Team:Heidelberg/Delftibactin/DelH
From 2013.igem.org
m |
Nils.kurzawa (Talk | contribs) m |
||
| Line 8: | Line 8: | ||
.methods { | .methods { | ||
margin-top:11%; | margin-top:11%; | ||
| + | } | ||
| + | p { | ||
| + | font-size:10px; | ||
} | } | ||
</style> | </style> | ||
Revision as of 19:59, 1 October 2013


Del H. This bitchy 18 kbp fragment.
Lorem ipsum dolor sit amet, consetetur sadipscing elitr, sed diam nonumy eirmod tempor invidunt ut labore et dolore magna aliquyam erat, sed diam voluptua. At vero eos et accusam et justo duo dolores et ea rebum. Stet clita kasd gubergren, no sea takimata sanctus est Lorem ipsum dolor sit amet.
Week 1
In order to transfer the gold precipitating NRPS from D. acidovorans to E.coli, the necessary modules will be amplified from the D. acidovorans genome and assembled as plasmids. Due to its large size of 18 Kb, the module DelH will be expressed on a separate plasmid. A strategy was developed, primers designed accordingly and necessary BioBricks obtained from the distribution. The D. acidovorans was obtained from the DSMZ and cultured in Acidovorax complex medium.
Week 2
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
At the beginning of the week, we could verify that the Gibson Assembly for Tripeptide I was indeed positive, however, the other Gibson Assemblies did not work properly. Instead of picking new colonies, we decided to optimize the Gibson recipe instead, as backbone religations were the most common problem. With these improved protocols, we used Gibson Assembly for the Dipeptide, Tripeptide II and Tetrapeptide I, later that week, Tetrapeptide II followed. After the Transformation to DH10β cells and screening by restriction digest we could send samples for the Dipeptide and Tetrapeptide I to sequencing and obtained a positive alignment. Hence we transformed BAP I cells with the positive constructs. The same was...
Week 3
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
At the beginning of the week, we could verify that the Gibson Assembly for Tripeptide I was indeed positive, however, the other Gibson Assemblies did not work properly. Instead of picking new colonies, we decided to optimize the Gibson recipe instead, as backbone religations were the most common problem. With these improved protocols, we used Gibson Assembly for the Dipeptide, Tripeptide II and Tetrapeptide I, later that week, Tetrapeptide II followed. After the Transformation to DH10β cells and screening by restriction digest we could send samples for the Dipeptide and Tetrapeptide I to sequencing and obtained a positive alignment. Hence we transformed BAP I cells with the positive constructs. The same was...
Week 4
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 5
At the beginning of the week, we could verify that the Gibson Assembly for Tripeptide I was indeed positive, however, the other Gibson Assemblies did not work properly. Instead of picking new colonies, we decided to optimize the Gibson recipe instead, as backbone religations were the most common problem. With these improved protocols, we used Gibson Assembly for the Dipeptide, Tripeptide II and Tetrapeptide I, later that week, Tetrapeptide II followed.
Week 6
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 7
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 8
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 9
In order to transfer the gold precipitating NRPS from D. acidovorans to E.coli, the necessary modules will be amplified from the D. acidovorans genome and assembled as plasmids. Due to its large size of 18 Kb, the module DelH will be expressed on a separate plasmid. A strategy was developed, primers designed accordingly and necessary BioBricks obtained from the distribution. The D. acidovorans was obtained from the DSMZ and cultured in Acidovorax complex medium.
Week 10
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
At the beginning of the week, we could verify that the Gibson Assembly for Tripeptide I was indeed positive, however, the other Gibson Assemblies did not work properly. Instead of picking new colonies, we decided to optimize the Gibson recipe instead, as backbone religations were the most common problem. With these improved protocols, we used Gibson Assembly for the Dipeptide, Tripeptide II and Tetrapeptide I, later that week, Tetrapeptide II followed. After the Transformation to DH10β cells and screening by restriction digest we could send samples for the Dipeptide and Tetrapeptide I to sequencing and obtained a positive alignment. Hence we transformed BAP I cells with the positive constructs. The same was...
Week 11
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
At the beginning of the week, we could verify that the Gibson Assembly for Tripeptide I was indeed positive, however, the other Gibson Assemblies did not work properly. Instead of picking new colonies, we decided to optimize the Gibson recipe instead, as backbone religations were the most common problem. With these improved protocols, we used Gibson Assembly for the Dipeptide, Tripeptide II and Tetrapeptide I, later that week, Tetrapeptide II followed. After the Transformation to DH10β cells and screening by restriction digest we could send samples for the Dipeptide and Tetrapeptide I to sequencing and obtained a positive alignment. Hence we transformed BAP I cells with the positive constructs. The same was...
Week 12
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 13
At the beginning of the week, we could verify that the Gibson Assembly for Tripeptide I was indeed positive, however, the other Gibson Assemblies did not work properly. Instead of picking new colonies, we decided to optimize the Gibson recipe instead, as backbone religations were the most common problem. With these improved protocols, we used Gibson Assembly for the Dipeptide, Tripeptide II and Tetrapeptide I, later that week, Tetrapeptide II followed.
Week 14
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 15
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 16
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 17
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Week 18
Cras justo odio, dapibus ac facilisis in, egestas eget quam. Donec id elit non mi porta gravida at eget metus. Nullam id dolor id nibh ultricies vehicula ut id elit.
Methods:
Lorem ipsum dolor sit amet, consetetur sadipscing elitr, sed diam nonumy eirmod tempor invidunt ut labore et dolore magna aliquyam erat, sed diam voluptua. At vero eos et accusam et justo duo dolores et ea rebum. Stet clita kasd gubergren, no sea takimata sanctus est Lorem ipsum dolor sit amet. Lorem ipsum dolor sit amet, consetetur sadipscing elitr, sed diam nonumy eirmod tempor invidunt ut labore et dolore magna aliquyam erat, sed diam voluptua. At vero eos et accusam et justo duo dolores et ea rebum. Stet clita kasd gubergren, no sea takimata sanctus est Lorem ipsum dolor sit amet.Lorem ipsum dolor sit amet, consetetur sadipscing elitr, sed diam nonumy eirmod tempor invidunt ut labore et dolore magna aliquyam erat, sed diam voluptua. .
29-04 - 05-05-13
Generation of Plasmid Backbones
Transformation of Biobricks
| Part of the registry | Plate of 2012 | Well | Resistance |
|---|---|---|---|
| pSB4K5 (insert=J04450) | 1 | 5G | Kanamycin |
| pSB6A1 (insert=J04450) | 1 | 1K | Ampicillin |
| lacZ (I732019) in pSB1AK3 | 4 | 12G | Kanamycin, Ampicillin |
| AraC (I0500) in pSB2K3 | 1 | 14N | Kanamycin |
| pSB1C3 (insert=J04450) | 1 | 3A | Chloramphenicol |
- Added 10 µl H2O to each well
- Incubated for 10 min at RT
- Thawed 5x chemical competent E.coli Top10
- 3 µl of plasmid DNA were added
- Incubated for 10 min on ice
- Heat shock for 40 s at 42.2°C
- Incubated for 10 min on ice
- Added 500 µl LB Medium
- Incubated at 37°C for 40 min
- Centrifuged 120 at 5,000 rpm, supernatant discarded
- Pellet resuspended in remaining medium
- Plated on plates with the corresponding antibiotics (as shown in the table above, section: resistances)
- Incubated for 2 days at RT
- One colony was picked from each plate and incubated over night at 37°C in LB medium with the antibiotic listed above
Result
All 5 parts from the Registry Distribution 2012 were successfully transformed in E.coli Top10 except for the one containing the AraC promotor.
Amplification of DelH Fragments
Design of Primers
| Identifier | Order date | Note | Sequence |
|---|---|---|---|
| DN01:DelH_f1_PacI_fw | 03-05-2013 | Amplification of DelH F1, with RBS and adding PacI restriction site | TTTT TTAATTAA TCACACAGGAAAGTACTAG ATGGACCGTGGCCGCCTGC GCCAAATCG |
| DN02:DelH_f1_SalI_rev | 03-05-2013 | Amplification of DelH F1 until SalI restriction site | TTTT GTCGACCAACACCTGTGCCTGC |
| DN03:DelH_f2_SalI_fw | 03-05-2013 | Amplification of DelH F2 starting at SalI restriction site | TTTT GTCGACTGGATGGAGCCTGGTGAAAG |
| DN04:DelH_f2_KpnI_rev | 03-05-2013 | Amplification of DelH F2, adding KpnII restriction site | TTTT GGTACC TCAGTCCAGCGCGTACTCCAG |
| DN05:AraCbb_KpnI_fw | 03-05-2013 | Amplification of backbone for DelH (pSB6A1-AraC-lacZ), adding KpnI site | TTTT GGTACC AAAGAGGAGAAATACTAGATGACCATG |
| DN08:AraCbb_PacI_rev | 03-05-2013 | Amplification of backbone for DelH (pSB6A1-AraC-lacZ), adding PacI site | TTTT TTAATTAA GCTAGCCCAAAAAAACGGGTATG |
Amplification of 2
06-05 - 12-05-13
Amplification of BioBricks
Miniprep of BioBricks
- Miniprep was performed using Qiagen kit and eluted in 35 µl ddH2O.
- From 500 µl of liquid culture, a glycerol stock was prepared as described in Preparing glycerol stocks.
Result
- DNA concentration was determined using nanovue.
| Sample | Concentration [ng/µl] |
|---|---|
| pSB4K5 | 21 |
| pSB6A1 | 47 |
| lacZ | 45 |
| pSB1C3 | 23 |
Generation of Plasmid Backbones
Restriction Digest of BioBrick pSB6A1
| Component | Amount [µl] |
|---|---|
| PSB6A1 DNA | 32.5 |
| BSA (10x) | 5 |
| NEB2 buffer (10x) | 5 |
| Enzymes (EcoRI-HF and PstI) | 1.5 each |
| ddH2O | 4.5 |
Result
The fragments of plasmids pSB4K5 and pSB1C3 show the expected pattern.
- => BioBrick backbone was cut and gel isolated.
Generation of LacZ Fragment
Restriction Digest of BioBrick pSB1AK3 Conditions A
| Component | Amount [µl] |
|---|---|
| pSB1AK3 DNA | 32.5 |
| BSA (10x) | 5 |
| NEB3 buffer (10x) | 5 |
| Enzymes (XbaI and PstI) | 1.5 each |
| ddH2O | 4.5 |
- Incubation for 1 h at 37°C
Result
- The restriction mix was loaded onto a 1% agarose gel using 6 µl gene ruler marker.
The fragments show the expected pattern.
- => LacZ insert were cut and gel isolated.
Restriction Digest of BioBrick K173004(lacZ) Conditions B
| Component | Amount [µl] |
|---|---|
| pSB1AK3 DNA | 8.3 |
| BSA (10x) | 5 |
| NEB4 buffer (10x) | 5 |
| Enzymes (XbaI & SalI-HF) | 1.5 each |
| ddH2O | 28.7 |
- Purification was performed with Qiagen Nucleotide removal Kit and eluted in 38.5 µl ddH2O
Result
2.5 µl were measured with the Nanovue. The measurement proved the failure of miniprep.
Restriction Digest of BioBrick K173004(lacZ) Conditions C
| Component | Amount [µl] |
|---|---|
| pSB1AK3 DNA | 32.5 |
| BSA (10x) | 5 |
| NEB 3 buffer (10x) | 5 |
| Enzyme (PstI) | 1.5 |
| ddH2O | - |
Result
- =>The band was too high on gel and discarded, because we cannot be sure if it really is lacZ or the backbone of the lacZ plasmid.
Generation of AraC Fragment
Amplification of BioBricks containing I13453 and K206000
- The transformation was performed as described on 03-05 and incubated ON at 37°.
Amplification of DelH F1
PCR Conditions F1.W2.A
| Reagent | DelH F1 | DelH F1 |
|---|---|---|
| Expected length [Kb] | 10 | 10 |
| Template | 1 µl D. acidovorans glycerol stock | 1 µl D. acidovorans DNA |
| Primer 100 µM fw | 0.5 µl DelH_f1_PacI_fw | 0.5 µl DelH_f1_PacI_fw |
| Primer 100 µM rev | 0.5 µl DelH_f1_SalI_rev | 0.5 µl DelH_f1_SalI_rev |
| Phusion Master Mix (2x) | 25 µl | 25 µl |
| ddH2O | 23 µl | 23 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 16 | 98 | 1 |
| 66(↓0.5°C) | 5 | |
| 72 | 3:40 min | |
| 14 | 98 | 1 |
| 62 | 5 | |
| 72 | 3:40 min | |
| 1 | 72 | 5 min |
| 1 | 4 | inf |
Result
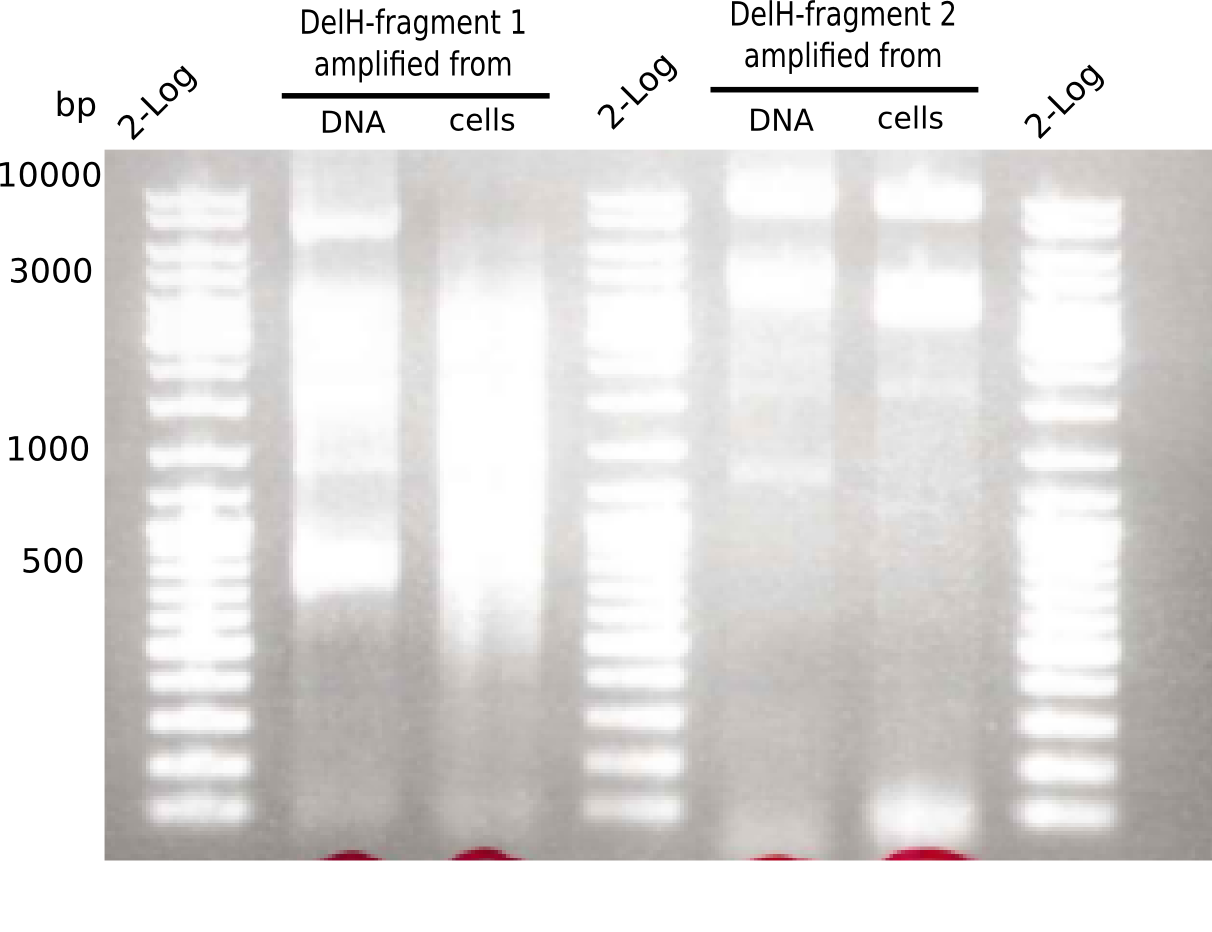
Expected band: 10 Kb
Different unspecific bands were observed.
- => Further optimize PCR conditions.
PCR Conditions F1.W2.B
| Reagent | DelH-fragment1 | DelH-fragment1 |
|---|---|---|
| Expected length [Kb] | 10 | 10 |
| Template | 1 µl D. acidovorans glycerol stock | 1 µl D. acidovorans DNA |
| Primer 100 µM fw | 0.5 µl DelH_f1_PacI_fw | 0.5 µl DelH_f1_PacI_fw |
| Primer 100 µM rev | 0.5 µl DelH_f1_SalI_rev | 0.5 µl DelH_f1_SalI_rev |
| Phusion Master Mix (2x) | 25 µl | 25 µl |
| ddH2O | 23 µl | 23 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 |
| 72 | 2:45 min | |
| 1 | 72 | 5 min |
| 1 | 4 | inf |
Result
PCRs showed only unspecific bands.
- => Further optimize PCR conditions.
PCR Conditions F1.W2.C
| Reagent | DelH F1 | DelH F1 | DelH F1 | DelH F1 |
|---|---|---|---|---|
| Expected length [Kb] | 10 | 10 | 10 | 10 |
| Template | 1 µl D. acidovorans normal | 1 µl D. acidovorans normal | 1 µl D. acidovorans 1:25 | 1 µl D. acidovorans 1:25 |
| DelH_f1_PacI_fw | 0.5 µl normal | 0.5 µl 1:10 | 0.5 µl normal | 0.5 µl 1:10 |
| DelH_f1_SalI_rev | 0.5 µl normal | 0.5 µl 1:10 | 0.5 µl normal | 0.5 µl 1:10 |
| Phusion Master Mix (2x) | 25 µl | 25 µl | 25 µl | 25 µl |
| DMSO | - | - | 2.5 µl | 2.5 µl |
| ddH2O | 23 µl | 23 µl | 20.5 µl | 20.5 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 16 | 98 | 1 |
| 62 | 5 | |
| 72 | 2:45 min | |
| 1 | 72 | 5 min |
| 1 | 4 | inf |
Result
PCR with DMSO and 1:10 diluted primer looked best.
- => Gel extraction performed.
Amplification of DelH F2
PCR Conditions F2.W2.A
| Reagent | DelH F2 | DelH F2 |
|---|---|---|
| Expected length [Kb] | 8 | 8 |
| Template | 1 µl D. acidovorans glycerol stock | 1 µl D. acidovorans DNA |
| Primer 100 µM fw | 0.5 µl DelH_f2_SalI_fw | 0.5 µl DelH_f2_SalI_fw |
| Primer 100 µM rev | 0.5 µl DelH_f2_KpnI_rev | 0.5 µl DelH_f2_KpnI_rev |
| Phusion Master Mix (2x) | 25 µl | 25 µl |
| ddH2O | 23 µl | 23 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 16 | 98 | 1 |
| 66(↓0.5°C) | 5 | |
| 72 | 3:40 min | |
| 14 | 98 | 1 s |
| 62 | 5 s | |
| 72 | 3:40 min | |
| 1 | 72 | 5 min |
| 1 | 4 | inf |
Result
Expected band: 8 Kb
Different unspecific bands were observed.
- => Further optimize PCR conditions.
PCR Conditions F2.W2.B
| Reagent | DelH F2 | DelH F2 |
|---|---|---|
| Expected length [Kb] | 8 | 8 |
| Template | 1 µl D. acidovorans glycerol stock | 1 µl D. acidovorans DNA |
| Primer 100 µM fw | 0.5 µl DelH_f2_SalI_fw | 0.5 µl DelH_f2_SalI_fw |
| Primer 100 µM rev | 0.5 µl DelH_f2_KpnI_rev | 0.5 µl DelH_f2_KpnI_rev |
| Phusion Master Mix (2x) | 25 µl | 25 µl |
| ddH2O | 23 µl | 23 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 s |
| 72 | 2:45 min | |
| 1 | 72 | 5 min |
| 1 | 4 | inf |
Results
Expected band: 8 Kb
Gel shows the expected specific band at 8 Kb.
- => DelH F2 was thus successfully amplified from the glycerol stock.
Amplification of 1
13-05 - 19-05-13
Amplification of DelH F1
Gel Purification
45 µl of DelH F1 PCR (08-05) product were run on a 0,8% agarose gel (1 h with 135V). Gel extraction was performed using gel extraction kit of Qiagen, and diluted in 25 µl ddH2O.
Result
| Sample | Concentration [ng/µl] |
|---|---|
| DelH F1 | 7 |
- => Because the concentration is so low, a re-PCR of the PCR product (08-05) is going to be done.
re-PCR Conditions F1.W3.A
| Reagent | DelH F1 |
|---|---|
| Expected length [Kb] | 10 |
| Template | 1 µl 1:10 gel purified F1 |
| Primer 10 µM fw | 0.5 µl DelH_f1_PacI_fw |
| Primer 10 µM rev | 0.5 µl DelH_f1_SalI_rev |
| Phusion Master Mix (2x) | 25 µl |
| DMSO | 2.5 µl |
| ddH2O | 20.5 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 |
| 66 | 5 | |
| 72 | 3:30 min | |
| 72 | 3:30 min | |
| 1 | 4 | inf |
Result
Expected length: 10 Kb
Different unspecific bands were observed.
- => Test digest to confirm identity.
Test Restriction Digest
| PCR product of DelH-F1 | BSA | NEBuffer 4 | Enzymes | ddH2O | Total amount |
|---|---|---|---|---|---|
| 18 µl | 5 µl | 5 µl | 2x 1.5 µl SalI-HF & PacI | 19 µl | 50 µl |
Afterwards, purification with the nucleotide removal kit was performed, but due to using wrong column, there was no product left.
- => Repeat amplification of DelH F1.
PCR Conditions F1.W3.B
| Reagent | DelH F1 | DelH F1 | DelH F1 |
|---|---|---|---|
| Expected length [Kb] | 10 | 10 | 10 |
| Template | Picked colony | Picked colony | Picked colony |
| Primer 10 µM fw | 0.5 µl DelH_f1_PacI_fw | 0.5 µl DelH_f1_PacI_fw | 0.5 µl DelH_f1_PacI_fw |
| Primer 10 µM rev | 0.5 µl DelH_f1_SalI_rev | 0.5 µl DelH_f1_SalI_rev | 0.5 µl DelH_f1_SalI_rev |
| Phusion Master Mix (2x) | 25 µl | 25 µl | 25 µl |
| DMSO | 1.5 µl | 2.5 µl | 5 µl |
| ddH2O | 21.5 µl | 20.5 µl | 18 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 3:30 min | |
| 1 | 72 | 3:30 min |
| 1 | 4 | inf |
Result
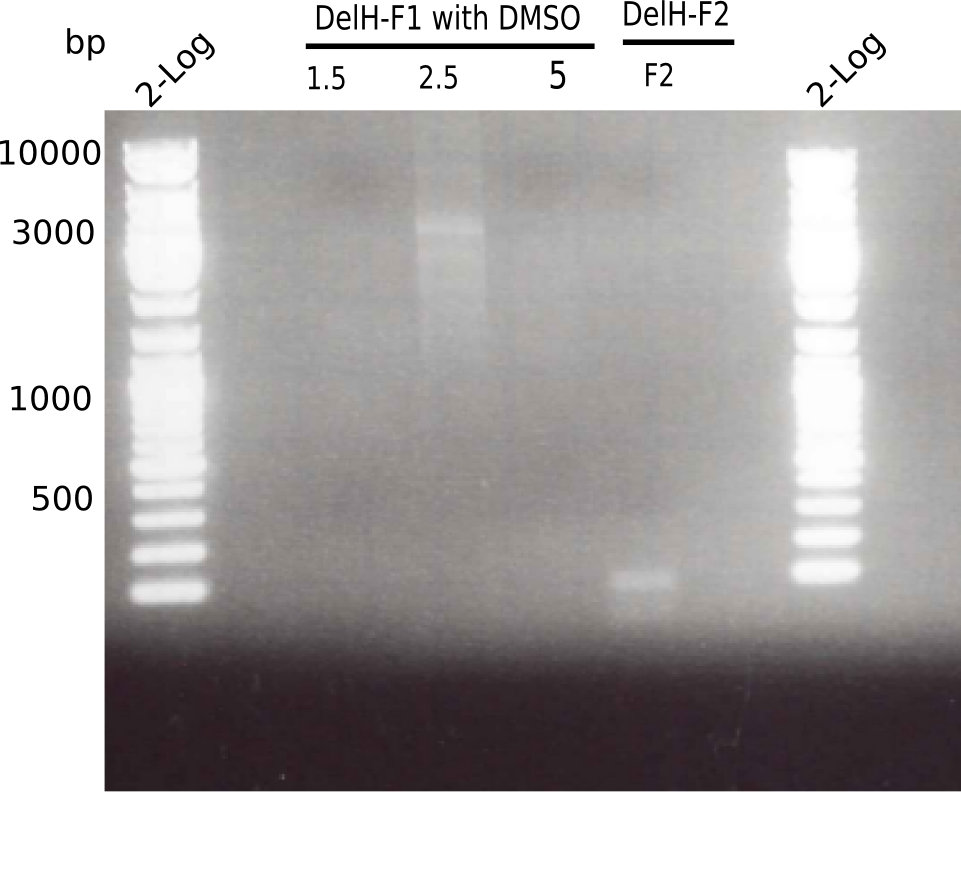
Expected length: 10 Kb
There is only an unspecific band at the second PCR of F1 with 2.5 µl DMSO.
- => Repeat using DMSO and altered PCR program.
PCR Conditions F1.W3.C
| Reagent | DelH F1 | DelH F1 |
|---|---|---|
| Expected length [Kb] | 10 | 10 |
| Template | 1 µl glycerol stock | 1 µl glycerol stock |
| Primer 10 µM fw | 0.5 µl DelH_f1_PacI_fw | 0.5 µl DelH_f1_PacI_fw |
| Primer 10 µM rev | 0.5 µl DelH_f1_SalI_rev | 0.5 µl DelH_f1_SalI_rev |
| Phusion Master Mix (2x) | 25 µl | 25 µl |
| DMSO | 2.5 µl | 2.5 µl |
| ddH2O | 20.5 µl | 20.5 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 3:30 min | |
| 1 | 72 | 3:30 min |
| 1 | 4 | inf |
Result
Expected length: 10 Kb
There was no band visible.
- => Repeat using DMSO and altered PCR program.
Amplification of DelH F2
Gel Purification
45 µl of DelH F1 PCR (07-05) product were run on a 0,8% agarose gel (1 h with 135V). Gel extraction was performed using gel extraction kit of Qiagen, and diluted in 25 µl ddH2O.
Result
| Sample | Concentration [ng/µl] |
|---|---|
| DelH F2 | 6 |
- => Because the concentration is so low, a re-PCR of the PCR product (07-05) is going to be done.
Re-PCR Conditions F2.W3.A
| Reagent | DelH-fragment2 |
|---|---|
| Expected length [Kb] | 8 |
| Template | 1 µl 1:10 gel purified F2 |
| Primer 10 µM fw | 0.5 µl DelH_f2_SalI_fw |
| Primer 10 µM rev | 0.5 µl DelH_f2_KpnI_rev |
| Phusion Master Mix (2x) | 25 µl |
| DMSO | 2.5 µl |
| ddH2O | 20.5 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 3:30 min | |
| 72 | 3:30 min | |
| 1 | 4 | inf |
Result
Expected length: 8 Kb
Specific band was observed.
- => Test digest to confirm identity.
Test Restriction Digest
| PCR product of DelH-F1 | BSA | NEBuffer 4 | Enzymes | ddH2O | Total amount |
|---|---|---|---|---|---|
| 18 µl | 5 µl | 5 µl | 2x 1.5 µl SalI-HF & KpnI-HF | 19 µl | 50 µl |
Afterwards, a purification with the nucleotide removal kit was performed, but due to using wrong column, there was no product left.
- => Repeat amplification of DelH F2.
PCR Conditions F2.W3.B
| Reagent | DelH F2 |
|---|---|
| Expected length [Kb] | 8 |
| Template | Picked colony |
| Primer 10 µM fw | 0.5 µl DelH_f2_SalI_fw |
| Primer 10 µM rev | 0.5 µl DelH_f2_KpnI_rev |
| Phusion Master Mix (2x) | 25 µl |
| DMSO | 0 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 |
| 66 | 5 | |
| 72 | 2:45 min | |
| 1 | 72 | 2:45 min |
| 1 | 4 | inf |
Result
Expected length: 8 Kb
There is no band visible.
- => Repeat using altered PCR program.
PCR Conditions F2.W3.B
| Reagent | DelH F2 |
|---|---|
| Expected length [Kb] | 8 |
| Template | 1 µl glycerol stock |
| Primer 10 µM fw | 0.5 µl DelH_f2_SalI_fw |
| Primer 10 µM rev | 0.5 µl DelH_f2_KpnI_rev |
| Phusion Master Mix (2x) | 25 µl |
| DMSO | 0 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 |
| 66 | 5 | |
| 72 | 2:45 min | |
| 1 | 72 | 2:45 min |
| 1 | 4 | inf |
Result
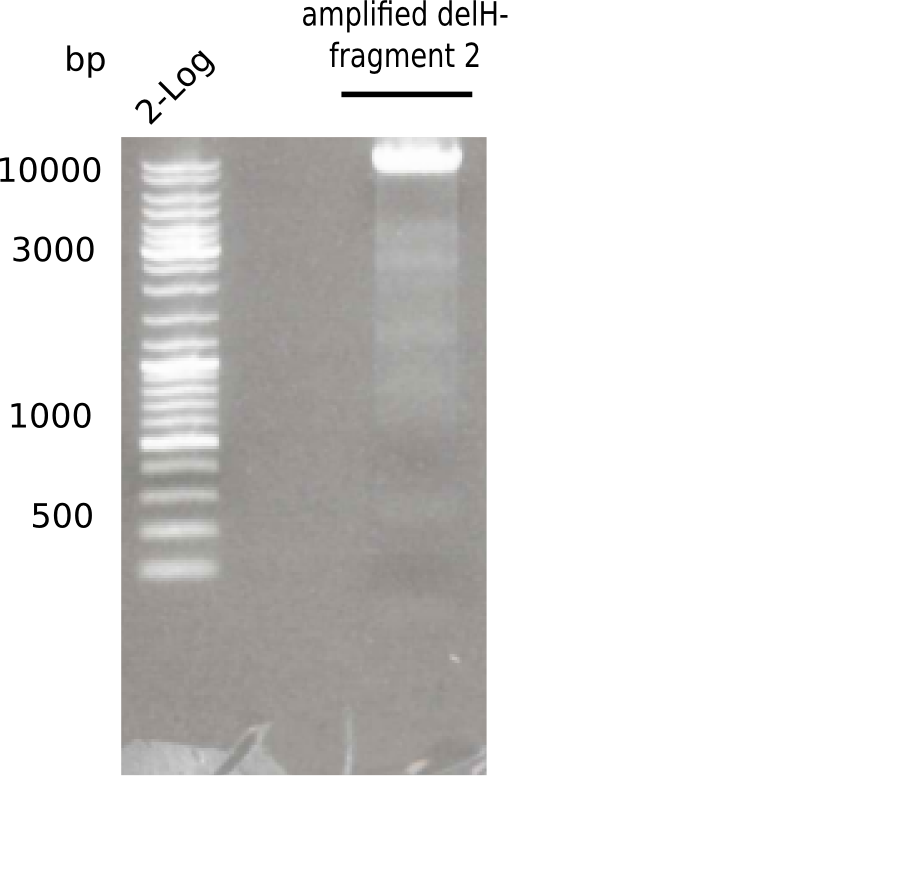
Expected length: 8 Kb
There was a specific band at 8 Kb
- => Fragment was cut and gel extracted.
Generation of Backbone pSB6A1-AraC-lacZ
Miniprep of Amplified Parts
DH10ß containing I13453 and K20600 (for AraC) and backbone pSB1AK3 (lacZ) were grown ON at 37°C and minipreped.
Result
| Sample | Concentration [ng/µl] |
|---|---|
| lacZ | 42.5 |
| AraC (5) | 26.5 |
| AraC (6) | 34.5 |
Restriction Digest
- AraC was cut with EcoRI-HF and SpeI buffered in NEB4
- LacZ was cut with XbaI and PstI buffered in NEB3
| Reagent | Amount [µl] |
|---|---|
| DNA | 22.5 |
| Enzymes | 1.5 each |
| Buffer (10x) | 5 |
| BSA (10x) | 5 |
| ddH2O | 14.5 |
- Incubated for 1h at 37°C
- Purified with Qiagen Kit and diluted in 20 µl ddH2O
Result
| Sample | Concentration [ng/µl] |
|---|---|
| lacZ | 33.5 |
| AraC (5) | 19.5 |
| AraC (6) | 25.5 |
Ligation of pSB6A1, AraC and lacZ
| Reagent | Amount [µl] |
|---|---|
| pSB6A1 | 1.5 |
| lacZ | 4 |
| AraC | 3 |
| Ligase | 1.5 |
| Buffer | 2 |
| ddH2O | 8.5 |
- Incubated at RT for 45 min
- Chemical transformation of competent DH10ß
- Streaked on LB Amp plates
- Incubation ON at 37°C
Miniprep of pSB6A1-AraC-lacZ
- One colony each was picked and grown in 2 ml LB Amp ON
- Cultures were minipreped.
Result
| Sample | Concentration [ng/µl] |
|---|---|
| pSB6A1-AraC-lacZ (5) | 57 |
| pSB6A1-AraC-lacZ (6) | 73 |
Restriction Digest of Backbone pSB6A1-AraC-lacZ
- AraC-lacZ was cut from pSB6A1-AraC-lacZ using PstI & EcoRI-HF
| Reagent | Amount [µl] |
|---|---|
| Miniprep DNA | 10 |
| Enzymes | 1.5 each |
| Buffer NEB 2(10x) | 5 |
| BSA (10x) | 5 |
| ddH2O | 27 |
- Incubated for 1 h at 37°C
- Purified with Qiagen kit and diluted in 20 µl ddH2O
Result
| Sample | Concentration [ng/µl] |
|---|---|
| pSB6A1-AraC-lacZ (5) D+P | 18 |
| pSB6A1-AraC-lacZ (6) D+P | 26 |
Ligation of AraC-lacZ with pSB1C3
| Reagent | Amount [µl] |
|---|---|
| DNA of pSB1C3 | 10 |
| DNA of AraC-lacZ (6) | 5 |
| Ligase | 1 |
| Buffer | 2 |
| ddH2O | 2 |
- Incubated at RT for 50 min
- Heat inactivation at 70°C for 5 min
- 10 µl were used for a chemical transformation in TOP10 and plated on LB Chlor
- Incubation ON at 37°C
Conlony-PCR Conditions BB.W3.A
| Reagent | psB1C3-AraC-lacZ (5) |
|---|---|
| Expected length [Kb] | ? |
| Template | Picked colony of psB1C3-AraC-lacZ (5) |
| Primer 10 µM fw | 0.5 µl |
| Primer 10 µM rev | 0.5 µl |
| Phusion Master Mix (2x) | 25 µl |
| DMSO | - |
| ddH2O | 24 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 10 |
| 30 | 98 | 1 |
| 66 | 5 | |
| 72 | 1:30 min | |
| 1 | 72 | 1:30 min |
| 1 | 4 | inf |
Result
Expected length:
There was no fragments visible.
- => Is picked coloniy negative or did entire ligation not work out?
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
Amplification of 1
Something completely different
test
 "
"