Team:ETH Zurich/Experiments 3
From 2013.igem.org
Contents[hide] |
Enzyme-substrate reactions
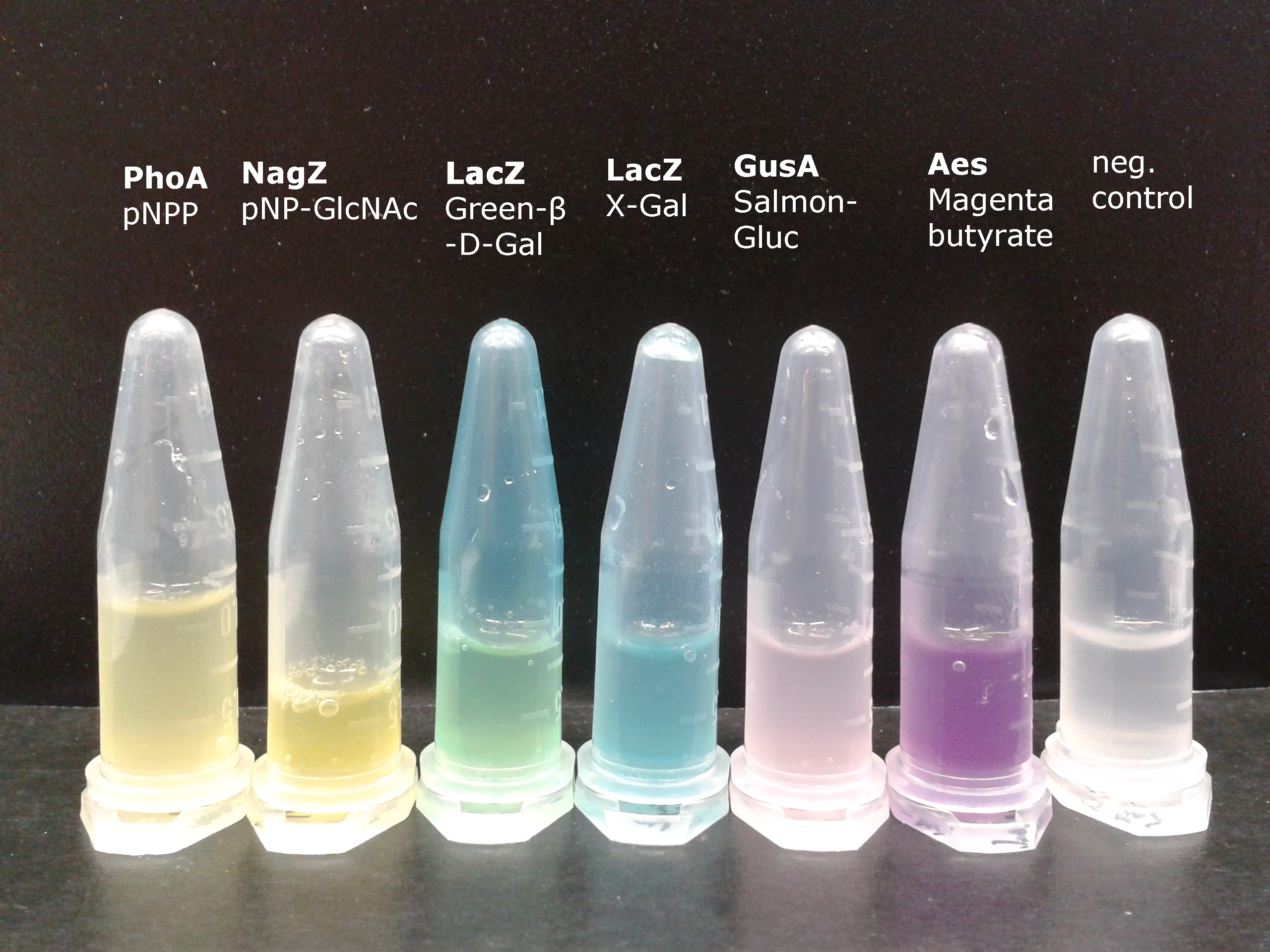
To generate visible output by adding substrates to colonies in Colisweeper, we made use of orthogonal enzyme-substrate reactions. A set of chromogenic substrates was chosen to produce different colors depending on the abundant hydrolases and thereby to uncover the identity of each colony to the player.
The set of enzyme-substrate pairs chosen for the Colisweeper project, and characterizations of their reactions, are described below. Information on other possible substrates that can be used for the enzymes of the Colisweeper reporter system can be found in the reporter system section.
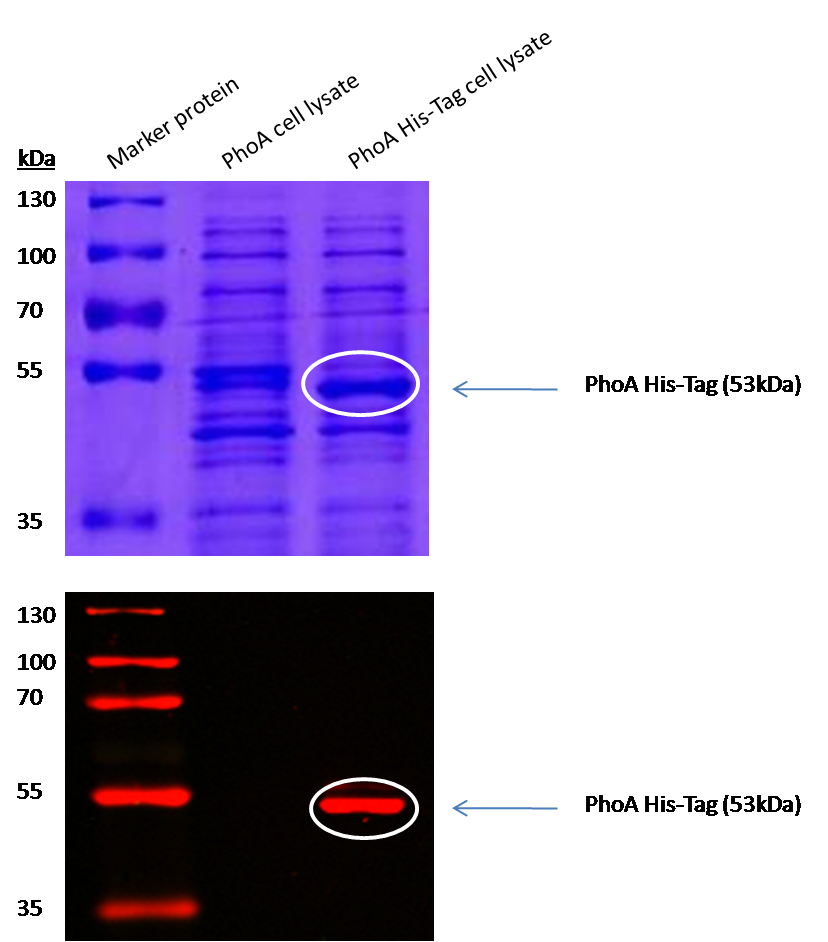
To characterize the hydrolases used in Colisweeper, we conducted substrate tests in liquid cultures as well as on colonies, enzyme kinetics and detection of HIS-tagged protein by Western Blot. Additionally, crosstalk assays and color overlay tests (multiple enzyme-substrate reactions together) have been performed for this project. An overview of chromogenic substrates used can be found in the Materials section.
Expand the boxes to see the characterization of the hydrolases
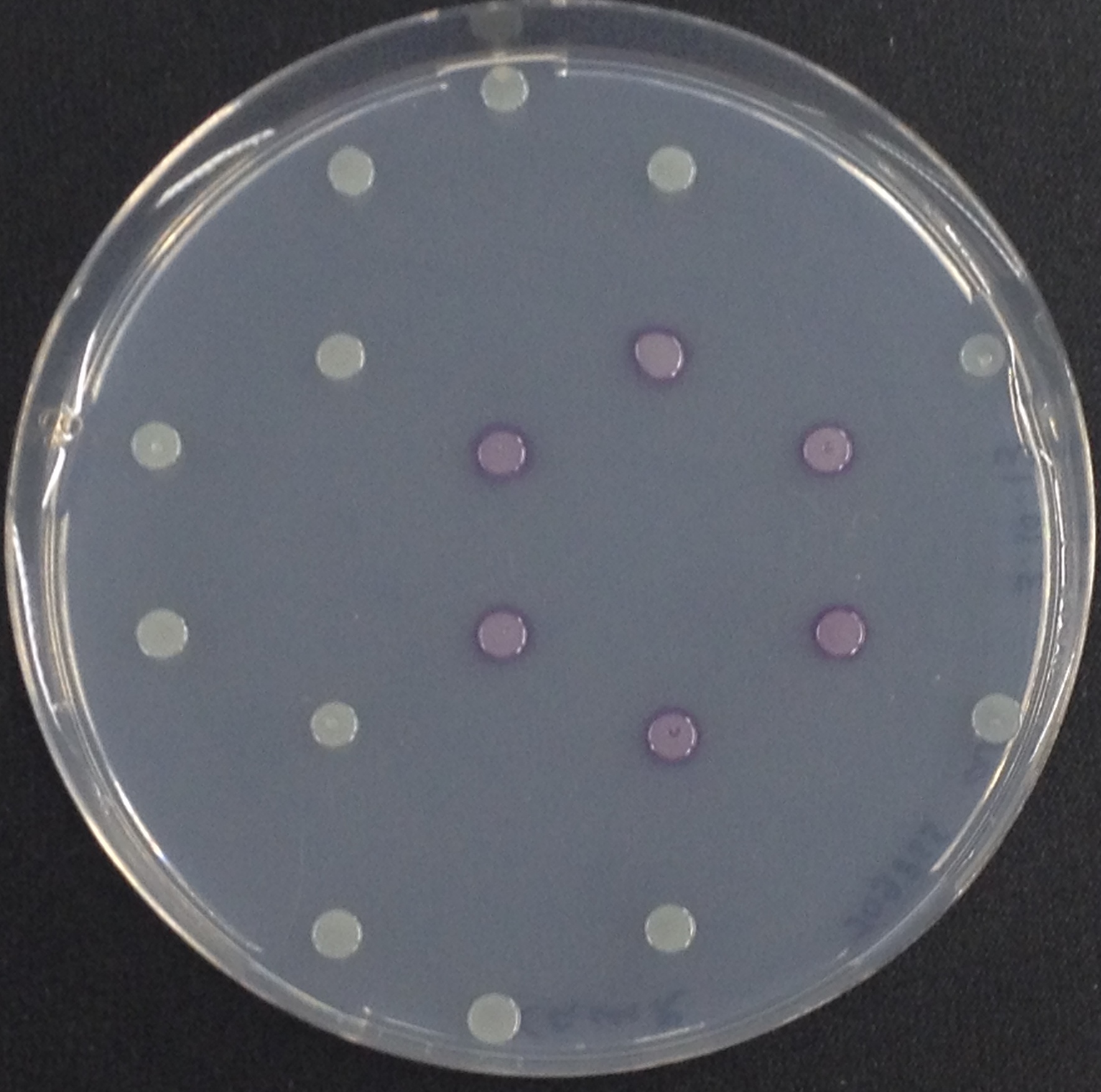
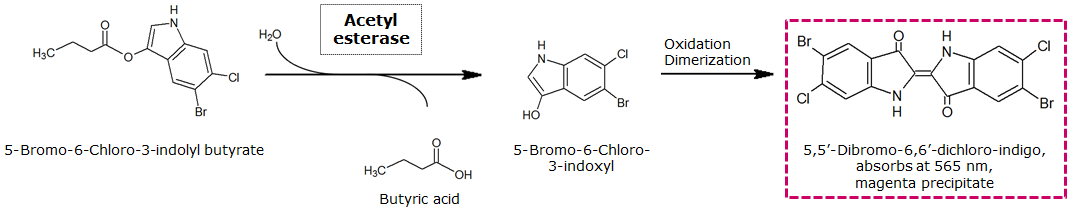
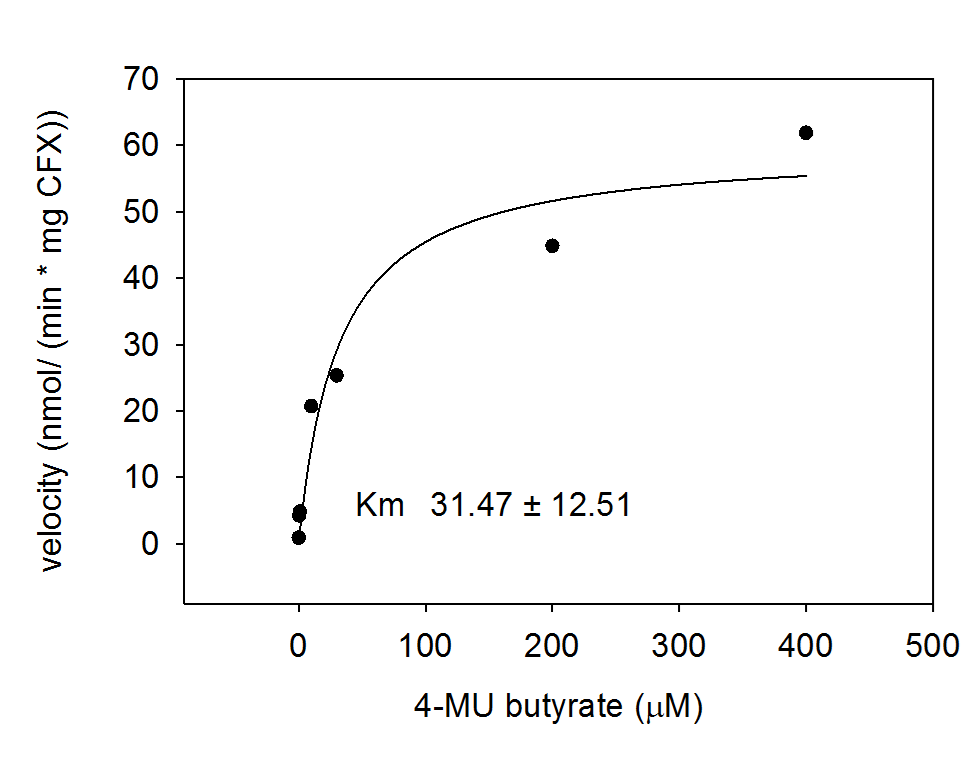
Acetyl esterase (Aes)
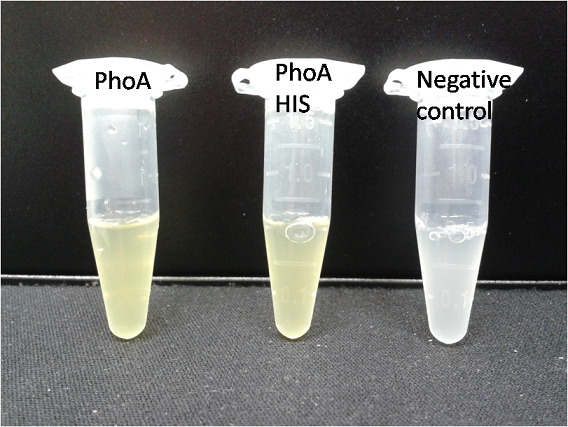
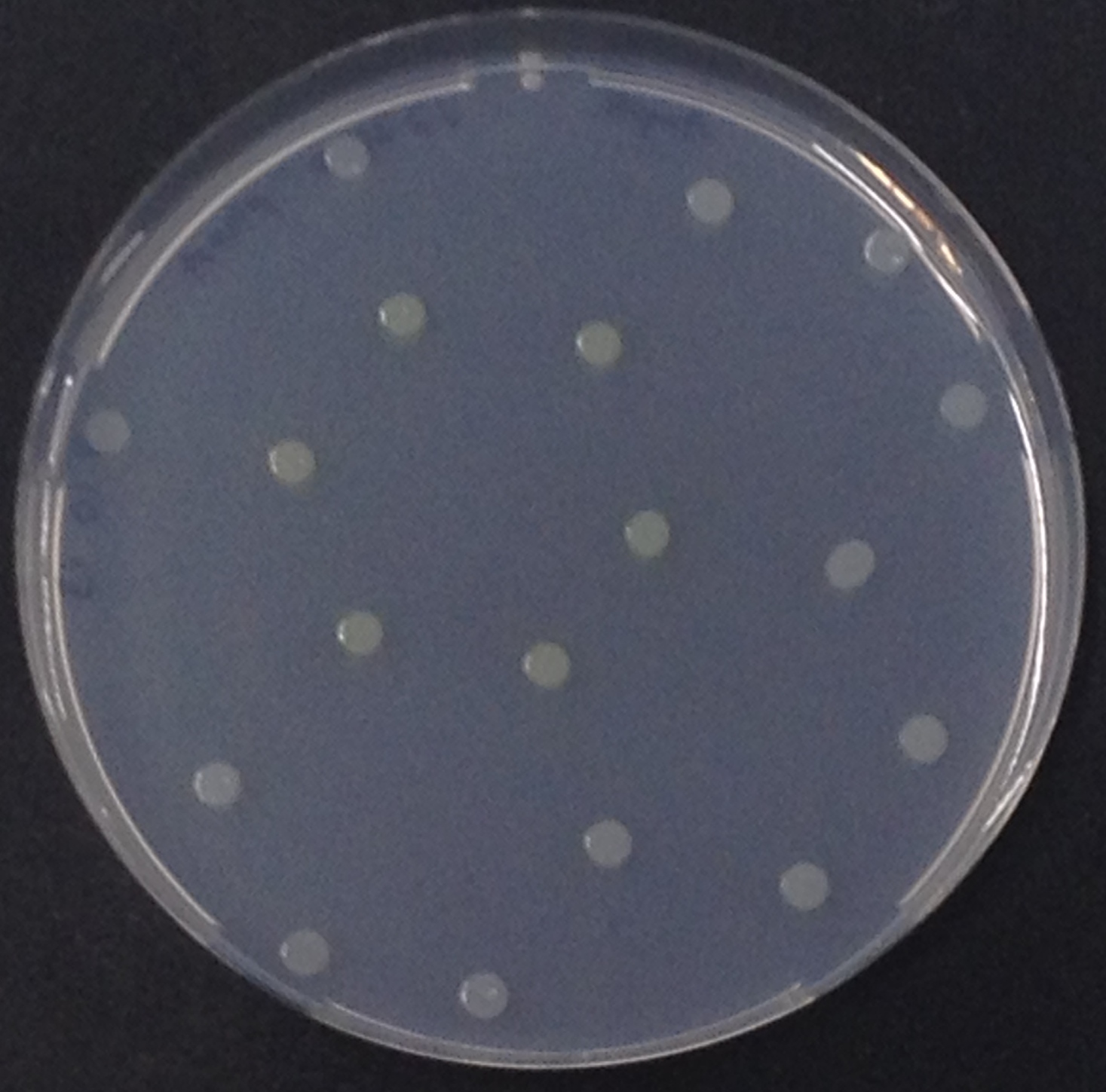
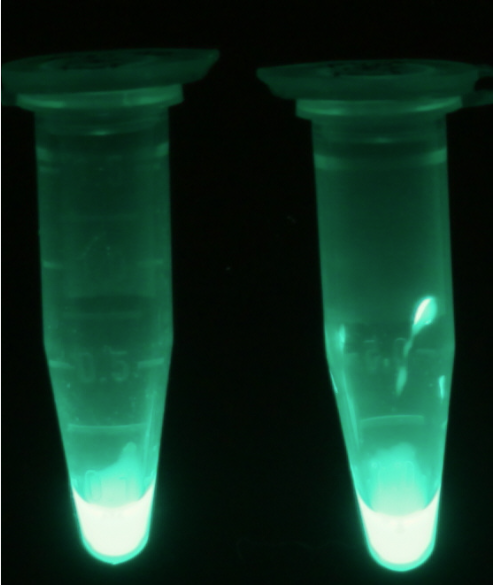
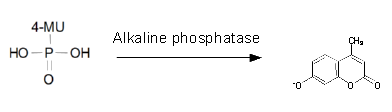
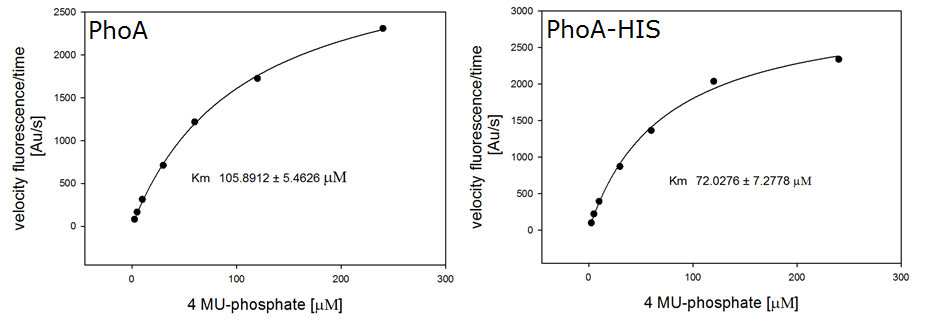
Alkaline phosphatase (PhoA)
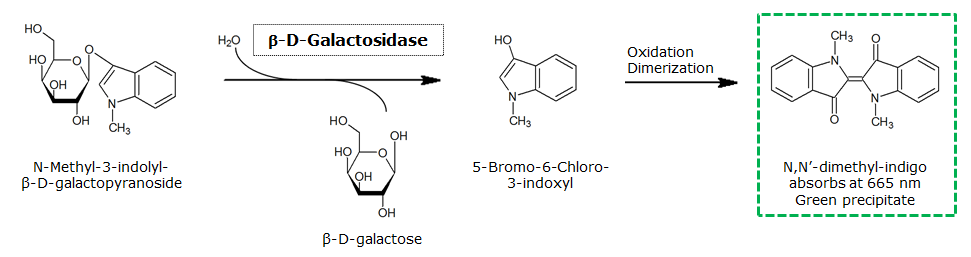
β-Galactosidase (LacZ)
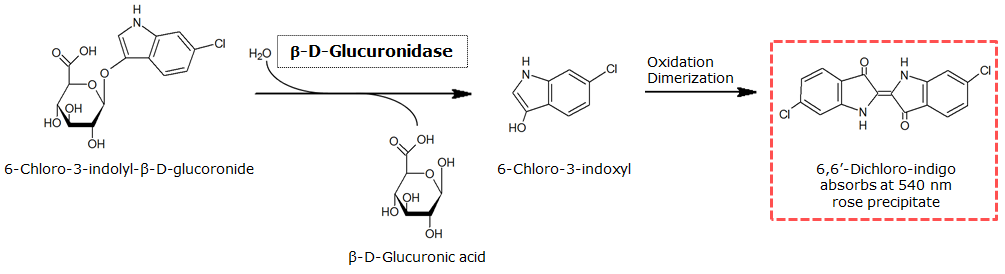
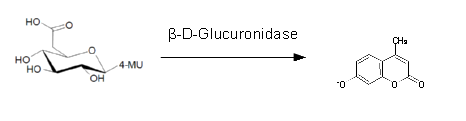
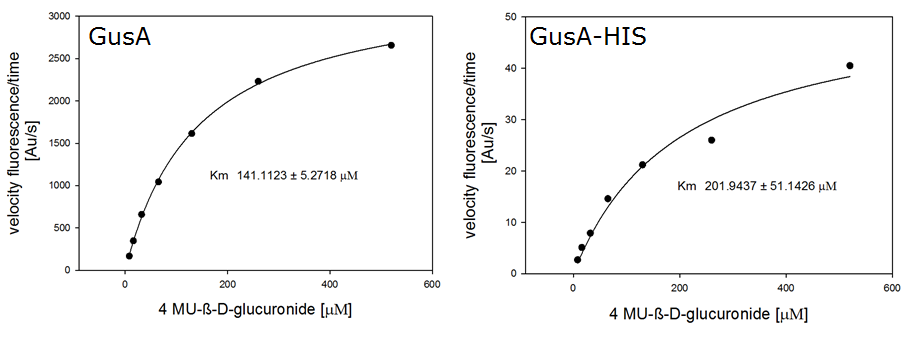
β-Glucuronidase (GusA)
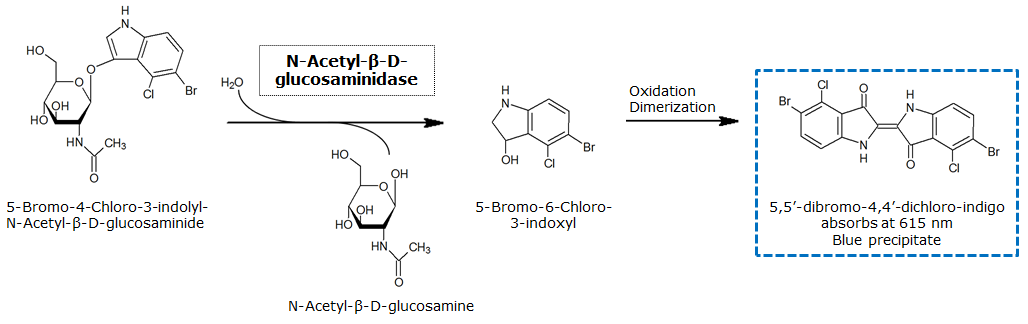
β-N-Acetylglucosaminidase (NagZ)
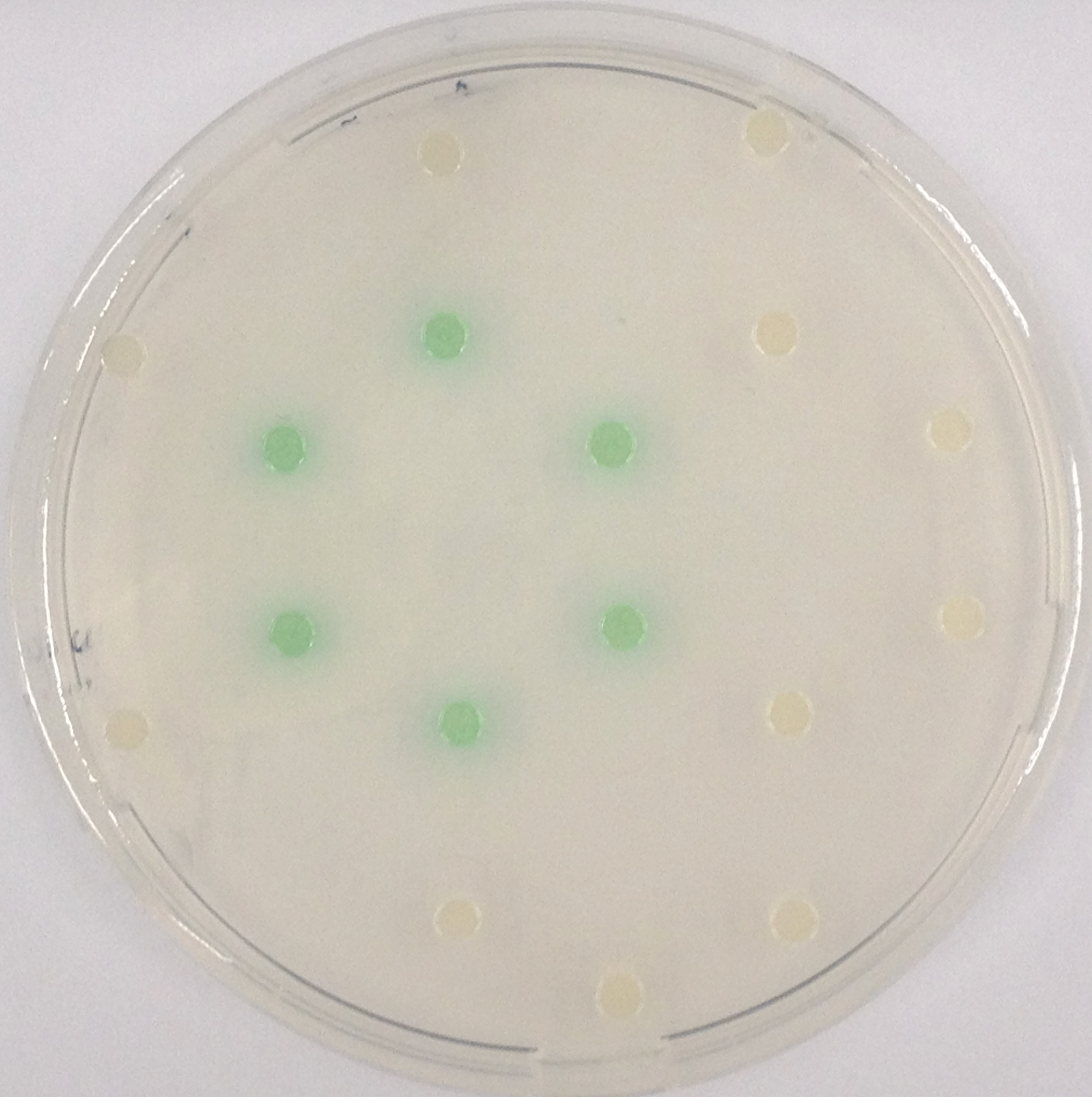
Crosstalk
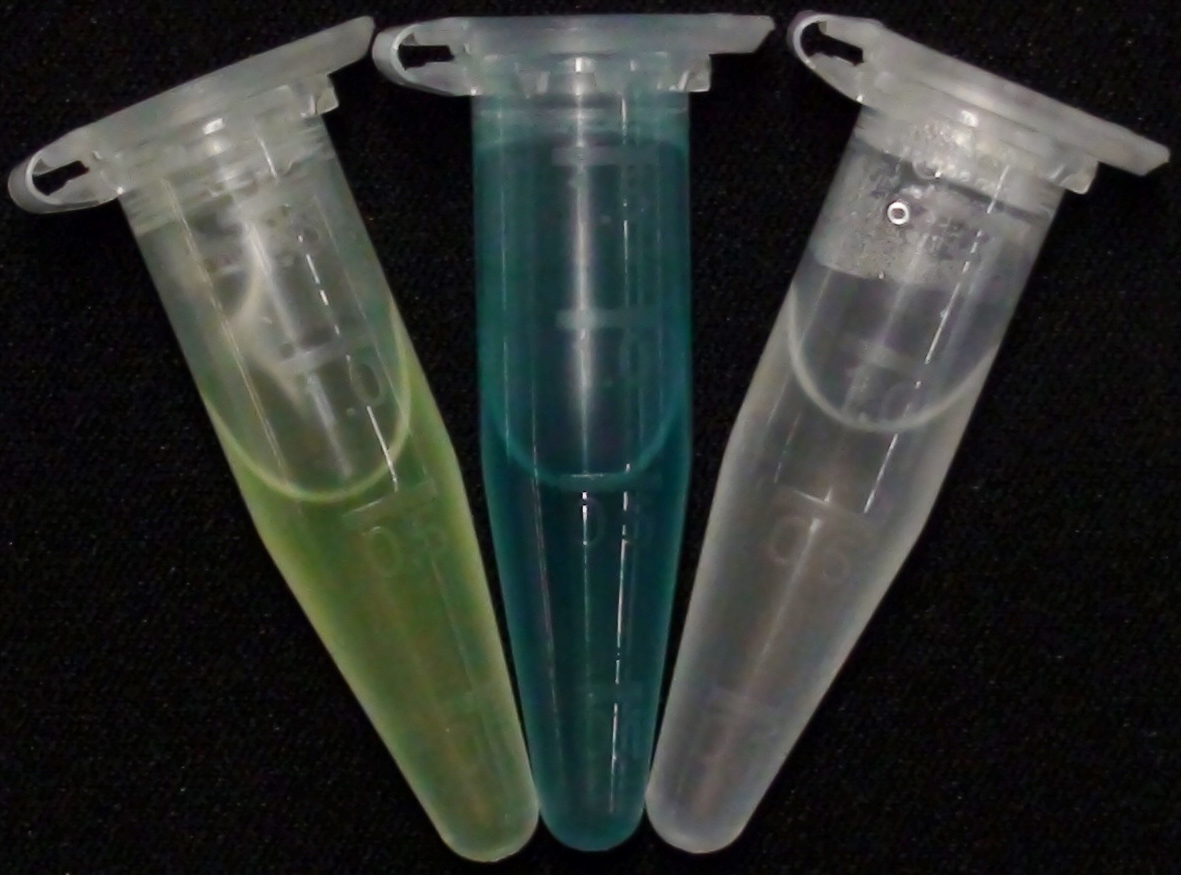
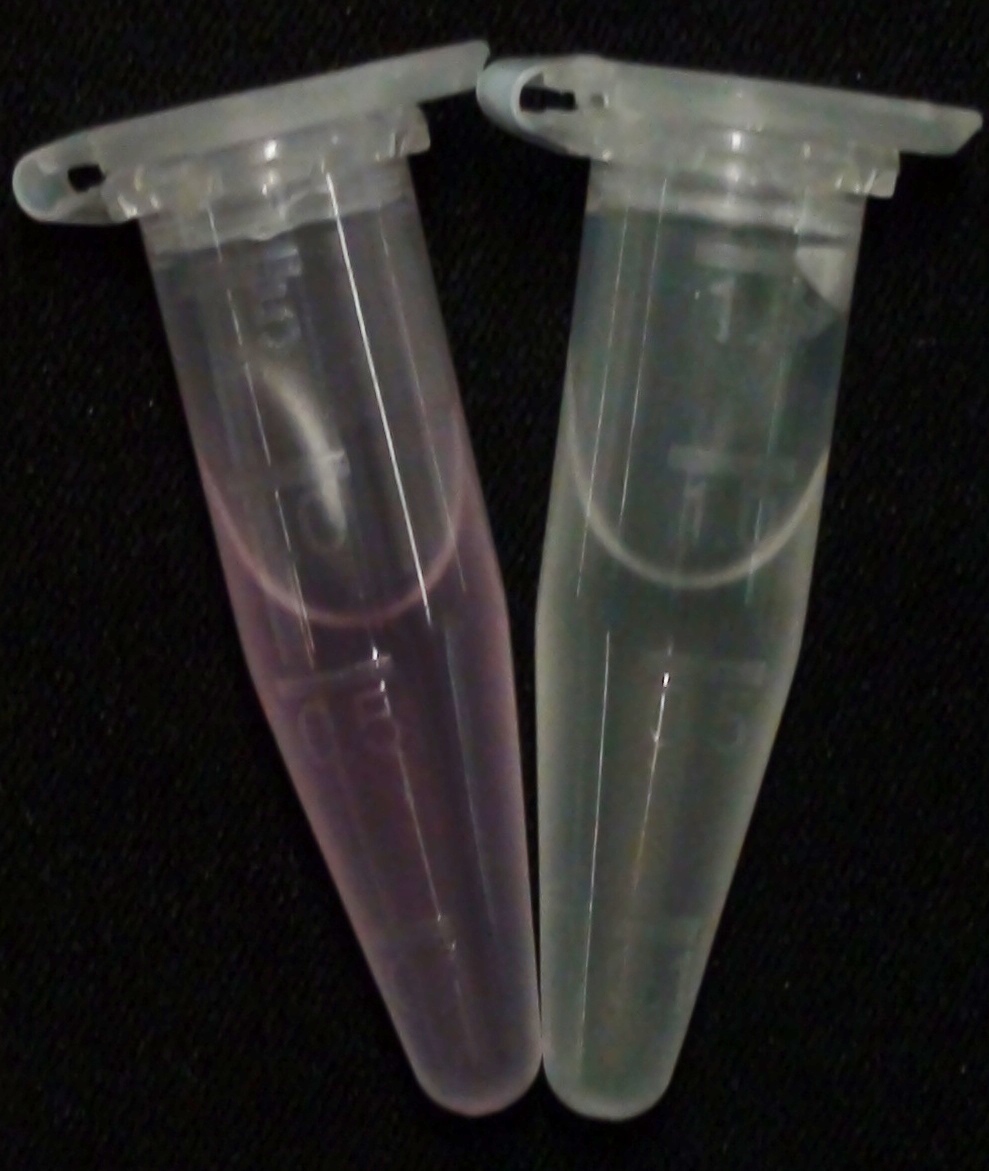
To ensure orthogonality between the enzyme-substrate reactions used in Colisweeper, a crosstalk test has been done to ensure that all overexpressed enzymes specifically cleave their assigned substrate.
Color overlay
To play Colisweeper, colors for each colony identity have to be clearly distinguishable by eye. Because we are using high-pass filters to differentially process certain AHL levels, we had to make sure that a mix of cleaved product would show distinguishable colors.
References
(1) Orenga S, Roger-Dalbert C, James A, Perry J, US Patent 20090017481 (2009).
Absorption: Biosynth
 "
"