Final Circuit
For the final Colisweeper circuit we plan a four plasmid system. The mine cells constitutively express LuxI for signal generation and NagZ as identifier hydrolase. In the non-mine cells LuxR is expressed constitutively to process the OHHL signal. PhoA is expressed constitutively as well as reporter for safe cells. Aes and GusA are expressed from pLux promoters with different sensitivities. You can find all the biobricks we used and our own new biobricks in the figure below.

Figure 1. Plasmids in mine and non-mine cells: move the cursor over the separate parts to check which biobricks we used.
Cloned Constructs
To get to the circuit mentioned above we tested different versions of the circuit. For example we started our experiments using GFP as a reporter instead of the hydrolases. Then we also tested different LuxI and LuxR generating constructs. In the following table we list all the biobricks we used, the plasmids we cloned and what experiments we used them for. In general we used standard biobrick cloning techniques as described in the methods section. Whenever we used PCR gene amplification for cloning, we list the primers used in the following table. To be able to co-transform different plasmids we used backbones with compatible origins of replication and resistance genes. In the table you can find which backbone versions we used for which constructs.
| Fluorescent protein reporter constructs |
| |
Description |
Cloning |
Maps |
| 1 |
Receiver cell construct for GFP diffusion experiments |
[http://parts.igem.org/Part:BBa_J09855 BBa_J09855] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_E0840 BBa_E0840] insert (XbaI, PstI) |
![300px][File:Pla2.png](/wiki/images/c/c9/Pla1.png) |
| 2 |
Library of the Receiver cell constructs |
Using the BBa_J09855.BBa_E0840 construct a library with mutated pLux promoters was created through site-saturation mutagenesis to screen for promoters with changed sensitivities
Primers:
5'-tatactagagacnnntaggatcgtacag
5'-gatcgtannngtttacgcaagaaaatg
5'-tagagacnnntaggatcgtannngtttacgcaagaaaatg
5'-tagagaccnntaggatcgtanangtttacgcaagaaaatg
5'-tagagacctntaggatcgtacangtttacgcaagaaaatg
Interesting versions of the promoter were sequenced and inserted into pSB1C3 backbone using custom-made oligos. They could then be used for further cloning. |
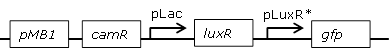 |
| 3 |
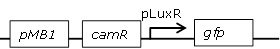
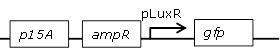
Receiver cell construct for GFP experiments without the LuxR generating part |
[http://parts.igem.org/Part:BBa_R0062 BBa_R0062] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_E0840 BBa_E0840] insert (XbaI, PstI) |
 |
| 4 |
Receiver cell construct for GFP experiments with positive feedback loop to reduce leakiness |
BBa_J09855.BBa_E0840 construct where the pLac promoter was replaced with pLuxR to build a positive feedback loop. The promoter was inserted with two pairs of custom-made oligos using XbaI and HindIII restriction sites.
Oligos:
5’- Ctagagacctgtaggatcgtacaggtttacgcaagaaaatggtttgttatagtcgaataaatactaga
5’- gattaaagaggagaaatactagatgaaaaacataaatgccgacgacacatacagaataattaataaaattaa
5’- aatctctagtatttattcgactataacaaaccattttcttgcgtaaacctgtacgatcctacaggtct
5’- agctttaattttattaattattctgtatgtgtcgtcggcatttatgtttttcatctagtatttctcctcttt |
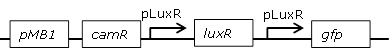 |
| 5 |
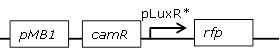
Receiver cell construct with mutated pLux promoter library and RFP reporter to test together with wild-type pLux GFP receiver cells in one cell |
Library of mutated pLux promoters (see above) backbone (SpeI,PstI) and RFP (SpeI,PstI) insert from [http://parts.igem.org/Part:BBa_J23118 BBa_J23118] |
 |
| LuxI generating constructs |
| |
Description |
Cloning |
Maps |
| 6 |
Sender cell construct with a very strong constitutive promoter from the BBa_J23100 promoter library for GFP and Hydrolase experiments |
[http://parts.igem.org/Part:BBa_J23100 BBa_J23100] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_K805016 BBa_K805016] insert (XbaI, PstI) |

 |
| 7 |
Sender cell construct with an intermediate constitutive promoter from the BBa_J23100 promoter library for GFP and Hydrolase experiments |
[http://parts.igem.org/Part:BBa_J23118 BBa_J23118] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_K805016 BBa_K805016] insert (XbaI, PstI) |

 |
| 8 |
Sender cell construct with an intermediate constitutive promoter from the BBa_J23100 promoter library for GFP and Hydrolase experiments |
[http://parts.igem.org/Part:BBa_J23110 BBa_J23110] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_K805016 BBa_K805016] insert (XbaI, PstI) |

 |
| 9 |
Sender cell construct with a weak constitutive promoter from the BBa_J23100 promoter library for GFP and Hydrolase experiments |
[http://parts.igem.org/Part:BBa_J23114 BBa_J23114] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_K805016 BBa_K805016] insert (XbaI, PstI) |

 |
| LuxR generating constructs |
| |
Description |
Cloning |
Maps |
| 10 |
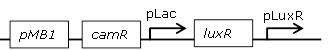
constitutive LuxR generating biobrick |
[http://parts.igem.org/Part:BBa_J09855 BBa_J09855] |
 |
| 11 |
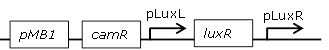
constitutive LuxR generating biobrick, with negative feedback-loop at high OHHL concentrations |
[http://parts.igem.org/Part:BBa_F2621 BBa_F2621] |
 |
| 12 |
negatively regulated pLuxL-LacI construct to improve the leakiness problem of the LuxR system |
[http://parts.igem.org/Part:BBa_R0063 BBa_R0063] backbone (SpeI, PstI) and [http://parts.igem.org/Part:BBa_C0012 BBa_C0012] insert (XbaI, PstI) |
 |
| Hydrolase constructs |
| |
Description |
Cloning |
Maps |
| |
Aes coding region with RBS in SB1C3 backbone |
BBa_K1216002 |
|
Groupparts / Biobricks
In the following table you can find all all the biobricks that were submitted by our group.
<groupparts>iGEM013 ETH_Zurich</groupparts>

 "
"





