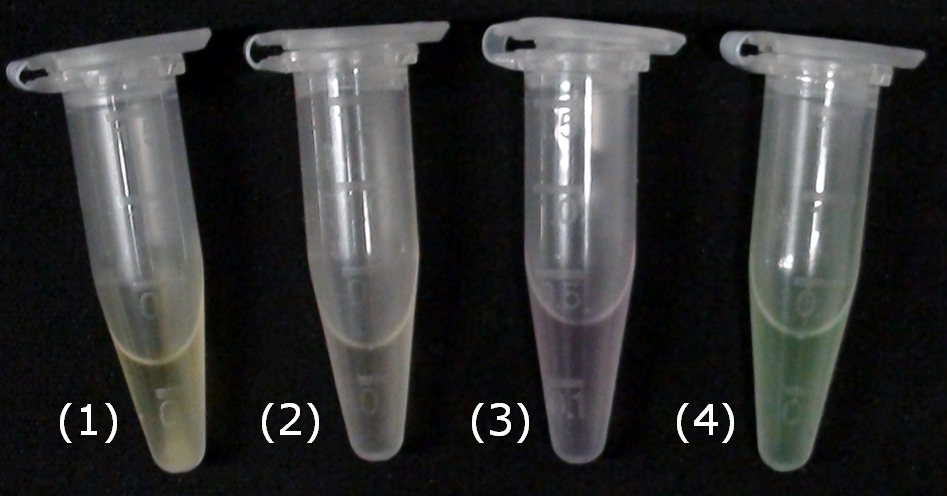
For this purpose, we mixed liquid cultures of our ΔaesΔgusAΔnagZ triple knockout strain which overexpress PhoA, GusA, Aes or NagZ, and added the corresponding substrates to the cultures. Figure 27 shows the resulting colors after hydrolysis of the substrates: PhoA with pNPP in (1), PhoA with pNPP and GusA with Salmon-Gluc in (2), PhoA with pNPP, GusA with Salmon-Gluc and Aes with Magenta butyrate in (3), and PhoA with pNPP and nagZ with X-GluNAc in (4). These colors are clearly distinguishable from one another and therefore can be used together as reporters of a system incorporating high-pass filters.
References
(1) Orenga S, Roger-Dalbert C, James A, Perry J, US Patent 20090017481 (2009).
(2) Magnelli PE, Bielik A, Guthrie E, Methods Mol. Biol., 801, 189-211 (2011).
Absorption: Biosynth
 "
"