Team:Heidelberg/NRPS
From 2013.igem.org
Nils.kurzawa (Talk | contribs) |
Nils.kurzawa (Talk | contribs) |
||
| Line 203: | Line 203: | ||
}); | }); | ||
$(".fancybox.fancyFigure").fancybox({ | $(".fancybox.fancyFigure").fancybox({ | ||
| - | |||
prevEffect : 'none', | prevEffect : 'none', | ||
nextEffect : 'none', | nextEffect : 'none', | ||
helpers : { | helpers : { | ||
title : { | title : { | ||
| - | type: ' | + | type: 'outside' |
}, | }, | ||
thumbs : { | thumbs : { | ||
Revision as of 13:50, 28 October 2013


NRPS. Get to Know the Theory.
Abstract
Everybody knows the dogma of molecular biology - DNA makes RNA makes protein, but it doesn't cover all nature's capabilities, as for example non-ribosomal peptide synthetases, or short NRPS. This alternative pathway for peptide formation is mainly found in bacteria and fungi, but can at also be functionally transferred to mamallian cells. Natural NRPs can have various functions from simple dyes up to metal chelators or antibiotics. NRPSs are large protein complexes adding amino or arylic acids, taken from a pool of more than 500 monomers, to a growing peptide chain without the need for a template. The molecular structure of NRPS is based on modules, each incorporating exactly one monomer and comprising several domains of different functions. The most important domains are the adenylation (specific substrate activation), the thiolation (peptide / monomer carrier) and the condensation domain (peptide bond formation), which are needed to form basic peptide chains. Besides these various other domains are known to introduce secondary modifications. Thanks to their remarkable modularity and the wide range of substrates, NRPSs bear the impressive biosynthetic potential to create novel non-ribosomal peptides of various functions in vivo or in vitro. The range of possible applications exceeding natural functions remains unforseen.
Modular Structure
The biggest benefit of NRPS for synthetic biology is it's very modular structure. This starts at the gene level and goes as deep as domain or even single residue level. This modular organisation can be exploited by reorganisation of the modules in order to achieve different products.

Figure 1: Modular organisation of NRPS.Non-ribosomal peptide synthetases can be subdivided in modules each incorporating one amino acid. Those are comprised of several different domains, here condensation (C), adenylation (A), thiolation (T) and N-methylation (NM).
Biosynthetic Gene Cluster
The genes necessary for the biosynthesis are normally organised in a gene cluster. This contains the genes coding for the non-ribosomal peptide synthetases, which can reach a size of up to 1.5 MDa (cyclosporin Fischbach), genes necessary for the monomers' biosynthesis, as well as tailoring enzymes introducing further modifications in the peptide. There is normally more than just one peptide synthetase. The proteins are often connected by communication domains in order to keep the structure of the assembly line. If one wants to transfer a whole pathway to a different host organisms one crucial apect, besides the successful cloning of the synthetases, is the monomer supply. Depending on the hosts's endogenous machinery one can leave out certain genes or has to include other pathways in order to keep up the supply.
One Module - One Amino Acid
Each NRP synthetase is organised in so called modules, where every single module is responsible for incorporating one amino acid in the growing peptide chain. Since all modules have similar minimal structure components one can reorder them easily to achive a different product. These minimal structure components are NRPS specific protein domains each providing the assembly line with a different function. The two levels of modularity - modules and domains are shown in figure 1.(Fischbach)
Chain Elongation
The mechansim of the peptide bond formation in NRPS is partially different from that in the ribosome. The biggest differences are the attachment of the growing peptide chain and the number of catalytic domains. In ribosomal synthesis one ribosome can add many amino acids, but in non-ribosomal synthesis the number of catalytic domains rises linearly with the number of amino acids incorporated. Thus the latter is only suitable for oligopeptide synthesis.
Thiolation Domain and PPTases
In the ribosome the growing peptide chain and the monomers to be incorporated are bound to tRNA, but never to the ribosome itself. NRP synthetases contain thiolation domains (T-domains) in every module, where the corresponding amino acid is covalently bound to the enzyme via a thioester bond. The essential amino acid residue of the T-domains is a serine which is posttranslationally modified to carry the sulfhydryl group required for the thioester bond. This modification is carried out by separate proteins - the phosphopantetheinyltranferases (PPTases), which use coenzyme A as cofactor. The reaction is shown in Fig. 2a). As the functionality of the thiolation domains is essential for the peptide synthesis one should consider transferring a suitable PPTase to a host organism together with the synthetases.
Adenylation Domain
The actual attachment of the amino acid to the already modified T-domain is carried out by the adenylation domain (A-domain). It is highly substrate specific for only a single monomer, besides some rare cases when it can also bind a second, very similar monomer. During the reaction the the monomer is activated with ATP in a first step and in a second one the tioester bond between the phosphopantetheinyl residue and the monomer's carboxylic acid residue is formed. The reaction is depicted in Fig. 2b).
Condensation Domain
After two neighbouring monomers have been activated, the condensation domain (C) is the one to form the peptide bond, which is shown in Fig. 2c). The C-domain is selective for the acceptor amino acid and thus one couple of C and A-domain always have the same substrate specificity. The reaction catalysed by the C-domain is a nucleophilic attack of the acceptor amino acid on donor peptide chain.
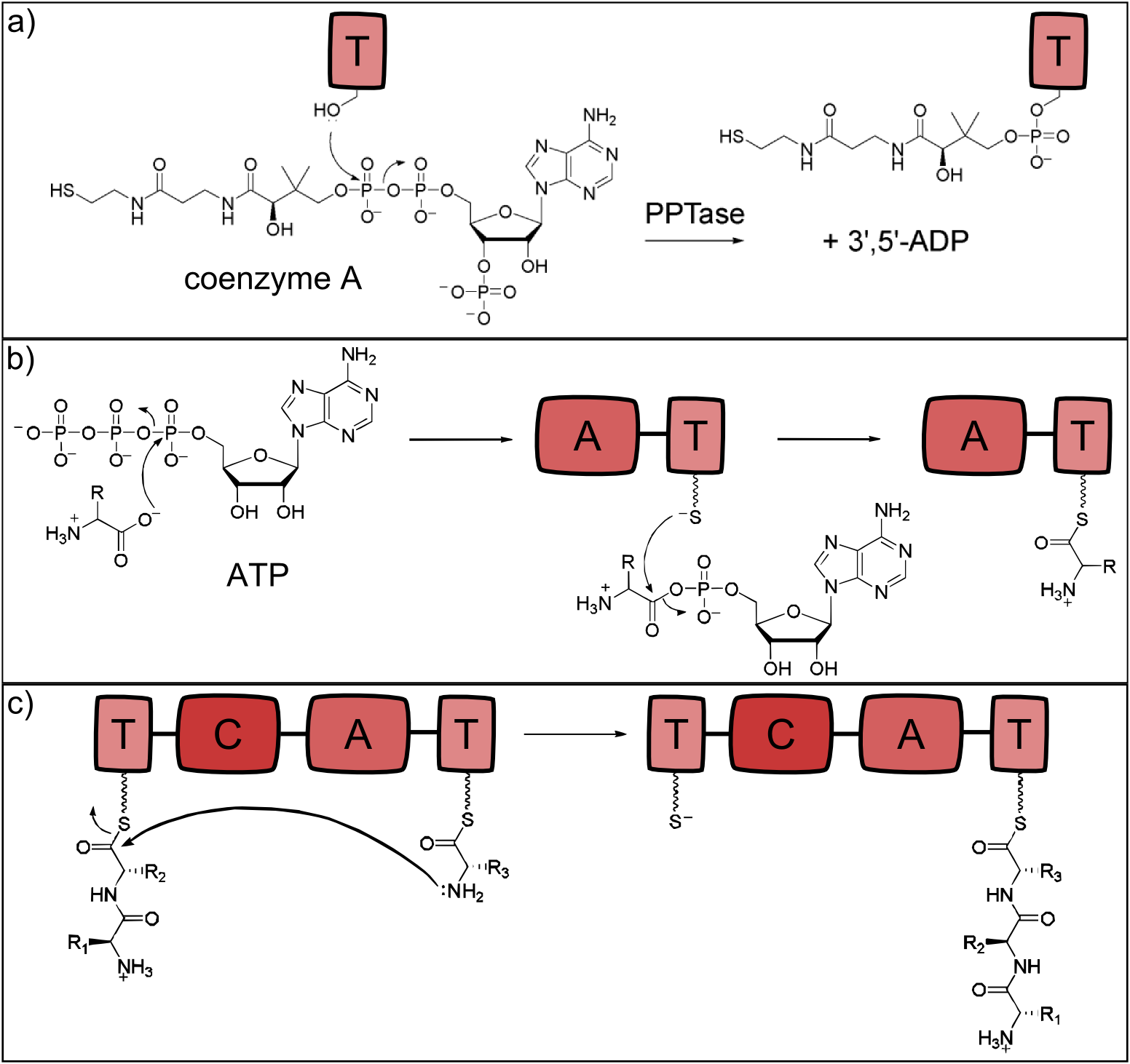
Figure 2: Reactions catalysed by the three basic NRPS domains. a) Thiolation (T) domains are the carrier domains of the monomers and the growing peptide chain. The transfer of phosphopantetheine from coenzyme A to a conserved serine in the T-domain is essential for non-ribosomal peptide synthesis as the thiole residue is necessary for monomer binding. b) The adenylation (A) domain selects the amino acids and covalently binds it to the T domain. It catalyzes two reactions: First, the activation of the monomer by ATP-binding and second, the acylation to the downstream T domain. c) Chain elongation is catalyzed by condensation (C) domains. The C domain enables peptide bond formation between the monomer (acceptor) and the growing peptide chain (donor), resulting in a translocation of the peptide chain to the acceptor T domain (adapted from [1]).
TErmination - ThioEsterase
In order to terminate a pathway a thioesterase (TE) is located at the end of most NRPS. This can have one out of several exact functions. These can roughly be classified in simple product cleavage or macrocyclisation. The first group cleaves the thioester bond between the product and the enzyme complex by transfering the peptide to it's own conserved serine residue and then releases it to the cytoplasm. The second group introduces a macrocylce in the product after cleaving it off the last T-domain. This macrocylce can either connect the two peptide termini or introduce any other peptide bond based cycle. In addition to those integral TE domains separated thioesterases are known to function as a rescue protein for stalled NRPS.
Monomer Modifications
As already mentioned in the overview NRPS can select monomers out of more than 500 different ones. These are either amino acids or arylic acids, which normally lack the amine residue and are thus only suitable for chain initiations. Most of the monomers are derived from more basic ones.
Epimerisation and Stereoselectivity
The most basic derivation is the introduction of D-amino acids. This is achieved by epimerisation (E) domains located between the T-domain of the donor and the C-domain of the acceptor module (see Fig. 3). It racemises the donor amino acid and the correct stereoconformation is selected by the acceptor C-domain. Thus the C-domains can be classified as either CD or CL.

Figure 5: A combination of epimerisation (E) and C domains allow for the incorporation of L-amino acids into NRPs. First, the E domain racemises the donor amino acid. Subsequently, a C domain specific for D-amino acids catalyzes the condesation reaction (lower reaction). For the L-stereoconformation, no peptide bond can be formed (upper reaction;adapted from [1]).
N-Methylation
The next basic modification of the peptides is the N-methylation carried out by N-methyltransferases (NM). They are located between the A and the T-domain of a module. The transfer CH3 from S-adenosylmethionine to the amino group of the module's monomer after it has been activated by the A-domain.
Heterocyclisation
Besides the macrocyclisation performed by TE-domains cyclisation (Cy) domains can introduce additional amide bonds between neighbouring amino acids in the middle of the assembly line. They need either serine, threonine or cystein as acceptor amino acid in order to form a heterocycle.
All of these modifying domains have in common, that the border to the C or A-domains are often very blurry. The Cy-domains and E-domains are thus often displayed as combined C/Cy or C/E-domains. Besides these most common modifications many others can be found within the pathway or as tailoring enzymes. These can for example be O-methylations, halogenations, glycosylations or oxidations - there is a lot more to explore!
1. Fischbach M, Walsh C (2006) Assembly-Line Enzymology for Polyketide and Nonribosomal Peptide Antibiotics: Logic, Machinery, and Mechanisms. Chemical Reviews: 106, 3468−3496.
 "
"









