Team:Heidelberg/Project/Indigoidine
From 2013.igem.org
m (Created page with "{{:Team:Heidelberg/Templates/Navigation}} <html lang="en"> <style type="text/css"> li { font-size:14px; } </style> <div class="container"> <!--Project D...") |
m |
||
| Line 86: | Line 86: | ||
<h2>Introduction</h2> | <h2>Introduction</h2> | ||
<p style="font-size:14px; text-align:justify"> | <p style="font-size:14px; text-align:justify"> | ||
| - | + | </html>{{:Team:Heidelberg/Templates/Indigoidine_Introduction}}<html> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</p> | </p> | ||
<h2>Experiments</h2> | <h2>Experiments</h2> | ||
Revision as of 17:55, 4 October 2013


Indigoidine. Proving Modularity of NRPS by Shuffling Domains.

Highlights
- Transfer of the whole delftibactin NRPS pathway from D. acidovorans into E. coli
- Novel approach for transfering a whole NRPS pathway more than 50 kb in size from one bacterial species into another
- Optimization of the Gibson Cloning Strategy for the creation of large plasmids (over 30 kb in size) with high GC content
- Precipitation of pure gold from electronic waste using delftibactin
Abstract
An integral characteristic of synthetic biology yet often undermined is the ability to learn fundamental knowledge by systematically perturbing a biological system. Non-ribosomal peptide synthetases (NRPS) are predestinated for such a trial and error approach. Their hierarchical organization into modules and domains offer a unique opportunity to spin around their inherent logical assembly and observe if their functionality is preserved. Following this idea, we prove the interchangeability of NRPS domains at the example of ''indC'' from ''Photorhabdus luminescens laumondii'' TT01 (DSM15139). The native NRPS domains have been replaced with domains from other bacterial organisms and fully synthetic domains. To quantify the NRPS efficiency we established an indigoidine assay based on OD measurement of the blue-colored pigment. Interestingly, we find that our data points out the dependence on the T-domain and the 4'-Phosphopanthetheinyl-transferases (PPTases), resulting in different levels of indigoidine synthesis. Furthermore, we introduce HiCT - High throughput protocols for circular polymerase extension Cloning and Transformation - a new standard for the assembly of combinatorial gene libraries (RFC 99).
Introduction
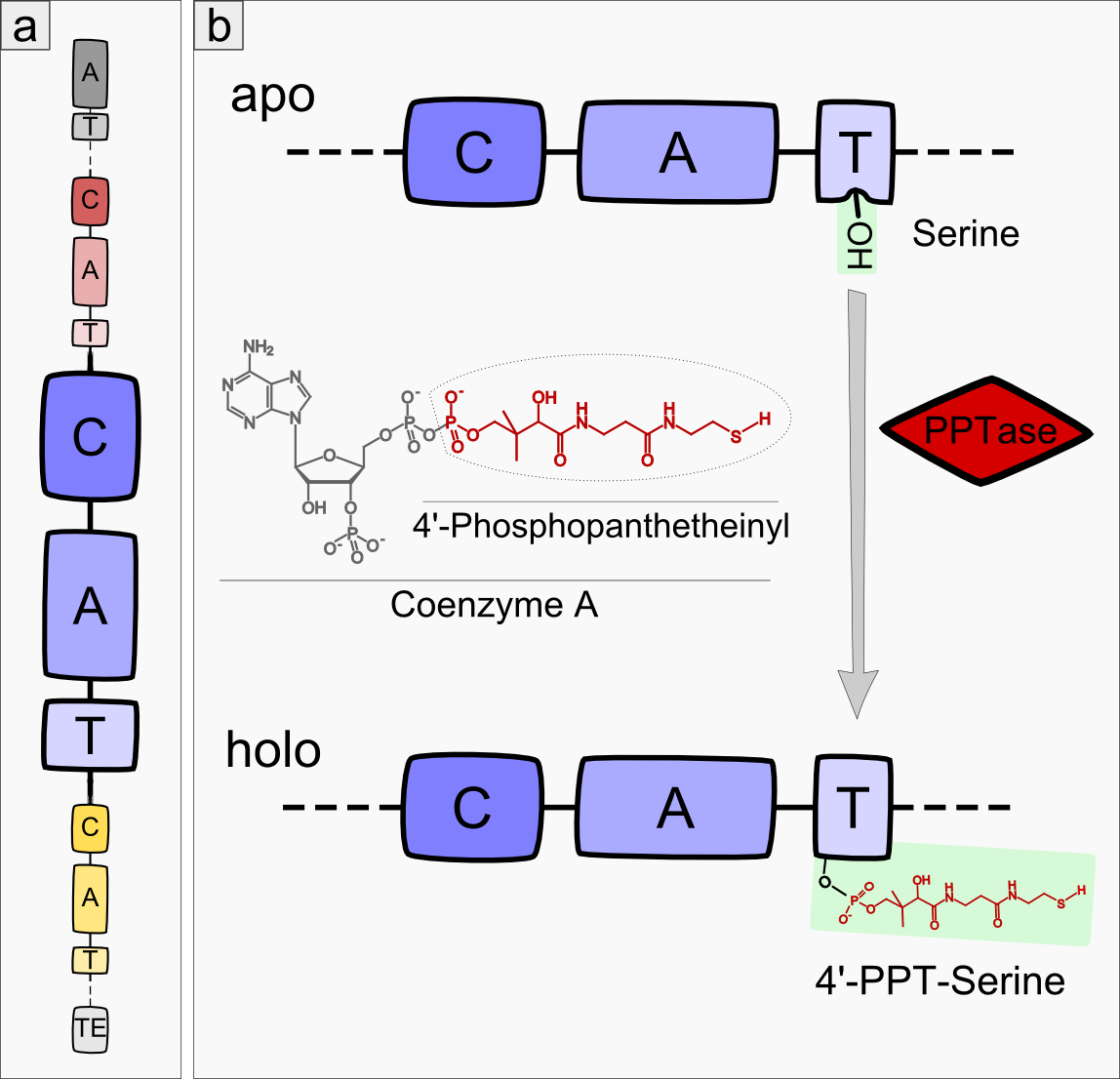
Most modules of non-ribosomal peptide synthetase (NRPS) pathways consist of three domain types: condensation, adenylation and thiolation domain (see Figure 1a), also called peptidyl-carrier-protein domain (PCP)-domain (reviewed in <bib id="pmid16895337"/>)). Remarkably, precisely this order of domains is highly conserved among different NRPS pathways except for the very first and last module of each NRPS. In the initial module, the A-domain is always followed by a T-domain. The last module of an NRPS usually ends with a TE-domain. During the process of non-ribisomal peptide (NRP) synthesis, a new amino acid is first adenylated by the A-domain and then bound to the T-domain via a thioester bond. The C-domain catalyzes the condensation of the substrate - which is bound to the T-domain of the previous module - and the amino acid of the next module. The T-domain itself shows no substrate specificity but acts as a carrier domain, which keeps the peptide attached to the NRPS module complex. The core element of every T-domain is a conserved 4’-phosphopanthetheinylated (4’-PPT) serine. The 4’-PPT residue is added by a 4’-Phosphopanthetheinyl-transferase (PPTase), which converts the NRPS apo-enzyme to its active holo-form (see Figure 1b).
 Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference].
Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference].
For example, a single module of P. luminescens laumondii TT01 (DSM15139) contains an internal oxidation domain (Ox-domain) in its A-domain and a special TE-domain (Figure 2a). This enzyme first adenylates L-glutamine (A-domain), which is then attached to the T-domain. The TE-domain cleaves and catalyzes the cyclization of the substrate, which is further oxidized by the Ox-domain. The oxidation of two cyclic glutamines results in the formation of an insoluble small molecule (Figure 2b)[Reference].
The small molecule produced by the pathway described above is a blue-colored pigment called indigoidine. Accordingly, the catalytic NRPS is referred to as indigoidine synthetase or blue pigment synthetase encoded by various bacterial strains such as S. lavendulae subsp. lavendulae (ATCC11924) or P. luminescens (<bib id="Takahashi2007"/><bib id="Brachmann2012"/>). Previous publications showed that replacing the T-domain of the blue pigment synthetase bpsA from S. lavendulae with T-domains of other NRPS modules results in a loss of function, i.e. the engineered indigoidine synthetase does not produce the blue pigment (<bib id="Owen2012"/). So far the only T-domain exchange that resulted in a functional bpsA was achieved using the T-domain of entF, - a gene involved in the enterobactin biosynthesis of E. coli. For this approach, random mutagenesis was performed to yield various mutated entF T-domains, some of which were capable to preserve the enzyme function when being introduced into bpsA (<bib id="Owen2012"/>). Other studies revealed that it is possible to exchange the A-domains of NRPS modules in B. subtilis resulting in modified non-ribosomal peptide products (<bib id="Doekel2000"/> ). Additionally, the selectivity of an NRPS module for specific amino acids could be modified by altering the conserved motif in the active site of the A-domain (<bib id="Thirlway2012"/>). Furthermore, since the endogenous 4'-Phoshopanthetheinyl-transferase (PPTase) entD from E. coli has been reported to exhibit low efficiency in activating heterologous NRPS pathways, most research in the field of NRPS involves co-expression of another PPTase (<bib id="Pfeifer2001"/>), bib id="Takahashi2007"/>).
Experiments
Our aim is to express delftibactin in E. coli. This will be achieved by introducing three different plasmids which contain parts of the delftibactin-cluster [File:Del cluster.gb] ,a Methylmalonyl-CoA pathway, a Pptase which replaces the DelC-function and a permeability device for the export of the desired NRP.

- Methylmalonyl-CoA, ppTase & permeability device
- DelH
- DelA-P - The rest of the genes of the Del-cluster Basic Strategy will be described in the following paragraphs. For further detailed experiments you can visit our LabJournal [Link to labjournal].
- Our first aim was to achieve a genomic integration of the genes that encode for components of the Methylmalonyl-CoA pathway into E. coli. The presence of this pathway is required for the production of NRPs. Because the genomic integration turned out to be more challenging then expected a new strategy was developed. Therefore, two plasmids were created (pIK2) containing MethylmalonylCoA amplified from Streptomyces coeliolor and a ppTase amplified from Bacillus subtilis in the Biobrick Backbone pSB3C5 and the permeability device (BBa_I746200) for the outer membrane of E. coli was inserted in another plasmid (pIK1). Team Cambridge revealed in 2007 that Bba_I746200 is toxic. It was itherefore inserted into pIK2 between the two terminators driven by a weak promoter (BBa_J23114) and a weak RBS (Bba_B0030), yielding pIK8 with a total size of 9,467 bp, which was inserted in DH10ß and BL21DE3 via electroporation.
-
As the gene encoding DelH alone has a size of 18 kb we decided to clone and introduce this huge gene on a separate plasmid. The first restriction enzyme strategy was problematic because of DelH amplification and the low yield in the ligation. A new GibsonAssembly-strategy was performed and DelH amplified in smaller pieces. It seemed to appear the same problem of as in the pIK1 that E. coli is selecting out the mutated DelH-constructs or is activly mutating it for toxic reasons. A plasmid was desi
 "
"











