Team:NTU-Taida/Notebook/Journal
From 2013.igem.org
(Difference between revisions)
(→7/8~7/12) |
(→7/8) |
||
| Line 77: | Line 77: | ||
Construct the circuit for new receptor CinR | Construct the circuit for new receptor CinR | ||
==7/8== | ==7/8== | ||
| + | ===Sequence:=== | ||
| + | ACB-4, ACE-5, ACG-1, DFB-1, DFE-2, DFE-3, DFG-1, RhlE-2, RhlG-1, RhlG-2, LasB3, LasE1, LasG4 | ||
| + | ===Primer list:=== | ||
| + | <html> | ||
| + | <table> | ||
| + | <tr class="tableizer-firstrow"><th>Primer order</th><th>Name</th><th>sequence</th><th> </th></tr> | ||
| + | <tr><td>BMRC-120</td><td>iGEM2013-pSB1A2</td><td>attaccgcctttgagtgagc</td><td>R</td></tr> | ||
| + | <tr><td>BMRC-153</td><td>iGEM2013-pSB1A2</td><td>gtgccacctgacgtctaagaa</td><td>F</td></tr> | ||
| + | <tr><td>BMRC-122</td><td>iGEM2013-mCherry</td><td>gccgtcctcgaagttcatcac</td><td>mcherry</td></tr> | ||
| + | <tr><td>BMRC-124</td><td>iGEM2013-mRFP</td><td>aacggtaacaccaccgtc</td><td>RFP</td></tr> | ||
| + | <tr><td>BMRC-126</td><td>iGEM2013-GFP</td><td>cttgtagttcccgtcatcttt</td><td>GFP</td></tr> | ||
| + | </table> | ||
| + | </html> | ||
| + | ===Digestion, ligation, transform: 3A assembly and standard assembly both=== | ||
| + | |||
| + | B0030+CinR | ||
==7/9== | ==7/9== | ||
Revision as of 01:12, 3 September 2013
Contents |
July
7/1~7/5
Finish RhlR and LasR circuit.
7/1
Transform:
- Positive feedback circuit:
Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G; - Control:
PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G;
(A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
- New:
Pcin, CinR (resistant=C)
7/2
Result:
There’s no colonies at plates of Pcin and CinR.
Transform:
Pcin(resistant=A), CinR (resistant=K)
Inoculation and incubation at LB broth
- Positive feedback circuit: Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G;
- Control: PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G; (A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
7/3
Inoculation and incubation at LB broth
Pcin(resistant=A), CinR (resistant=K)
Check
- Positive feedback circuit: Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G;
- Control: PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G; (A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
7/4
Check and Plasmid DNA extraction (mini-prep):
- Positive feedback circuit: Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G;
- Control: PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G; (A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
- Pcin(resistant=A), CinR (resistant=K)
7/5
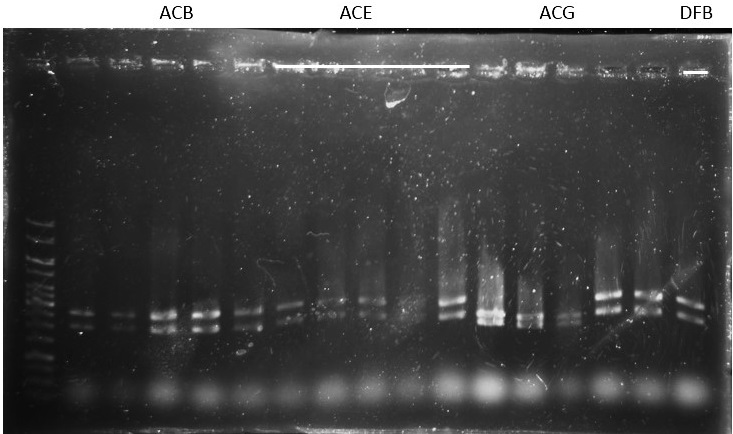
Check: (for plasmid ready to be sequenced)
ACB-4, ACE-5, ACG-1, DFB-1, DFE-2, DFE-3, DFE-4, DFE-5, DFG-1, RhlB-1, RhlE-2, RhlG-1, RhlG-2, LasB3, LasE1, LasG4 (8,9,11 well failed :DFE-4/DFE-5/ RhlB-1)
7/8~7/12
Construct the circuit for new receptor CinR
7/8
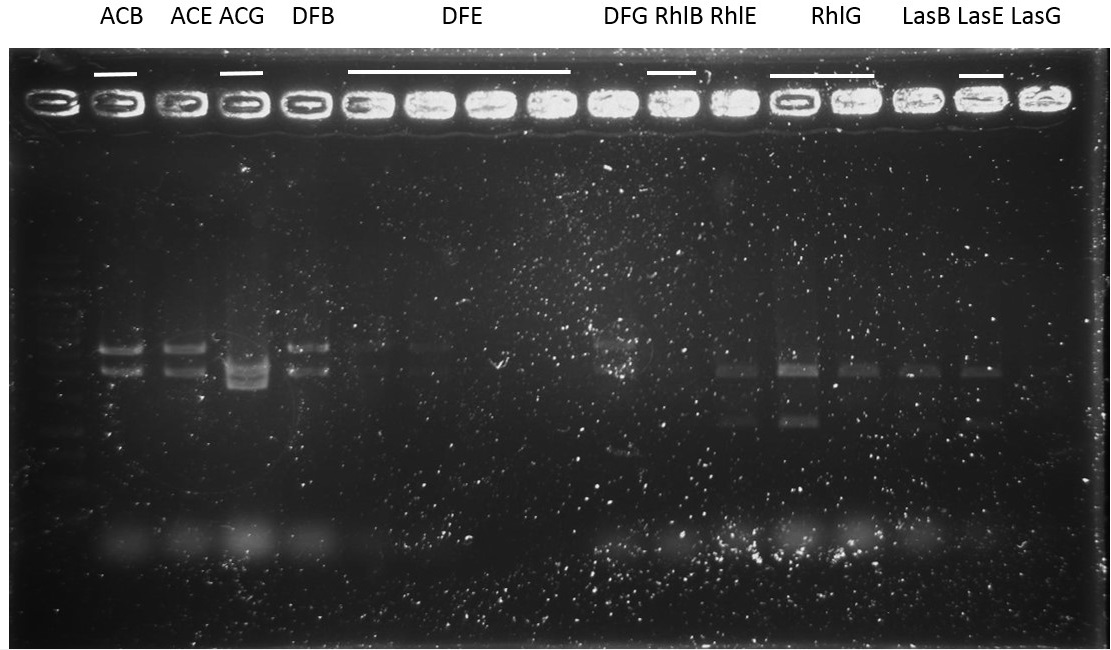
Sequence:
ACB-4, ACE-5, ACG-1, DFB-1, DFE-2, DFE-3, DFG-1, RhlE-2, RhlG-1, RhlG-2, LasB3, LasE1, LasG4
Primer list:
| Primer order | Name | sequence | |
|---|---|---|---|
| BMRC-120 | iGEM2013-pSB1A2 | attaccgcctttgagtgagc | R |
| BMRC-153 | iGEM2013-pSB1A2 | gtgccacctgacgtctaagaa | F |
| BMRC-122 | iGEM2013-mCherry | gccgtcctcgaagttcatcac | mcherry |
| BMRC-124 | iGEM2013-mRFP | aacggtaacaccaccgtc | RFP |
| BMRC-126 | iGEM2013-GFP | cttgtagttcccgtcatcttt | GFP |
Digestion, ligation, transform: 3A assembly and standard assembly both
B0030+CinR
7/9
7/10
7/11
7/12
7/15~7/19
7/15
7/16
7/17
7/18
7/19
7/22~7/31
7/22
7/23
7/24
7/25
7/26
7/27
7/28
7/29
7/30
7/31
July
2
 "
"