Team:Heidelberg/Project/Indigoidine
From 2013.igem.org
m |
m |
||
| Line 137: | Line 137: | ||
<div id="tyrocidineText" class="col-sm-12"> | <div id="tyrocidineText" class="col-sm-12"> | ||
<h2 id="introduction">Introduction</h2> | <h2 id="introduction">Introduction</h2> | ||
| - | + | <p> | |
Most modules of non-ribosomal peptide synthetase (NRPS) pathways consist of three domain types: condensation, adenylation and thiolation domain (see <a class="fancybox fancyFigure" title="NRPS module and domain structure and activation of T-domains." href="https://static.igem.org/mediawiki/2013/a/a9/Heidelberg_IndPD_Fig1.png" rel="gallery1">Figure 1a</a>), also called peptidyl-carrier-protein domain (PCP)-domain (reviewed in <bib id="pmid16895337"/>)). Remarkably, precisely this order of domains is highly conserved among different NRPS pathways except for the very first and last module of each NRPS. In the initial module, the A-domain is always followed by a T-domain. The last module of an NRPS usually ends with a TE-domain. During the process of non-ribisomal peptide (NRP) synthesis, a new amino acid is first adenylated by the A-domain and then bound to the T-domain via a thioester bond. The C-domain catalyzes the condensation of the substrate - which is bound to the T-domain of the previous module - and the amino acid of the next module. The T-domain itself shows no substrate specificity but acts as a carrier domain, which keeps the peptide attached to the NRPS module complex. The core element of every T-domain is a conserved 4’-phosphopanthetheinylated (4’-PPT) serine. The 4’-PPT residue is added by a 4’-Phosphopanthetheinyl-transferase (PPTase), which converts the NRPS apo-enzyme to its active holo-form (see <a class="fancybox fancyFigure" title="NRPS module and domain structure and activation of T-domains." href="https://static.igem.org/mediawiki/2013/a/a9/Heidelberg_IndPD_Fig1.png" rel="gallery1">Figure 1b</a>). | Most modules of non-ribosomal peptide synthetase (NRPS) pathways consist of three domain types: condensation, adenylation and thiolation domain (see <a class="fancybox fancyFigure" title="NRPS module and domain structure and activation of T-domains." href="https://static.igem.org/mediawiki/2013/a/a9/Heidelberg_IndPD_Fig1.png" rel="gallery1">Figure 1a</a>), also called peptidyl-carrier-protein domain (PCP)-domain (reviewed in <bib id="pmid16895337"/>)). Remarkably, precisely this order of domains is highly conserved among different NRPS pathways except for the very first and last module of each NRPS. In the initial module, the A-domain is always followed by a T-domain. The last module of an NRPS usually ends with a TE-domain. During the process of non-ribisomal peptide (NRP) synthesis, a new amino acid is first adenylated by the A-domain and then bound to the T-domain via a thioester bond. The C-domain catalyzes the condensation of the substrate - which is bound to the T-domain of the previous module - and the amino acid of the next module. The T-domain itself shows no substrate specificity but acts as a carrier domain, which keeps the peptide attached to the NRPS module complex. The core element of every T-domain is a conserved 4’-phosphopanthetheinylated (4’-PPT) serine. The 4’-PPT residue is added by a 4’-Phosphopanthetheinyl-transferase (PPTase), which converts the NRPS apo-enzyme to its active holo-form (see <a class="fancybox fancyFigure" title="NRPS module and domain structure and activation of T-domains." href="https://static.igem.org/mediawiki/2013/a/a9/Heidelberg_IndPD_Fig1.png" rel="gallery1">Figure 1b</a>). | ||
| + | </p> | ||
| + | |||
| + | |||
<a class="fancybox fancyGraphical" href="https://static.igem.org/mediawiki/2013/a/a9/Heidelberg_IndPD_Fig1.png"> | <a class="fancybox fancyGraphical" href="https://static.igem.org/mediawiki/2013/a/a9/Heidelberg_IndPD_Fig1.png"> | ||
| Line 159: | Line 162: | ||
</figcaption> | </figcaption> | ||
</a> | </a> | ||
| - | + | <p> | |
Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference]. <br/> | Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference]. <br/> | ||
| - | For example, a single module of <em>P. luminescens laumondii</em> TT01 (DSM15139) contains an internal oxidation domain (Ox-domain) in its A-domain and a special TE-domain (<a class="fancybox fancyFigure" title="Exchange of the indC T-domain on a pSB1C3 derived expression plasmid" href="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png" rel="gallery1">Figure 2a</a>). This enzyme first adenylates L-glutamine (A-domain), which is then attached to the T-domain. The TE-domain cleaves and catalyzes the cyclization of the substrate, which is further oxidized by the Ox-domain. The oxidation of two cyclic glutamines results in the formation of an insoluble small molecule (<a class="fancybox fancyFigure" title="Exchange of the indC T-domain on a pSB1C3 derived expression plasmid" href="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png" rel="gallery1">Figure 2b</a>)[Reference]. < | + | For example, a single module of <em>P. luminescens laumondii</em> TT01 (DSM15139) contains an internal oxidation domain (Ox-domain) in its A-domain and a special TE-domain (<a class="fancybox fancyFigure" title="Exchange of the indC T-domain on a pSB1C3 derived expression plasmid" href="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png" rel="gallery1">Figure 2a</a>). This enzyme first adenylates L-glutamine (A-domain), which is then attached to the T-domain. The TE-domain cleaves and catalyzes the cyclization of the substrate, which is further oxidized by the Ox-domain. The oxidation of two cyclic glutamines results in the formation of an insoluble small molecule (<a class="fancybox fancyFigure" title="Exchange of the indC T-domain on a pSB1C3 derived expression plasmid" href="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png" rel="gallery1">Figure 2b</a>)[Reference]. |
| + | </p> | ||
<a class="fancybox fancyGraphical" href="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png"> | <a class="fancybox fancyGraphical" href="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png"> | ||
<img style="width:60%; margin-bottom:10px; padding:1%;border-style:solid;border-width:1px;border-radius: 5px;" src="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png"></img> | <img style="width:60%; margin-bottom:10px; padding:1%;border-style:solid;border-width:1px;border-radius: 5px;" src="https://static.igem.org/mediawiki/2013/c/ca/Heidelberg_IndPD_Fig2.png"></img> | ||
| Line 170: | Line 174: | ||
The small molecule produced by the pathway described above is a blue-colored pigment called <em>indigoidine</em>. Accordingly, the catalytic NRPS is referred to as <em>indigoidine synthetase</em> or <em>blue pigment synthetase</em> encoded by various bacterial strains such as <em>S. lavendulae subsp. lavendulae</em> (ATCC11924) or <em>P. luminescens</em> (<bib id="Takahashi2007"/><bib id="Brachmann2012"/>). Previous publications showed that replacing the T-domain of the blue pigment synthetase bpsA from <em>S. lavendulae</em> with T-domains of other NRPS modules results in a loss of function, i.e. the engineered indigoidine synthetase does not produce the blue pigment (<bib id="Owen2012"/). So far the only T-domain exchange that resulted in a functional bpsA was achieved using the T-domain of entF, - a gene involved in the enterobactin biosynthesis of <em>E. coli</em>. For this approach, random mutagenesis was performed to yield various mutated entF T-domains, some of which were capable to preserve the enzyme function when being introduced into bpsA (<bib id="Owen2012"/>). Other studies revealed that it is possible to exchange the A-domains of NRPS modules in <em>B. subtilis</em> resulting in modified non-ribosomal peptide products (<bib id="Doekel2000"/> ). Additionally, the selectivity of an NRPS module for specific amino acids could be modified by altering the conserved motif in the active site of the A-domain (<bib id="Thirlway2012"/>). Furthermore, since the endogenous 4'-Phoshopanthetheinyl-transferase (PPTase) entD from <em>E. coli</em> has been reported to exhibit low efficiency in activating heterologous NRPS pathways, most research in the field of NRPS involves co-expression of another PPTase (<bib id="Pfeifer2001"/>), bib id="Takahashi2007"/>). | The small molecule produced by the pathway described above is a blue-colored pigment called <em>indigoidine</em>. Accordingly, the catalytic NRPS is referred to as <em>indigoidine synthetase</em> or <em>blue pigment synthetase</em> encoded by various bacterial strains such as <em>S. lavendulae subsp. lavendulae</em> (ATCC11924) or <em>P. luminescens</em> (<bib id="Takahashi2007"/><bib id="Brachmann2012"/>). Previous publications showed that replacing the T-domain of the blue pigment synthetase bpsA from <em>S. lavendulae</em> with T-domains of other NRPS modules results in a loss of function, i.e. the engineered indigoidine synthetase does not produce the blue pigment (<bib id="Owen2012"/). So far the only T-domain exchange that resulted in a functional bpsA was achieved using the T-domain of entF, - a gene involved in the enterobactin biosynthesis of <em>E. coli</em>. For this approach, random mutagenesis was performed to yield various mutated entF T-domains, some of which were capable to preserve the enzyme function when being introduced into bpsA (<bib id="Owen2012"/>). Other studies revealed that it is possible to exchange the A-domains of NRPS modules in <em>B. subtilis</em> resulting in modified non-ribosomal peptide products (<bib id="Doekel2000"/> ). Additionally, the selectivity of an NRPS module for specific amino acids could be modified by altering the conserved motif in the active site of the A-domain (<bib id="Thirlway2012"/>). Furthermore, since the endogenous 4'-Phoshopanthetheinyl-transferase (PPTase) entD from <em>E. coli</em> has been reported to exhibit low efficiency in activating heterologous NRPS pathways, most research in the field of NRPS involves co-expression of another PPTase (<bib id="Pfeifer2001"/>), bib id="Takahashi2007"/>). | ||
| - | <h2 id=" | + | <h2 id="aim">Aim</h2> |
We want to find the best approach to determine the linker structure between the A-, T- and TE-domain of indC by exchanging the native T-domain with the T-domain of bpsA. | We want to find the best approach to determine the linker structure between the A-, T- and TE-domain of indC by exchanging the native T-domain with the T-domain of bpsA. | ||
We want to proof that it is possible to exchange the indC T-domain with T-domains from other NRPS modules that are not related to indigoidine synthetases and/or the <em>P. luminescens</em> strain | We want to proof that it is possible to exchange the indC T-domain with T-domains from other NRPS modules that are not related to indigoidine synthetases and/or the <em>P. luminescens</em> strain | ||
| Line 195: | Line 199: | ||
As model system, we used the unimodular indigoidine synthetase NRPS from <em>P. luminescens subsp. Laumondii</em> TT01. We predicted the modular composition and domain borders of IndC using our own NRPS-Designer software. | As model system, we used the unimodular indigoidine synthetase NRPS from <em>P. luminescens subsp. Laumondii</em> TT01. We predicted the modular composition and domain borders of IndC using our own NRPS-Designer software. | ||
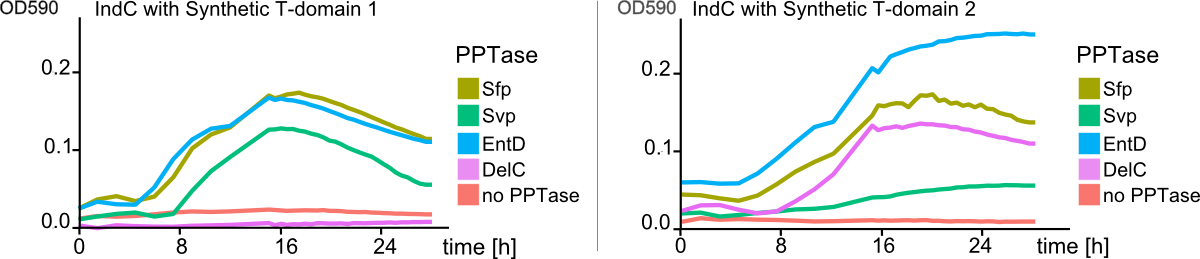
| - | We then started by replacing the native IndC T-domain with T-domains derived from different NRPS pathways from different bacterial strains, among those the T-domain from the BpsA indigoidine synthetase from <em>S. lavendulae<em> ATCC11924. Constructs were transformed into E. coli alongside with an PPTase expression cassette in order to screen for functional IndC variants. As hoped, a subset of the natural T-domains were functioning in context of the IndC scaffold module, leading to indigoidine production and thereby blue coloring of colonies and corresponding liquid cultures. We then engineered a variety of synthetic T-domains derived from consensus sequences of different natural T-domains. Again, a subset of these T-domains successfully maintained indigoidine production (Fig. 9). | + | We then started by replacing the native IndC T-domain with T-domains derived from different NRPS pathways from different bacterial strains, among those the T-domain from the BpsA indigoidine synthetase from <em>S. lavendulae<em> ATCC11924. Constructs were transformed into E. coli alongside with an PPTase expression cassette in order to screen for functional IndC variants. As hoped, a subset of the natural T-domains were functioning in context of the IndC scaffold module, leading to indigoidine production and thereby blue coloring of colonies and corresponding liquid cultures. We then engineered a variety of synthetic T-domains derived from consensus sequences of different natural T-domains. Again, a subset of these T-domains successfully maintained indigoidine production (<a class="fancybox fancyFigure" title="Figure 3" href="https://static.igem.org/mediawiki/2013/6/6e/Heidelberg_IndPD_Fig9.png" rel="gallery1">Fig. 9</a>). |
Notably, one of our engineered IndC construcats showed an indigoidine production even higher compared to the wild-type IndC (T-domain Plu2642; <a class="fancybox fancyFigure" title="Figure 3" href="https://static.igem.org/mediawiki/2013/6/6c/Heidelberg_IndPD_Fig3.png" rel="gallery1">Figure 3</a>). | Notably, one of our engineered IndC construcats showed an indigoidine production even higher compared to the wild-type IndC (T-domain Plu2642; <a class="fancybox fancyFigure" title="Figure 3" href="https://static.igem.org/mediawiki/2013/6/6c/Heidelberg_IndPD_Fig3.png" rel="gallery1">Figure 3</a>). | ||
Revision as of 01:54, 5 October 2013


Indigoidine. Proving Modularity of NRPS by Shuffling Domains.
Abstract
An integral characteristic of synthetic biology yet often undermined is the ability to learn fundamental knowledge by systematically perturbing a biological system. Non-ribosomal peptide synthetases (NRPS) are predestinated for such a trial and error approach. Their hierarchical organization into modules and domains offer a unique opportunity to spin around their inherent logical assembly and observe if their functionality is preserved. Following this idea, we prove the interchangeability of NRPS domains at the example of indC from Photorhabdus luminescens laumondii TT01 (DSM15139). The native NRPS domains have been replaced with domains from other bacterial organisms and fully synthetic domains. To quantify the NRPS efficiency we established an indigoidine assay based on OD measurement of the blue-colored pigment. Interestingly, we find that our data points out the dependence on the T-domain and the 4'-Phosphopanthetheinyl-transferases (PPTases), resulting in different levels of indigoidine synthesis. Furthermore, we introduce HiCT - High throughput protocols for circular polymerase extension Cloning and Transformation - a new standard for the assembly of combinatorial gene libraries (RFC 99).
Introduction
Most modules of non-ribosomal peptide synthetase (NRPS) pathways consist of three domain types: condensation, adenylation and thiolation domain (see Figure 1a), also called peptidyl-carrier-protein domain (PCP)-domain (reviewed in

Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference].
For example, a single module of P. luminescens laumondii TT01 (DSM15139) contains an internal oxidation domain (Ox-domain) in its A-domain and a special TE-domain (Figure 2a). This enzyme first adenylates L-glutamine (A-domain), which is then attached to the T-domain. The TE-domain cleaves and catalyzes the cyclization of the substrate, which is further oxidized by the Ox-domain. The oxidation of two cyclic glutamines results in the formation of an insoluble small molecule (Figure 2b)[Reference].

Aim
We want to find the best approach to determine the linker structure between the A-, T- and TE-domain of indC by exchanging the native T-domain with the T-domain of bpsA. We want to proof that it is possible to exchange the indC T-domain with T-domains from other NRPS modules that are not related to indigoidine synthetases and/or the P. luminescens strain We want to point out the great potential of NRPS for synthetic approaches by designing synthetic T-domains that will result in a functional indigoidine synthetase when introduced to indC We want to show that the activity of NRPS is dependant on the PPTase used to activate them and that the right combination of PPTase and T-domain plays a crucial role when trying to increase the overall product yield. We want to exhibit the potential of NRPS modularity by converting native NRPS modules with glutamine specificity into an indigoidine synthetase by introducing an oxidation domain into the Adenylation-domain. Additionally we will try out various domain combinations to assemble and configure a indigoidine synthetase comprised of different NRPS modules.Results
Engineered indigoidine synthetases retain functionality
We successfully engineered the nonribosomal peptide synthetase indC from P. luminescens subsp. Laumondii TT01 by replacing its native T-domain with both T-domains of other NRPS modules from different bacterial strains and synthetic T-domains of own design, thus creating a library of 58 engineered variants of the indC indigoidine synthetase. So far only the exchange of the T-domain in the related indigoidine synthetase bpsA from S. lavendulae ATCC11924 with the T-domain of the E. coli entF NRPS module was reported, postulating that the T-domain cannot be replaced by other native T-domains retaining the overall protein function, since the engineered variants of bpsA were incapable of producing the blue pigment indigoidine (However, our data shows that the T-domain of indC can be replaced by the T-domain of bpsA only if specific T-domain border combinations are used (see Fig. x) to define the replaced fragment (Figure xa). We proved that indC retained its functionality when the T-domain of the native indC gene was replaced with T-domains of other NRPS modules (see Figure X), applying the T-domain border combination "A2" (see Fig. x). We also tried the T-domain border combination "B2" with the same T-domains, as these T-domain borders were proposed by the Pfam domain prediction tool (pfam.sanger.ac.uk). In this approach less transformants were capable of producing the plue pigment (data not shown), suggesting that the T-domain border combination "A2" can be seen as an improved variant compared to the Pfam prediction.
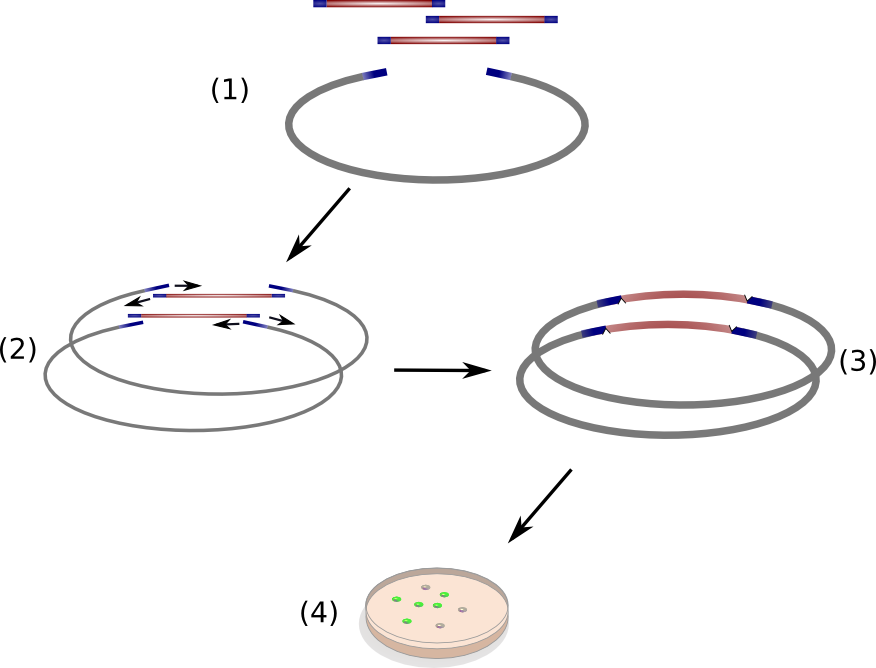
Discussion
In this subproject, we wanted to set the basis for engineering entirely synthetic NRPS modules composed of user-defined domains. As model system, we used the unimodular indigoidine synthetase NRPS from P. luminescens subsp. Laumondii TT01. We predicted the modular composition and domain borders of IndC using our own NRPS-Designer software. We then started by replacing the native IndC T-domain with T-domains derived from different NRPS pathways from different bacterial strains, among those the T-domain from the BpsA indigoidine synthetase from S. lavendulae ATCC11924. Constructs were transformed into E. coli alongside with an PPTase expression cassette in order to screen for functional IndC variants. As hoped, a subset of the natural T-domains were functioning in context of the IndC scaffold module, leading to indigoidine production and thereby blue coloring of colonies and corresponding liquid cultures. We then engineered a variety of synthetic T-domains derived from consensus sequences of different natural T-domains. Again, a subset of these T-domains successfully maintained indigoidine production (Fig. 9). Notably, one of our engineered IndC construcats showed an indigoidine production even higher compared to the wild-type IndC (T-domain Plu2642; Figure 3). This is particularly remarkably as our results contradict to previous studies of NRPS domains that reported the native T-domain of the indigoidine synthetase BpsA to be absolutely essential for protein function (and therefore not replaceable by other T-domains). ([Owen2012]). However, to our surprise, the BpsA T-domain-containing IndC construct did not yield any detectable indigoidine production, although BpsA shares strong sequence homology with IndC. We hypothesized, that the selection of the exact border could be critical for maintaining domain functionality when introduced into a novel NRPS module scaffold. Therefore, we amplified different BpsA T-domain variants differing in their domain border and introduced them into the IndC scaffold. Remarkably, a subset of the resulting IndC variants showed successful indigoidine production. We thus revaluated all native and synthetic T-domains in light of this finding and performed a second screening round in which we were able to rescue even more functional IndC variants, proofing our abovementioned hypothesis. We also co-transformed all engineered IndC construct bearing different natural and synthetic T-domains with four different PPTase expression constructs. To our surprise, the T-domains used not only determined general efficiency of indigoidine production, but also the efficiency of NRPS activation by the different activating PPTases. In conclusion we were able to demonstrate, that it is indeed possible to replace single Domains from NRPS modules, while preserving or even enhancing its functionality. In addition, we established an approach for the design of synthetic T-domains and proved their functionality by introducing them into the indigoidine synthetase indC scaffold. Moreover, we established a high throughput protocol for circular polymerase extension cloning and transformation (Hi-CT) (BBF RFC 99), which we applied for our domain shuffling approach. In summary, we created a library of 58 engineered indC variants. In addition we perforemd measurement of blue pigement production over time, which gave us novel insights in how NRPS domains should be designed, where the domain borders between different domains in a single NRPS module have to be set and which domains from respective NRPS pathways and bacterial strains can be used, when creating novel engineered NRPS pathways. We implemented our findings into the "NRPS-Designer" Software, so that the underlying algorithm for NRPS design takes into consideration the abovementioned findings (e.g. domain border setting) which are certainly crucial for successful in silico prediction of functional NRPSs. Thereby, our project pioneers the research on high-throughput methods for creation of synthetic NRPS modules composed of user-defined domains. We believe that our findings will highly contribute to future development of custom NRPSs.







 "
"


















