Team:NTU-Taida/Notebook/Journal
From 2013.igem.org
(→Transform:) |
(→7/18) |
||
| Line 172: | Line 172: | ||
==7/18== | ==7/18== | ||
| + | ===Inoculation and Incubation at LB broth:=== | ||
| + | B0030-CinR, B0030, LuxR, pLux, pConst(J23119), mTagBFP, Luciferase, simple Las detecting system(K575024), simple Rhl detecting system(K575033) | ||
==7/19== | ==7/19== | ||
Revision as of 01:42, 3 September 2013
Contents |
July
7/1~7/5
Finish RhlR and LasR circuit.
7/1
Transform:
- Positive feedback circuit:
Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G; - Control:
PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G;
(A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
- New:
Pcin, CinR (resistant=C)
7/2
Result:
There’s no colonies at plates of Pcin and CinR.
Transform:
Pcin(resistant=A), CinR (resistant=K)
Inoculation and incubation at LB broth
- Positive feedback circuit: Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G;
- Control: PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G; (A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
7/3
Inoculation and incubation at LB broth
Pcin(resistant=A), CinR (resistant=K)
Check
- Positive feedback circuit: Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G;
- Control: PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G; (A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
7/4
Check and Plasmid DNA extraction (mini-prep):
- Positive feedback circuit: Pc-A-C-B, Pc-A-C-E, Pc-A-C-B, Pc-D-F-B, Pc-D-F-E, Pc-D-F-G;
- Control: PLas-B, PLas-E, PLas-G, PRhl-B, PRhl-E, PRhl-G; (A: RBS-RhlR-tt C: PRhl-RBS-RhlR D: RBS-LasR-tt F: PLas-RBS-LasR B: RBS-mCherry-tt E: RBS-GFPmut-tt G: RBS-mRFP-tt )
- Pcin(resistant=A), CinR (resistant=K)
7/5
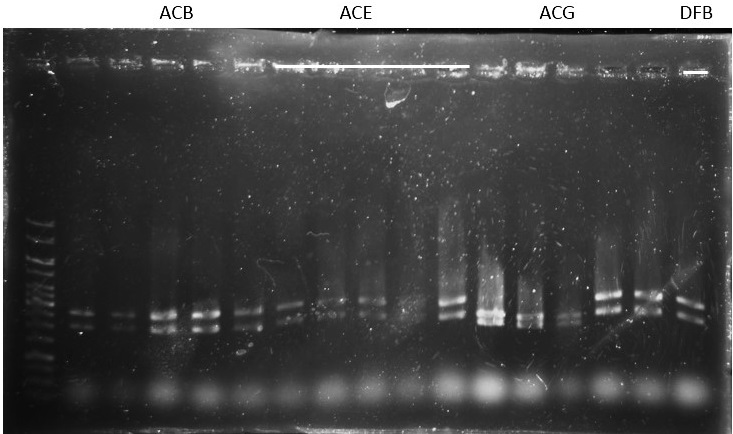
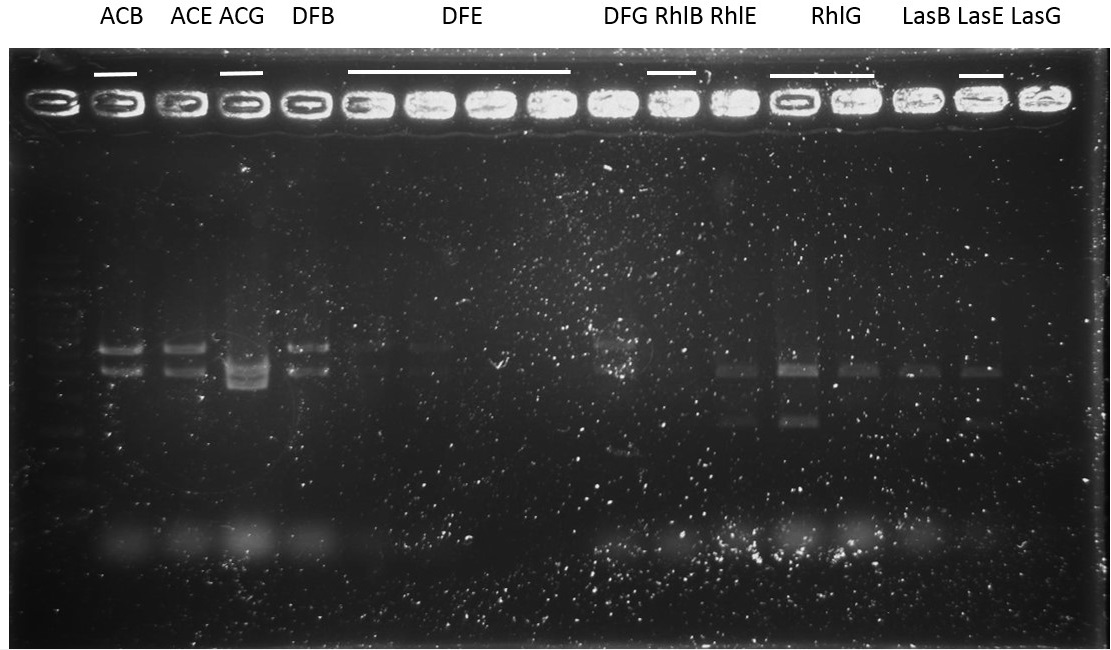
Check: (for plasmid ready to be sequenced)
ACB-4, ACE-5, ACG-1, DFB-1, DFE-2, DFE-3, DFE-4, DFE-5, DFG-1, RhlB-1, RhlE-2, RhlG-1, RhlG-2, LasB3, LasE1, LasG4 (8,9,11 well failed :DFE-4/DFE-5/ RhlB-1)
7/8~7/12
Construct the circuit for new receptor CinR
7/8
Sequence:
ACB-4, ACE-5, ACG-1, DFB-1, DFE-2, DFE-3, DFG-1, RhlE-2, RhlG-1, RhlG-2, LasB3, LasE1, LasG4
Primer list:
| Primer order | Name | sequence | |
|---|---|---|---|
| BMRC-120 | iGEM2013-pSB1A2 | attaccgcctttgagtgagc | R |
| BMRC-153 | iGEM2013-pSB1A2 | gtgccacctgacgtctaagaa | F |
| BMRC-122 | iGEM2013-mCherry | gccgtcctcgaagttcatcac | mcherry |
| BMRC-124 | iGEM2013-mRFP | aacggtaacaccaccgtc | RFP |
| BMRC-126 | iGEM2013-GFP | cttgtagttcccgtcatcttt | GFP |
Digestion, ligation, transform: 3A assembly and standard assembly both
B0030+CinR
7/9
Result:
The plate of B0030+CinR following 3A assembly was OK. The other following standard assembly failed unexpectedly.
Inoculation and incubation:
B0030+cinR
7/10
Plasmid DNA extraction (mini-prep):
B0030-cinR
Result:
A260/A280=1.3
Re-inoculation and incubation of plate of B0030-CinR
7/11
=Plasmid DNA extraction (mini-prep):
B0030-cinR
Result:
A260/A280=1.3
7/12
Digestion: standard assembly only:
B0030 CinR
Result:
There’s no band for B0030
7/15~7/19
CinR, LuxR
7/15
Ligation:
B0030+CinR (The certain content in the ependorf labeled B0030 undergone digestion was actually previously digested; thus, concentration was assumed to be low)
Transform:
B0030-CinR pCI CI
7/16
Result:
Only one colony of plate(B0030-CinR) grew. CI are successfully incubated. pCI failed to grow.
Conclusion:
B0030 threw away.
7/17
Digestion and ligation:
B0030 (eppendorf from early stage)
Transform:
B0030-CinR
RBS(B0030)
pCI(R0051)
LuxR(C0062)
pLux(Lux pR, R0062)
pConst(J23119)
mTagBFP(K592100)
Luciferase(J52008)
simple Las detecting system(K575024)
simple Rhl detecting system(K575033)
Sequence:
PcDFE, PcDFG, Pc ACB, Rhl B, Rhl E Total 16 tube
Sequence results:
No single plasmid was correct at all.
7/18
Inoculation and Incubation at LB broth:
B0030-CinR, B0030, LuxR, pLux, pConst(J23119), mTagBFP, Luciferase, simple Las detecting system(K575024), simple Rhl detecting system(K575033)
7/19
7/22~7/31
7/22
7/23
7/24
7/25
7/26
7/27
7/28
7/29
7/30
7/31
July
2
 "
"